1YBU
 
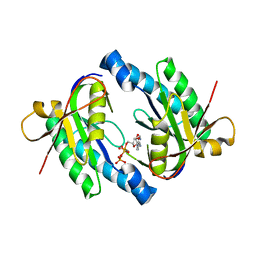 | | Mycobacterium tuberculosis adenylyl cyclase Rv1900c CHD, in complex with a substrate analog. | | Descriptor: | DIPHOSPHOMETHYLPHOSPHONIC ACID ADENOSYL ESTER, MANGANESE (II) ION, lipJ | | Authors: | Sinha, S.C, Wetterer, M, Sprang, S.R, Schultz, J.E, Linder, J.U. | | Deposit date: | 2004-12-21 | | Release date: | 2005-02-15 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Origin of asymmetry in adenylyl cyclases: structures of Mycobacterium tuberculosis Rv1900c.
Embo J., 24, 2005
|
|
1Y11
 
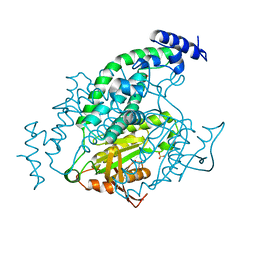 | | Mycobacterial adenylyl cyclase Rv1264, holoenzyme, active state | | Descriptor: | GLYCEROL, Hypothetical protein Rv1264/MT1302, PENTAETHYLENE GLYCOL, ... | | Authors: | Tews, I, Findeisen, F, Sinning, I, Schultz, A, Schultz, J.E, Linder, J.U. | | Deposit date: | 2004-11-16 | | Release date: | 2005-05-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | The structure of a pH-sensing mycobacterial adenylyl cyclase holoenzyme
Science, 308, 2005
|
|
1ZDR
 
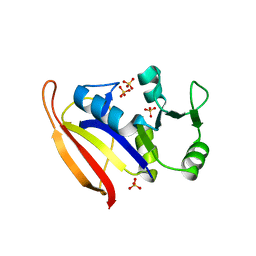 | | DHFR from Bacillus Stearothermophilus | | Descriptor: | GLYCEROL, SULFATE ION, dihydrofolate reductase | | Authors: | Kim, H.S, Damo, S.M, Lee, S.Y, Wemmer, D, Klinman, J.P. | | Deposit date: | 2005-04-14 | | Release date: | 2005-08-30 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure and hydride transfer mechanism of a moderate thermophilic dihydrofolate reductase from Bacillus stearothermophilus and comparison to its mesophilic and hyperthermophilic homologues.
Biochemistry, 44, 2005
|
|
4B1Q
 
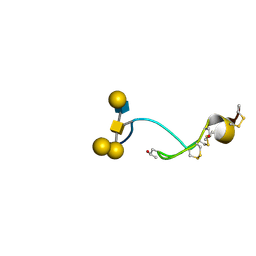 | | NMR structure of the glycosylated conotoxin CcTx from Conus consors | | Descriptor: | CONOTOXIN CCTX, alpha-L-galactopyranose-(1-2)-beta-D-galactopyranose-(1-3)-[alpha-L-galactopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-6)]2-acetamido-2-deoxy-alpha-D-galactopyranose | | Authors: | Hocking, H.G, Gerwig, G.J, Favreau, P, Stocklin, R, Kamerling, J.P, Boelens, R. | | Deposit date: | 2012-07-12 | | Release date: | 2013-02-06 | | Last modified: | 2020-07-29 | | Method: | SOLUTION NMR | | Cite: | Structure of the O-Glycosylated Conopeptide Cctx from Conus Consors Venom.
Chemistry, 19, 2013
|
|
1Y10
 
 | | Mycobacterial adenylyl cyclase Rv1264, holoenzyme, inhibited state | | Descriptor: | CALCIUM ION, Hypothetical protein Rv1264/MT1302, PENTAETHYLENE GLYCOL | | Authors: | Tews, I, Findeisen, F, Sinning, I, Schultz, A, Schultz, J.E, Linder, J.U. | | Deposit date: | 2004-11-16 | | Release date: | 2005-05-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The structure of a pH-sensing mycobacterial adenylyl cyclase holoenzyme
Science, 308, 2005
|
|
1YBT
 
 | | MYCOBACTERIUM TUBERCULOSIS ADENYLYL CYCLASE, RV1900C CHD | | Descriptor: | hydrolase, alpha/beta hydrolase fold family | | Authors: | Sinha, S.C, Wetterer, M, Sprang, S.R, Schultz, J.E, Linder, J.U. | | Deposit date: | 2004-12-21 | | Release date: | 2005-02-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Origin of asymmetry in adenylyl cyclases: structures of Mycobacterium tuberculosis Rv1900c.
Embo J., 24, 2005
|
|
1KNF
 
 | | Crystal Structure of 2,3-dihydroxybiphenyl 1,2-dioxygenase Complexed with 3-methyl Catechol under Anaerobic Condition | | Descriptor: | 2,3-DIHYDROXYBIPHENYL 1,2-DIOXYGENASE, 3-METHYLCATECHOL, FE (II) ION, ... | | Authors: | Han, S, Bolin, J.T. | | Deposit date: | 2001-12-18 | | Release date: | 2002-03-27 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular basis for the stabilization and inhibition of 2, 3-dihydroxybiphenyl 1,2-dioxygenase by t-butanol.
J.Biol.Chem., 273, 1998
|
|
7K3T
 
 | | Crystal Structure of SARS-CoV-2 Main Protease (3CLpro/Mpro) at 1.2 A Resolution and a Possible Capture of Zinc Binding Intermediate | | Descriptor: | 3C-like proteinase, DIMETHYL SULFOXIDE, ZINC ION | | Authors: | Andi, B, Kumaran, D, Kreitler, D.F, Soares, A.S, Shi, W, Jakoncic, J, Fuchs, M.R, Keereetaweep, J, Shanklin, J, McSweeney, S. | | Deposit date: | 2020-09-13 | | Release date: | 2020-09-23 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Hepatitis C virus NS3/4A inhibitors and other drug-like compounds as covalent binders of SARS-CoV-2 main protease.
Sci Rep, 12, 2022
|
|
1KMY
 
 | | Crystal Structure of 2,3-dihydroxybiphenyl 1,2-dioxygenase Complexed with 2,3-dihydroxybiphenyl under Anaerobic Condition | | Descriptor: | 2,3-DIHYDROXYBIPHENYL 1,2-DIOXYGENASE, BIPHENYL-2,3-DIOL, FE (II) ION, ... | | Authors: | Han, S, Bolin, J.T. | | Deposit date: | 2001-12-17 | | Release date: | 2002-02-20 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Molecular basis for the stabilization and inhibition of 2, 3-dihydroxybiphenyl 1,2-dioxygenase by t-butanol.
J.Biol.Chem., 273, 1998
|
|
7TR2
 
 | | Apo CaKip3[2-436]-L2-mutant(HsKHC) in complex with a microtubule | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, Kinesin-like protein,Kinesin-1 heavy chain, ... | | Authors: | Benoit, M.P.M.H, Asenjo, A.B, Hunter, B, Allingham, J.S, Sosa, H. | | Deposit date: | 2022-01-27 | | Release date: | 2022-07-13 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Kinesin-8-specific loop-2 controls the dual activities of the motor domain according to tubulin protofilament shape.
Nat Commun, 13, 2022
|
|
7TR3
 
 | | CaKip3[2-482] - AMP-PNP in complex with a dolastatin-10-stabilized tubulin ring | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, Kinesin-like protein, ... | | Authors: | Benoit, M.P.M.H, Asenjo, A.B, Hunter, B, Allingham, J.S, Sosa, H. | | Deposit date: | 2022-01-27 | | Release date: | 2022-07-20 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Kinesin-8-specific loop-2 controls the dual activities of the motor domain according to tubulin protofilament shape.
Nat Commun, 13, 2022
|
|
1YBJ
 
 | | Structural and Dynamics studies of both apo and holo forms of the hemophore HasA | | Descriptor: | Hemophore HasA | | Authors: | Wolff, N, Izadi-Pruneyre, N, Couprie, J, Habeck, M, Linge, J, Rieping, W, Wandersman, C, Nilges, M, Delepierre, M, Lecroisey, A. | | Deposit date: | 2004-12-21 | | Release date: | 2005-12-27 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Comparative analysis of structural and dynamic properties of the loaded and unloaded hemophore HasA: functional implications.
J.Mol.Biol., 376, 2008
|
|
1YKD
 | | Crystal Structure of the Tandem GAF Domains from a Cyanobacterial Adenylyl Cyclase: Novel Modes of Ligand-Binding and Dimerization | | Descriptor: | ADENOSINE-3',5'-CYCLIC-MONOPHOSPHATE, adenylate cyclase | | Authors: | Martinez, S.E, Bruder, S, Schultz, A, Zheng, N, Schultz, J.E, Beavo, J.A, Linder, J.U. | | Deposit date: | 2005-01-17 | | Release date: | 2005-02-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the tandem GAF domains from a cyanobacterial adenylyl cyclase: Modes of ligand binding and dimerization
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
1YGE
 
 | | LIPOXYGENASE-1 (SOYBEAN) AT 100K | | Descriptor: | FE (III) ION, LIPOXYGENASE-1 | | Authors: | Minor, W, Steczko, J, Stec, B, Otwinowski, Z, Bolin, J.T, Walter, R, Axelrod, B. | | Deposit date: | 1996-06-04 | | Release date: | 1997-07-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Crystal structure of soybean lipoxygenase L-1 at 1.4 A resolution.
Biochemistry, 35, 1996
|
|
1LKD
 | | CRYSTAL STRUCTURE OF 2,3-DIHYDROXYBIPHENYL 1,2-DIOXYGENASE (DHBD) COMPLEXED WITH 2',6'-DICL DIHYDROXYBIPHENYL (DHB) | | Descriptor: | 2',6'-DICHLORO-BIPHENYL-2,6-DIOL, BIPHENYL-2,3-DIOL 1,2-DIOXYGENASE, FE (II) ION, ... | | Authors: | Dai, S, Bolin, J.T. | | Deposit date: | 2002-04-24 | | Release date: | 2002-11-27 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Identification and analysis of a bottleneck in PCB biodegradation
Nat.Struct.Biol., 9, 2002
|
|
1LA2
 | | Structural analysis of Saccharomyces cerevisiae myo-inositol phosphate synthase | | Descriptor: | Myo-inositol-1-phosphate synthase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Kniewel, R, Buglino, J.A, Shen, V, Chadna, T, Beckwith, A, Lima, C.D, Burley, S.K, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2002-03-27 | | Release date: | 2002-04-10 | | Last modified: | 2021-02-03 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structural analysis of Saccharomyces cerevisiae myo-inositol phosphate synthase
J.STRUCT.FUNCT.GENOM., 2, 2002
|
|
1JQS
 
 | | Fitting of L11 protein and elongation factor G (domain G' and V) in the cryo-em map of E. coli 70S ribosome bound with EF-G and GMPPCP, a nonhydrolysable GTP analog | | Descriptor: | 50S Ribosomal protein L11, Elongation Factor G | | Authors: | Agrawal, R.K, Linde, J, Segupta, J, Nierhaus, K.H, Frank, J. | | Deposit date: | 2001-08-07 | | Release date: | 2001-09-07 | | Last modified: | 2024-02-07 | | Method: | ELECTRON MICROSCOPY (18 Å) | | Cite: | Localization of L11 protein on the ribosome and elucidation of its involvement in EF-G-dependent translocation.
J.Mol.Biol., 311, 2001
|
|
1LGT
 
 | | CRYSTAL STRUCTURE OF 2,3-DIHYDROXYBIPHENYL 1,2-DIOXYGENASE (DHBD) COMPLEXED WITH 2'-Cl DIHYDROXYBIPHENYL (DHB) | | Descriptor: | 2'-CHLORO-BIPHENYL-2,3-DIOL, BIPHENYL-2,3-DIOL 1,2-DIOXYGENASE, FE (II) ION | | Authors: | Dai, S, Bolin, J.T. | | Deposit date: | 2002-04-16 | | Release date: | 2002-11-27 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Identification and analysis of a bottleneck in PCB biodegradation
Nat.Struct.Biol., 12, 2002
|
|
1YM3
 
 | | Crystal Structure of carbonic anhydrase RV3588c from Mycobacterium tuberculosis | | Descriptor: | CARBONIC ANHYDRASE (CARBONATE DEHYDRATASE) (CARBONIC DEHYDRATASE), MAGNESIUM ION, ZINC ION | | Authors: | Covarrubias, A.S, Larsson, A.M, Hogbom, M, Lindberg, J, Bergfors, T, Bjorkelid, C, Mowbray, S.L, Unge, T, Jones, T.A, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2005-01-20 | | Release date: | 2005-03-08 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure and function of carbonic anhydrases from Mycobacterium tuberculosis.
J.Biol.Chem., 280, 2005
|
|
1JQM
 
 | | Fitting of L11 protein and elongation factor G (EF-G) in the cryo-em map of e. coli 70S ribosome bound with EF-G, GDP and fusidic acid | | Descriptor: | 50S Ribosomal protein L11, Elongation Factor G | | Authors: | Agrawal, R.K, Linde, J, Segupta, J, Nierhaus, K.H, Frank, J. | | Deposit date: | 2001-08-07 | | Release date: | 2001-09-07 | | Last modified: | 2024-02-07 | | Method: | ELECTRON MICROSCOPY (18 Å) | | Cite: | Localization of L11 protein on the ribosome and elucidation of its involvement in EF-G-dependent translocation.
J.Mol.Biol., 311, 2001
|
|
1JQT
 
 | | Fitting of L11 protein in the low resolution cryo-EM map of E.coli 70S ribosome | | Descriptor: | 50S Ribosomal protein L11 | | Authors: | Agrawal, R.K, Linde, J, Segupta, J, Nierhaus, K.H, Frank, J. | | Deposit date: | 2001-08-07 | | Release date: | 2001-09-07 | | Last modified: | 2024-02-07 | | Method: | ELECTRON MICROSCOPY (18 Å) | | Cite: | Localization of L11 protein on the ribosome and elucidation of its involvement in EF-G-dependent translocation.
J.Mol.Biol., 311, 2001
|
|
1VRU
 
 | | HIGH RESOLUTION STRUCTURES OF HIV-1 RT FROM FOUR RT-INHIBITOR COMPLEXES | | Descriptor: | ALPHA-(2,6-DICHLOROPHENYL)-ALPHA-(2-ACETYL-5-METHYLANILINO)ACETAMIDE, HIV-1 REVERSE TRANSCRIPTASE | | Authors: | Ren, J, Esnouf, R, Garman, E, Somers, D, Ross, C, Kirby, I, Keeling, J, Darby, G, Jones, Y, Stuart, D, Stammers, D. | | Deposit date: | 1995-04-19 | | Release date: | 1996-04-03 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | High resolution structures of HIV-1 RT from four RT-inhibitor complexes.
Nat.Struct.Biol., 2, 1995
|
|
1VRT
 
 | | HIGH RESOLUTION STRUCTURES OF HIV-1 RT FROM FOUR RT-INHIBITOR COMPLEXES | | Descriptor: | 11-CYCLOPROPYL-5,11-DIHYDRO-4-METHYL-6H-DIPYRIDO[3,2-B:2',3'-E][1,4]DIAZEPIN-6-ONE, HIV-1 REVERSE TRANSCRIPTASE, MAGNESIUM ION | | Authors: | Ren, J, Esnouf, R, Garman, E, Somers, D, Ross, C, Kirby, I, Keeling, J, Darby, G, Jones, Y, Stuart, D, Stammers, D. | | Deposit date: | 1995-04-19 | | Release date: | 1996-04-03 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | High resolution structures of HIV-1 RT from four RT-inhibitor complexes.
Nat.Struct.Biol., 2, 1995
|
|
1ZTZ
 
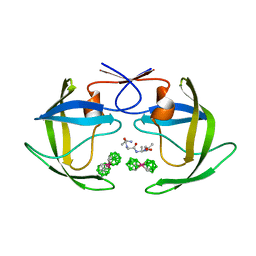 | | Crystal structure of HIV protease- metallacarborane complex | | Descriptor: | COBALT BIS(1,2-DICARBOLLIDE), PROTEASE RETROPEPSIN, autoproteolytic tetrapeptide | | Authors: | Cigler, P, Kozisek, M, Rezacova, P, Brynda, J, Otwinowski, Z, Sedlacek, J, Bodem, J, Kraeusslich, H.-G, Kral, V, Konvalinka, J. | | Deposit date: | 2005-05-28 | | Release date: | 2005-11-01 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | From nonpeptide toward noncarbon protease inhibitors: Metallacarboranes as specific and potent inhibitors of HIV protease
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
1ZYE
 
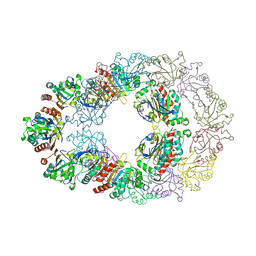 | | Crystal structure analysis of Bovine Mitochondrial Peroxiredoxin III | | Descriptor: | Thioredoxin-dependent peroxide reductase | | Authors: | Cao, Z, Roszak, A.W, Gourlay, L.J, Lindsay, J.G, Isaacs, N.W. | | Deposit date: | 2005-06-10 | | Release date: | 2005-09-20 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Bovine Mitochondrial Peroxiredoxin III Forms a Two-Ring Catenane
Structure, 13, 2005
|
|
