9FCF
 
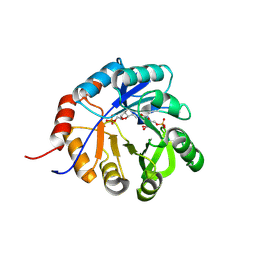 | | Medicago truncatula 5'-ProFAR isomerase (HISN3) D57N mutant in complex with ProFAR | | Descriptor: | 1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino] imidazole-4-carboxamide isomerase, chloroplastic, CHLORIDE ION, ... | | Authors: | Witek, W, Imiolczyk, B, Ruszkowski, M. | | Deposit date: | 2024-05-15 | | Release date: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | Structural, kinetic, and evolutionary peculiarities of HISN3, a plant 5'-ProFAR isomerase.
Plant Physiol Biochem., 215, 2024
|
|
9FGO
 
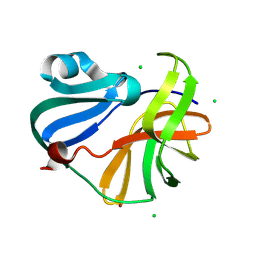 | | Crystal structure of Enterovirus 71 2A protease mutant C110A containing VP1-2A junction in the active site | | Descriptor: | CHLORIDE ION, Polyprotein, ZINC ION | | Authors: | Ni, X, Koekemoer, L, Williams, E.P, Wang, S, Wright, N.D, Godoy, A.S, Aschenbrenner, J.C, Balcomb, B.H, Lithgo, R.M, Marples, P.G, Fairhead, M, Thompson, W, Kirkegaard, K, Fearon, D, Walsh, M.A, von Delft, F. | | Deposit date: | 2024-05-24 | | Release date: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | Crystal structure of Enterovirus 71 2A protease mutant C110A containing VP1-2A junction in the active site
To Be Published
|
|
8BD2
 
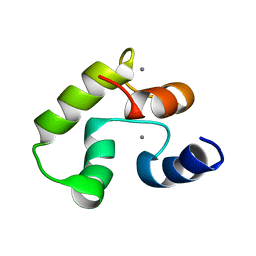 | | Calcium-bound Calmodulin variant G113R | | Descriptor: | CALCIUM ION, Calmodulin-3 | | Authors: | Wimmer, R, Holler, C.V, Petersson, N.M, Iwai, H, Niemelae, M.A, Brohus, M, Overgaard, M.T. | | Deposit date: | 2022-10-18 | | Release date: | 2023-11-08 | | Last modified: | 2024-01-17 | | Method: | SOLUTION NMR | | Cite: | Allosteric changes in protein stability and dynamics as pathogenic mechanism for calmodulin variants not affecting Ca 2+ coordinating residues.
Cell Calcium, 117, 2023
|
|
9J4L
 
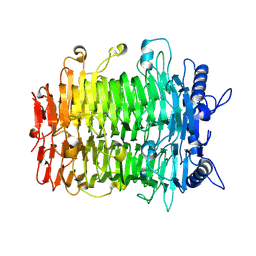 | | Crystal structure of GH9l Inulin fructotransferases (IFTase) | | Descriptor: | DFA-III-forming inulin fructotransferase | | Authors: | Chen, G, Wang, Z.X, Yang, Y.Q, Li, Y.G, Zhang, T, Ouyang, S.Y, Zhang, L, Chen, Y, Ruan, X.L, Miao, M. | | Deposit date: | 2024-08-09 | | Release date: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Elucidation of the mechanism underlying the sequential catalysis of inulin by fructotransferase.
Int.J.Biol.Macromol., 2024
|
|
7PUE
 
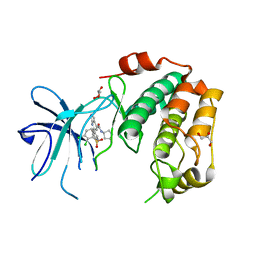 | | Human serum and glucocorticoid-regulated kinase 1 in complex with pyrazolopyridine inhibitor 3a | | Descriptor: | 6-[4-[[2,3-bis(chloranyl)phenyl]sulfonylamino]phenyl]-~{N}-[(3~{R})-pyrrolidin-3-yl]-2~{H}-pyrazolo[3,4-b]pyridine-4-carboxamide, GLYCEROL, Serine/threonine-protein kinase Sgk1 | | Authors: | Dreyer, M.K, Halland, N, Nazare, M. | | Deposit date: | 2021-09-29 | | Release date: | 2021-12-01 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.506 Å) | | Cite: | Rational Design of Highly Potent, Selective, and Bioavailable SGK1 Protein Kinase Inhibitors for the Treatment of Osteoarthritis.
J.Med.Chem., 65, 2022
|
|
8BT3
 
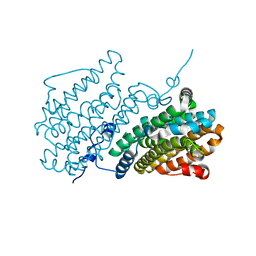 | |
6OBN
 
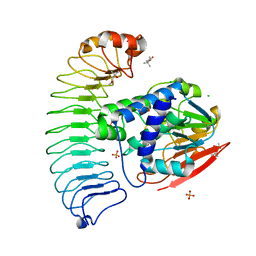 | | The crystal structure of coexpressed SDS22:PP1 complex | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CHLORIDE ION, FE (III) ION, ... | | Authors: | Choy, M.S, Moon, T.M, Bray, J.A, Archuleta, T.L, Shi, W, Peti, W, Page, R. | | Deposit date: | 2019-03-21 | | Release date: | 2019-09-18 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | SDS22 selectively recognizes and traps metal-deficient inactive PP1.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
8BT4
 
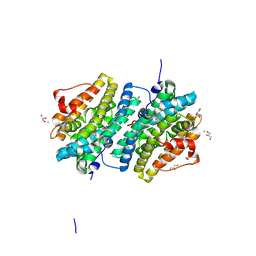 | | Ribonucleotide Reductase class Ie R2 from Mesoplasma florum, radical-lost ground state | | Descriptor: | CALCIUM ION, GLYCEROL, Ribonucleoside-diphosphate reductase | | Authors: | Lebrette, H, Srinivas, V, Hogbom, M. | | Deposit date: | 2022-11-27 | | Release date: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Structure of a ribonucleotide reductase R2 protein radical.
Science, 382, 2023
|
|
5HV9
 
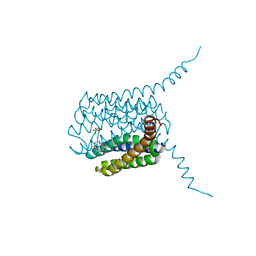 | | Human LTC4S mutant-S36E | | Descriptor: | GLUTATHIONE, Leukotriene C4 synthase, SULFATE ION | | Authors: | Thulasingam, M, Ahmad, H.R.S, Rinaldo-Matthis, A, Haeggstrom, J.Z. | | Deposit date: | 2016-01-28 | | Release date: | 2016-07-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Phosphorylation of Leukotriene C4 Synthase at Serine 36 Impairs Catalytic Activity.
J.Biol.Chem., 291, 2016
|
|
7PQ5
 
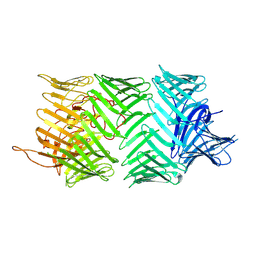 | | Photorhabdus laumondii T6SS-associated Rhs protein carrying the Tre23 toxin domain | | Descriptor: | Tre23 | | Authors: | Jurenas, D, Talachia Rosa, L, Rey, M, Chamot-Rooke, J, Fronzes, R, Cascales, E. | | Deposit date: | 2021-09-16 | | Release date: | 2021-12-01 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (3.17 Å) | | Cite: | Mounting, structure and autocleavage of a type VI secretion-associated Rhs polymorphic toxin.
Nat Commun, 12, 2021
|
|
5EB2
 
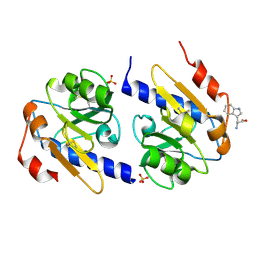 | | Trp-bound YfiR | | Descriptor: | SULFATE ION, TRYPTOPHAN, YfiR | | Authors: | Xu, M, Yang, X, Yang, X.-A, Zhou, L, Liu, T.-Z, Fan, Z, Jiang, T. | | Deposit date: | 2015-10-17 | | Release date: | 2016-05-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.709 Å) | | Cite: | Structural insights into the regulatory mechanism of the Pseudomonas aeruginosa YfiBNR system
Protein Cell, 7, 2016
|
|
8W7Y
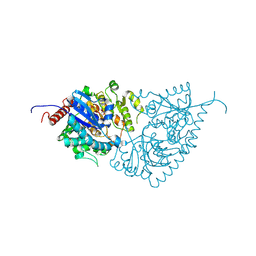 
 | |
4K2P
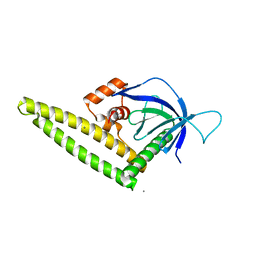 
 | |
7POW
 
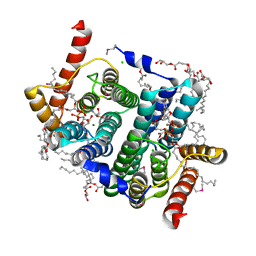 | | Crystal structure of phosphatidyl serine synthase (PSS) in transition state. | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, (2S)-2-amino-3-[[[(2R,3S,4R,5R)-5-(4-amino-2-oxo-pyrimidin-1-yl)-3,4-dihydroxy-tetrahydrofuran-2-yl]methoxy-hydroxy-phosphoryl]oxy-[(2R)-2,3-bis[[(Z)-octadec-9-enoyl]oxy]propoxy]-dihydroxy-lambda^5-phosphanyl]oxy-propanoic acid, 5'-O-[(R)-{[(S)-{(2R)-2,3-bis[(9E)-octadec-9-enoyloxy]propoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]cytidine, ... | | Authors: | Yildiz, O, Centola, M. | | Deposit date: | 2021-09-10 | | Release date: | 2021-12-08 | | Last modified: | 2023-03-01 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Crystal structures of phosphatidyl serine synthase PSS reveal the catalytic mechanism of CDP-DAG alcohol O-phosphatidyl transferases
Nat Commun, 12, 2021
|
|
8BIA
 
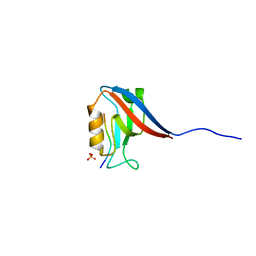 | | Crystal structure of Scribble PDZ1 with PTHR | | Descriptor: | PHOSPHATE ION, Parathyroid hormone/parathyroid hormone-related peptide receptor, Protein scribble homolog | | Authors: | Stewart, B.Z, Kvansakul, M. | | Deposit date: | 2022-11-02 | | Release date: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of Scribble PDZ1 with human papillomavirus strain 16 E6 peptide
To Be Published
|
|
1INW
 
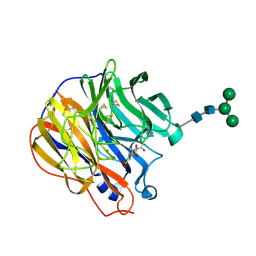 | | A SIALIC ACID DERIVED PHOSPHONATE ANALOG INHIBITS DIFFERENT STRAINS OF INFLUENZA VIRUS NEURAMINIDASE WITH DIFFERENT EFFICIENCIES | | Descriptor: | (1S)-4-acetamido-1,5-anhydro-2,4-dideoxy-1-phosphono-D-glycero-D-galacto-octitol, 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | White, C.L, Janakiraman, M.N, Laver, W.G, Philippon, C, Vasella, A, Air, G.M, Luo, M. | | Deposit date: | 1994-09-26 | | Release date: | 1995-02-07 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | A sialic acid-derived phosphonate analog inhibits different strains of influenza virus neuraminidase with different efficiencies.
J.Mol.Biol., 245, 1995
|
|
8BOE
 
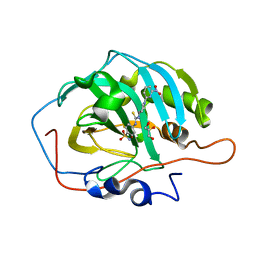 | |
8U1B
 
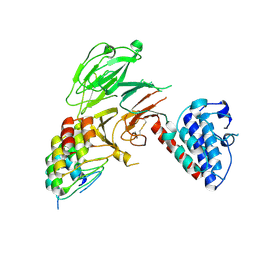 | | C-terminal LRRK2 bound to E11 DARPin | | Descriptor: | E11 DARPin, Leucine-rich repeat serine/threonine-protein kinase 2 | | Authors: | Sanz-Murillo, M, Mathea, S, Dederer, V, Knapp, S, Leschziner, A. | | Deposit date: | 2023-08-31 | | Release date: | 2024-09-04 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Development and characterization of DARPins that bind to LRRK2 WD40 domain
To Be Published
|
|
5HN6
 
 | | Crystal structure of beta-decarboxylating dehydrogenase (TK0280) from Thermococcus kodakarensis complexed with Mn and 3-isopropylmalate | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 3-ISOPROPYLMALIC ACID, Homoisocitrate dehydrogenase, ... | | Authors: | Shimizu, T, Tomita, T, Nishiyama, M. | | Deposit date: | 2016-01-18 | | Release date: | 2016-12-07 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure and function of an ancestral-type beta-decarboxylating dehydrogenase from Thermococcus kodakarensis
Biochem. J., 474, 2017
|
|
8BJ0
 
 | | Crystal structure of Scribble PDZ1 with PTHR | | Descriptor: | CHLORIDE ION, Parathyroid hormone/parathyroid hormone-related peptide receptor, Protein scribble homolog | | Authors: | Stewart, B.Z, Kvansakul, M. | | Deposit date: | 2022-11-03 | | Release date: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of Scribble PDZ1 with human papillomavirus strain 16 E6 peptide
To Be Published
|
|
5OVL
 
 | | crystal structure of MabA bound to NADP+ from M. smegmatis | | Descriptor: | 3-oxoacyl-[acyl-carrier-protein] reductase FabG, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Kussau, T, Van Wyk, N, Viljoen, A, Olieric, V, Flipo, M, Kremer, L, Blaise, M. | | Deposit date: | 2017-08-29 | | Release date: | 2018-02-28 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural rearrangements occurring upon cofactor binding in the Mycobacterium smegmatis beta-ketoacyl-acyl carrier protein reductase MabA.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
5DZ7
 
 | |
5HNF
 
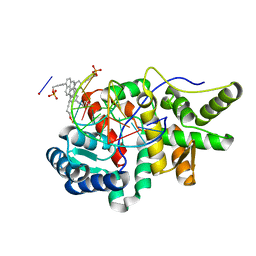 | | Crystal structure of pyrene- and phenanthrene-modified DNA in complex with the BpuJ1 endonuclease binding domain | | Descriptor: | DNA (5'-D(*GP*(YPE)P*AP*CP*CP*CP*GP*TP*GP*GP*A)-3'), DNA (5'-D(*TP*CP*CP*AP*CP*GP*GP*GP*TP*(YPF)P*C)-3'), Restriction endonuclease R.BpuJI | | Authors: | Probst, M, Aeschimann, W, Chau, T.-T.-H, Langenegger, S.M, Stocker, A, Haener, R. | | Deposit date: | 2016-01-18 | | Release date: | 2016-08-17 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.546 Å) | | Cite: | Structural insight into DNA-assembled oligochromophores: crystallographic analysis of pyrene- and phenanthrene-modified DNA in complex with BpuJI endonuclease.
Nucleic Acids Res., 44, 2016
|
|
8BK3
 
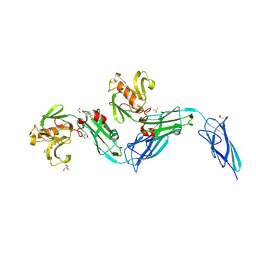 | | Crystal structure of the transpeptidase LdtMt2 from Mycobacterium tuberculosis in complex with diepoxide ketone 1 | | Descriptor: | 1,2-ETHANEDIOL, 1-[(2~{S},3~{R})-3-phenyloxiran-2-yl]ethanone, DIMETHYL SULFOXIDE, ... | | Authors: | de Munnik, M, Schofield, C.J. | | Deposit date: | 2022-11-08 | | Release date: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | alpha beta , alpha ' beta '-Diepoxyketones are mechanism-based inhibitors of nucleophilic cysteine enzymes.
Chem.Commun.(Camb.), 59, 2023
|
|
7PTX
 
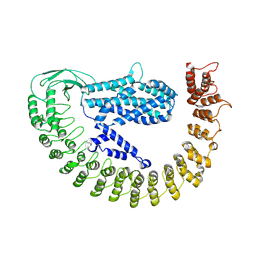 | | Alpha-latrocrustotoxin monomer | | Descriptor: | Alpha-latrocrustotoxin-Lt1a | | Authors: | Chen, M, Gatsogiannis, C. | | Deposit date: | 2021-09-27 | | Release date: | 2021-12-08 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (4.03 Å) | | Cite: | Molecular architecture of black widow spider neurotoxins.
Nat Commun, 12, 2021
|
|
