2W8H
 
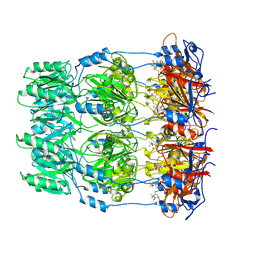 | | Crystal structure of spin labeled Wza24-345. | | Descriptor: | CHLORIDE ION, PUTATIVE OUTER MEMBRANE LIPOPROTEIN WZA, S-[(1-oxyl-2,2,5,5-tetramethyl-2,5-dihydro-1H-pyrrol-3-yl)methyl] methanesulfonothioate | | Authors: | Hagelueken, G, Ingledew, W.J, Huang, H, Petrovic-Stojanovska, B, Whitfield, C, ElMkami, H, Schiemann, O, Naismith, J.H. | | Deposit date: | 2009-01-16 | | Release date: | 2009-02-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.76 Å) | | Cite: | Peldor Distance Fingerprinting of the Octameric Outer-Membrane Protein Wza from Escherichia Coli.
Angew.Chem.Int.Ed.Engl., 48, 2009
|
|
2V7U
 
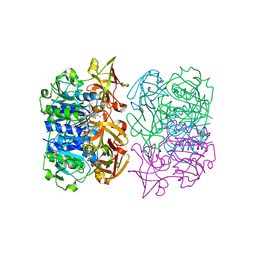 | |
2V7X
 
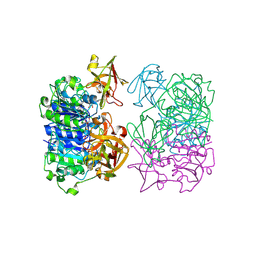 | | X-RAY CRYSTAL STRUCTURE OF 5'-FLUORODEOXYADENOSINE SYNTHASE S158A mutant FROM STREPTOMYCES CATTLEYA COMPLEXED WITH the PRODUCTS, FDA and Met | | Descriptor: | 5'-FLUORO-5'-DEOXYADENOSINE, 5'-FLUORO-5'-DEOXYADENOSINE SYNTHASE, METHIONINE | | Authors: | Robinson, D.A, Zhu, X, O'Hagan, D, Naismith, J.H. | | Deposit date: | 2007-08-02 | | Release date: | 2008-08-12 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Mechanism of enzymatic fluorination in Streptomyces cattleya.
J. Am. Chem. Soc., 129, 2007
|
|
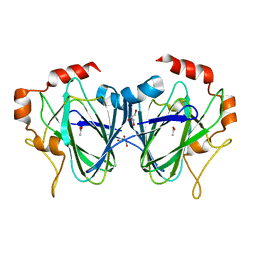
1PM7
 
 | |
1OFN
 
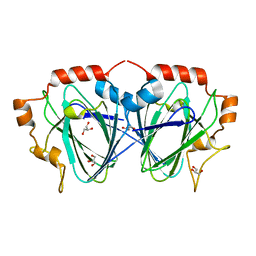 | | Purification, crystallisation and preliminary structural studies of dTDP-4-keto-6-deoxy-glucose-5-epimerase (EvaD) from Amycolatopsis orientalis; the fourth enzyme in the dTDP-L-epivancosamine biosynthetic pathway. | | Descriptor: | GLYCEROL, PCZA361.16 | | Authors: | Merkel, A.B, Naismith, J.H. | | Deposit date: | 2003-04-17 | | Release date: | 2004-04-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Purification, Crystallization and Preliminary Structural Studies of Dtdp-4-Keto-6-Deoxy-Glucose-5-Epimerase (Evad) from Amycolatopsis Orientalis, the Fourth Enzyme in the Dtdp-L-Epivancosamine Biosynthetic Pathway.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
5N4I
 
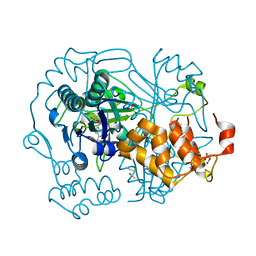 | |
2W0M
 
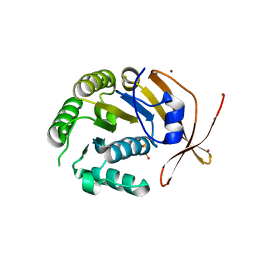 | | Crystal Structure of sso2452 from Sulfolobus solfataricus P2 | | Descriptor: | PYROPHOSPHATE 2-, SSO2452, ZINC ION | | Authors: | McRobbie, A, Carter, L, Johnson, K.A, Kerou, M, Liu, H, Mcmahon, S, Oke, M, Naismith, J.H, White, M.F. | | Deposit date: | 2008-08-19 | | Release date: | 2009-05-19 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Functional Characterisation of a Conserved Archaeal Rada Paralog with Antirecombinase Activity.
J.Mol.Biol., 389, 2009
|
|
2W82
 
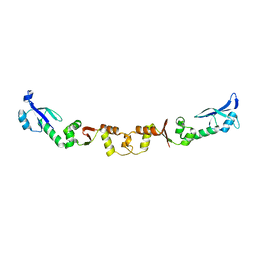 | | The structure of ArdA | | Descriptor: | ORF18 | | Authors: | McMahon, S.A, Roberts, G.A, Carter, L.G, Cooper, L.P, Liu, H, White, J.H, Johnson, K.A, Sanghvi, B, Oke, M, Walkinshaw, M.D, Blakely, G, Naismith, J.H, Dryden, D.T.F. | | Deposit date: | 2009-01-08 | | Release date: | 2009-01-27 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Extensive DNA Mimicry by the Arda Anti-Restriction Protein and its Role in the Spread of Antibiotic Resistance.
Nucleic Acids Res., 37, 2009
|
|
2W8I
 
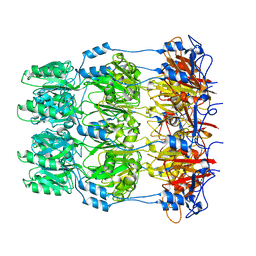 | | Crystal structure of Wza24-345. | | Descriptor: | PUTATIVE OUTER MEMBRANE LIPOPROTEIN WZA | | Authors: | Hagelueken, G, Ingledew, W.J, Huang, H, Petrovic-Stojanovska, B, Whitfield, C, ElMkami, H, Schiemann, O, Naismith, J.H. | | Deposit date: | 2009-01-16 | | Release date: | 2009-02-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Peldor Distance Fingerprinting of the Octameric Outer-Membrane Protein Wza from Escherichia Coli.
Angew.Chem.Int.Ed.Engl., 48, 2009
|
|
2WJA
 
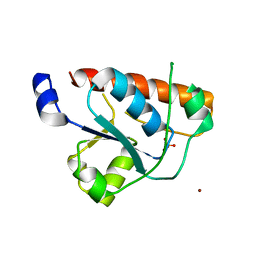 | | Crystal structure of the tyrosine phosphatase Wzb from Escherichia coli K30 in complex with phosphate. | | Descriptor: | NICKEL (II) ION, PHOSPHATE ION, PUTATIVE ACID PHOSPHATASE WZB | | Authors: | Huang, H, Hagelueken, G, Whitfield, C, Naismith, J.H. | | Deposit date: | 2009-05-25 | | Release date: | 2009-07-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structures of Wzb of Escherichia Coli and Cpsb of Streptococcus Pneumoniae, Representatives of Two Families of Tyrosine Phosphatases that Regulate Capsule Assembly.
J.Mol.Biol., 392, 2009
|
|
5N4D
 
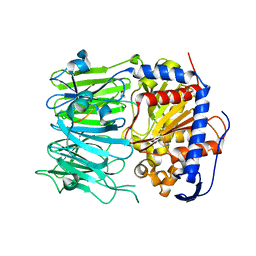 | | Prolyl oligopeptidase B from Galerina marginata bound to 25mer macrocyclization substrate - D661A mutant | | Descriptor: | Alpha-amanitin proprotein, GLYCEROL, Prolyl oligopeptidase | | Authors: | Czekster, C.M, McMahon, S.A, Ludewig, H, Naismith, J.H. | | Deposit date: | 2017-02-10 | | Release date: | 2017-11-01 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Characterization of a dual function macrocyclase enables design and use of efficient macrocyclization substrates.
Nat Commun, 8, 2017
|
|
5O3V
 
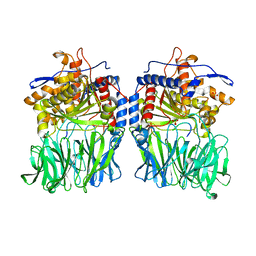 | | Structural characterization of the fast and promiscuous macrocyclase from plant - PCY1-S562A bound to Presegetalin B1 | | Descriptor: | MAGNESIUM ION, Peptide cyclase 1, Putative presegetalin B1, ... | | Authors: | Ludewig, H, Czekster, C.M, Bent, A.F, Naismith, J.H. | | Deposit date: | 2017-05-25 | | Release date: | 2018-02-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Characterization of the Fast and Promiscuous Macrocyclase from Plant PCY1 Enables the Use of Simple Substrates.
ACS Chem. Biol., 13, 2018
|
|
5O3U
 
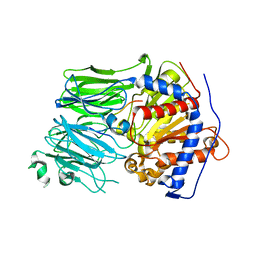 | | Structural characterization of the fast and promiscuous macrocyclase from plant - PCY1-S562A bound to Presegetalin F1 | | Descriptor: | Peptide cyclase 1, Putative presegetalin F1 | | Authors: | Ludewig, H, Czekster, C.M, Bent, A.F, Naismith, J.H. | | Deposit date: | 2017-05-25 | | Release date: | 2018-02-07 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Characterization of the Fast and Promiscuous Macrocyclase from Plant PCY1 Enables the Use of Simple Substrates.
ACS Chem. Biol., 13, 2018
|
|
2X0D
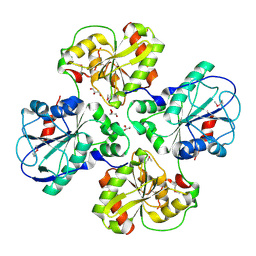 
 | | APO structure of WsaF | | Descriptor: | GLYCEROL, WSAF | | Authors: | Steiner, K, Hagelueken, G, Naismith, J.H. | | Deposit date: | 2009-12-08 | | Release date: | 2010-02-02 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Structural Basis of Substrate Binding in Wsaf, a Rhamnosyltransferase from Geobacillus Stearothermophilus.
J.Mol.Biol., 397, 2010
|
|
5O9C
 
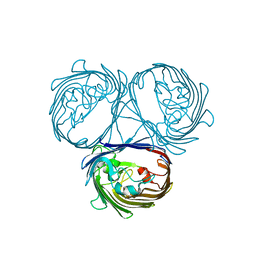 | |
5N4C
 
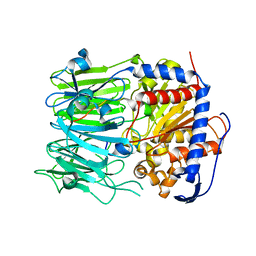 | | Prolyl oligopeptidase B from Galerina marginata bound to 35mer hydrolysis and macrocyclization substrate - S577A mutant | | Descriptor: | Alpha-amanitin proprotein, GLYCEROL, Prolyl oligopeptidase | | Authors: | Czekster, C.M, McMahon, S.A, Ludewig, H, Naismith, J.H. | | Deposit date: | 2017-02-10 | | Release date: | 2017-11-01 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Characterization of a dual function macrocyclase enables design and use of efficient macrocyclization substrates.
Nat Commun, 8, 2017
|
|
5N0R
 
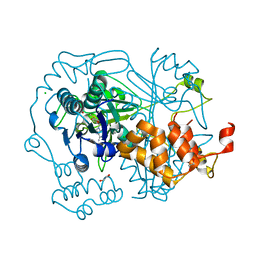 | |
2X0E
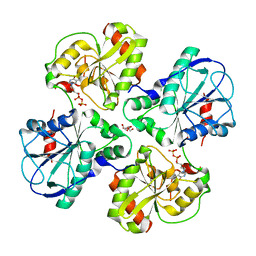 
 | | Complex structure of WsaF with dTDP | | Descriptor: | GLYCEROL, THYMIDINE-5'-DIPHOSPHATE, WSAF | | Authors: | Steiner, K, Hagelueken, G, Naismith, J.H. | | Deposit date: | 2009-12-08 | | Release date: | 2010-02-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Structural Basis of Substrate Binding in Wsaf, a Rhamnosyltransferase from Geobacillus Stearothermophilus.
J.Mol.Biol., 397, 2010
|
|
5O3W
 
 | | Structural characterization of the fast and promiscuous macrocyclase from plant - PCY1-S562A bound to Presegetalin A1 | | Descriptor: | MAGNESIUM ION, Peptide cyclase 1, Presegetalin A1, ... | | Authors: | Ludewig, H, Czekster, C.M, Bent, A.F, Naismith, J.H. | | Deposit date: | 2017-05-25 | | Release date: | 2018-02-07 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Characterization of the Fast and Promiscuous Macrocyclase from Plant PCY1 Enables the Use of Simple Substrates.
ACS Chem. Biol., 13, 2018
|
|
2WJE
 
 | |
2WJF
 
 | | Crystal structure of the tyrosine phosphatase Cps4B from Steptococcus pneumoniae TIGR4 in complex with phosphate. | | Descriptor: | MANGANESE (II) ION, PHOSPHATE ION, TYROSINE-PROTEIN PHOSPHATASE CPSB | | Authors: | Hagelueken, G, Huang, H, Naismith, J.H. | | Deposit date: | 2009-05-25 | | Release date: | 2009-07-14 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Crystal Structures of Wzb of Escherichia Coli and Cpsb of Streptococcus Pneumoniae, Representatives of Two Families of Tyrosine Phosphatases that Regulate Capsule Assembly.
J.Mol.Biol., 392, 2009
|
|
2X68
 
 | | The ternary complex of PrnB (the second enzyme in pyrrolnitrin biosynthesis pathway), 7-Cl-L-tryptophan and cyanide | | Descriptor: | 7-CHLOROTRYPTOPHAN, CYANIDE ION, PRNB, ... | | Authors: | Zhu, X, van pee, K.-H, Naismith, J.H. | | Deposit date: | 2010-02-15 | | Release date: | 2010-03-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | The Ternary Complex of Prnb (the Second Enzyme in Pyrrolnitrin Biosynthesis Pathway), Tryptophan and Cyanide Yields New Mechanistic Insights Into the Indolamine Dioxygenase Superfamily.
J.Biol.Chem., 285, 2010
|
|
2X66
 
 | | The binary complex of PrnB (the second enzyme in pyrrolnitrin biosynthesis pathway) and cyanide | | Descriptor: | CYANIDE ION, PRNB, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Zhu, X, van pee, K.-H, Naismith, J.H. | | Deposit date: | 2010-02-15 | | Release date: | 2010-03-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | The Ternary Complex of Prnb (the Second Enzyme in Pyrrolnitrin Biosynthesis Pathway), Tryptophan and Cyanide Yields New Mechanistic Insights Into the Indolamine Dioxygenase Superfamily.
J.Biol.Chem., 285, 2010
|
|
2X67
 
 | | The ternary complex of PrnB (the second enzyme in pyrrolnitrin biosynthesis pathway), tryptophan and cyanide | | Descriptor: | CYANIDE ION, PRNB, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Zhu, X, van pee, K.-H, Naismith, J.H. | | Deposit date: | 2010-02-15 | | Release date: | 2010-03-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | The Ternary Complex of Prnb (the Second Enzyme in Pyrrolnitrin Biosynthesis Pathway), Tryptophan and Cyanide Yields New Mechanistic Insights Into the Indolamine Dioxygenase Superfamily.
J.Biol.Chem., 285, 2010
|
|
2V7V
 
 | |
