8HOY
 
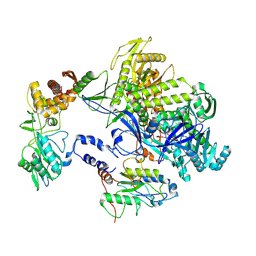 | | Cryo-EM structure of monkeypox virus DNA replication holoenzyme F8, A22 and E4 complex without DNA at 2.76 angostram | | 分子名称: | DNA polymerase, DNA polymerase processivity factor component A20, E4R | | 著者 | Xu, Y, Wu, Y, Zhang, Y, Fan, R, Yang, Y, Li, D, Yang, B, Zhang, Z, Dong, C. | | 登録日 | 2022-12-11 | | 公開日 | 2023-12-13 | | 実験手法 | ELECTRON MICROSCOPY (2.76 Å) | | 主引用文献 | Structure of DNA replication machinery from human monkeypox virus
To Be Published
|
|
8HPA
 
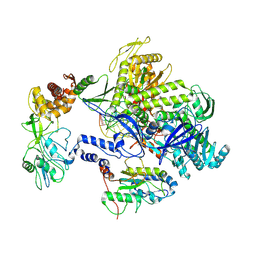 | | Monkeypox virus DNA replication holoenzyme F8, A22 and E4 complex in a DNA binding form | | 分子名称: | DNA (5'-D(*CP*GP*AP*TP*CP*CP*TP*TP*CP*CP*CP*CP*TP*AP*C)-3'), DNA (5'-D(P*AP*TP*GP*GP*TP*AP*GP*GP*GP*GP*AP*AP*GP*GP*AP*TP*CP*G)-3'), DNA polymerase, ... | | 著者 | Xu, Y, Wu, Y, Zhang, Y, Fan, R, Yang, Y, Li, D, Yang, B, Zhang, Z, Dong, C. | | 登録日 | 2022-12-12 | | 公開日 | 2024-01-31 | | 実験手法 | ELECTRON MICROSCOPY (3.01 Å) | | 主引用文献 | Structure of DNA replication machinery from human monkeypox virus
To Be Published
|
|
7V6Z
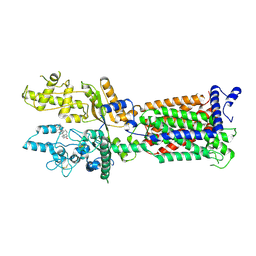 
 | | Cryo-EM structure of Patched1 (V1084A mutant) in lipid nanodisc, 3.64 angstrom (reprocessed with the dataset of 7dzp) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHOLESTEROL, ... | | 著者 | Luo, Y, Zhao, Y, Qu, Q, Li, D. | | 登録日 | 2021-08-20 | | 公開日 | 2021-09-22 | | 最終更新日 | 2022-02-16 | | 実験手法 | ELECTRON MICROSCOPY (3.64 Å) | | 主引用文献 | Cryo-EM study of patched in lipid nanodisc suggests a structural basis for its clustering in caveolae.
Structure, 29, 2021
|
|
7V6Y
 
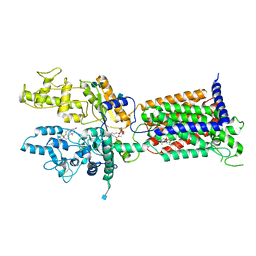 | | Cryo-EM structure of Patched in lipid nanodisc - the wildtype, 3.5 angstrom (re-processed with dataset of 7dzq) | | 分子名称: | (2S)-2-azanyl-3-[[(2S)-3-butanoyloxy-2-dec-9-enoyloxy-propoxy]-oxidanyl-phosphoryl]oxy-propanoic acid, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | 著者 | Luo, Y, Zhao, Y, Qu, Q, Li, D. | | 登録日 | 2021-08-20 | | 公開日 | 2021-09-22 | | 最終更新日 | 2022-02-16 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | Cryo-EM study of patched in lipid nanodisc suggests a structural basis for its clustering in caveolae.
Structure, 29, 2021
|
|
7V4D
 
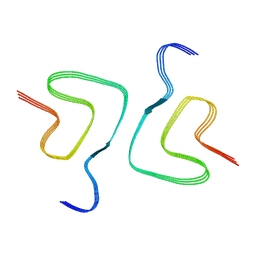 | | Heparin-remodelled alpha-synuclein fibrils | | 分子名称: | Alpha-synuclein | | 著者 | Tao, Y.Q, Sun, Y.P, Xia, W.C, Zhao, Q.Y, Liu, C, Li, D. | | 登録日 | 2021-08-12 | | 公開日 | 2022-08-17 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | Heparin-remodelled alpha-synuclein fibrils
To Be Published
|
|
7WEK
 
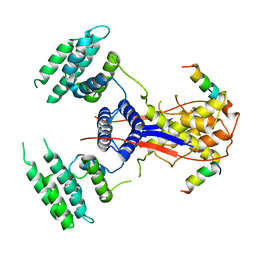 | |
7WEJ
 
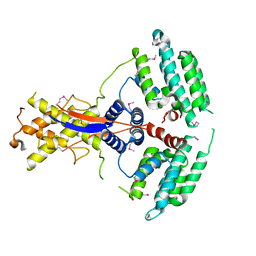 | | Crystal structure of the mouse Wdr47 NTD. | | 分子名称: | WD repeat-containing protein 47 | | 著者 | Ren, J.Q, Li, D, Feng, W. | | 登録日 | 2021-12-23 | | 公開日 | 2022-11-30 | | 実験手法 | X-RAY DIFFRACTION (3.09 Å) | | 主引用文献 | Intertwined Wdr47-NTD dimer recognizes a basic-helical motif in Camsap proteins for proper central-pair microtubule formation.
Cell Rep, 41, 2022
|
|
7FIF
 
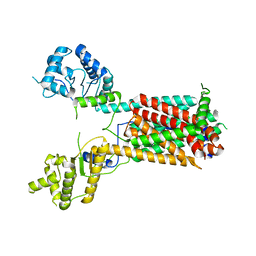 | | Cryo-EM structure of the hedgehog release protein Disp from water bear (Hypsibius dujardini) | | 分子名称: | Protein dispatched-like protein 1 | | 著者 | Luo, Y, Wan, G, Wang, Q, Zhao, Y, Cong, Y, Li, D. | | 登録日 | 2021-07-31 | | 公開日 | 2021-09-08 | | 最終更新日 | 2022-02-16 | | 実験手法 | ELECTRON MICROSCOPY (6.5 Å) | | 主引用文献 | Architecture of Dispatched, a Transmembrane Protein Responsible for Hedgehog Release.
Front Mol Biosci, 8, 2021
|
|
7TPR
 
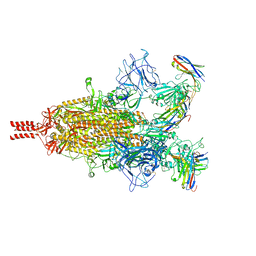 | | Camel nanobodies 7A3 and 8A2 broadly neutralize SARS-CoV-2 variants | | 分子名称: | Nanobody 7A3, Nanobody 8A2, Spike glycoprotein | | 著者 | Butay, K.J, Zhu, J, Dandey, V.P, Hong, J, Kwon, H.J, Chen, C.Z, Duan, Z, Li, D, Ren, H, Liang, T, Martin, N, Esposito, D, Ortega-Rodriguez, U, Xu, M, Xie, H, Ho, M, Cachau, R, Borgnia, M.J. | | 登録日 | 2022-01-25 | | 公開日 | 2022-04-20 | | 実験手法 | ELECTRON MICROSCOPY (2.39 Å) | | 主引用文献 | Camel nanobodies broadly neutralize SARS-CoV-2 variants
bioRxiv, 2021
|
|
6A73
 
 | | Complex structure of CSN2 with IP6 | | 分子名称: | COP9 signalosome complex subunit 2,Endolysin, INOSITOL HEXAKISPHOSPHATE, SULFATE ION | | 著者 | Liu, L, Li, D, Rao, F, Wang, T. | | 登録日 | 2018-07-02 | | 公開日 | 2019-07-03 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.447 Å) | | 主引用文献 | Basis for metabolite-dependent Cullin-RING ligase deneddylation by the COP9 signalosome.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
