8X2V
 
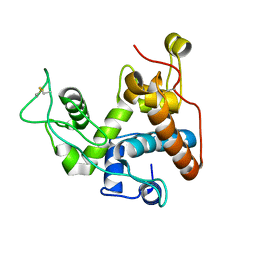 | |
8X2W
 | |
8HNE
 
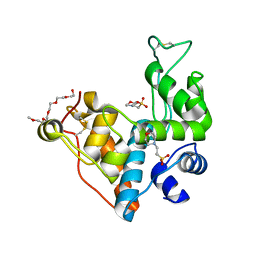 | | Crystal structure of the ancestral GH19 chitinase Anc4 | | Descriptor: | 2-(2-{2-[2-(2-METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHANOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Anc4, ... | | Authors: | Kozome, D, Laurino, P. | | Deposit date: | 2022-12-07 | | Release date: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | Emergence of antifungal activity in GH19 chitinases through gain-of-function in a remote loop.
To Be Published
|
|
8HNF
 
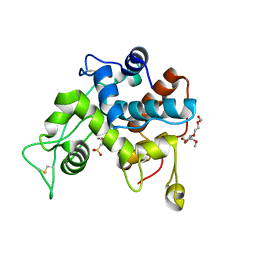 | | Crystal structure of the ancestral GH19 chitinase Anc5 | | Descriptor: | 2-(2-{2-[2-(2-METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHANOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Anc5, ... | | Authors: | Kozome, D, Laurino, P. | | Deposit date: | 2022-12-07 | | Release date: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | Emergence of antifungal activity in GH19 chitinases through gain-of-function in a remote loop.
To Be Published
|
|
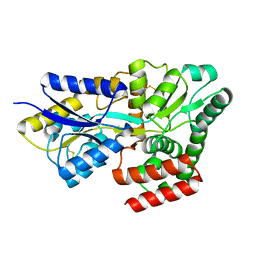
8HQQ
 
 | |
8HQR
 
 | |
7XQN
 
 | |
7XQM
 
 | |
5HQ3
 
 | | Stable, high-expression variant of human acetylcholinesterase | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Acetylcholinesterase, O-ETHYLMETHYLPHOSPHONIC ACID ESTER GROUP | | Authors: | Goldenzweig, A, Goldsmith, M, Hill, S.E, Gertman, O, Laurino, P, Ashani, Y, Dym, O, Albeck, S, Unger, T, Prilusky, J, Lieberman, R.L, Aharoni, A, Silman, I, Sussman, J.L, Tawfik, D.S, Fleishman, S.J. | | Deposit date: | 2016-01-21 | | Release date: | 2016-07-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Automated Structure- and Sequence-Based Design of Proteins for High Bacterial Expression and Stability.
Mol.Cell, 63, 2016
|
|
7LOO
 
 | | S-adenosyl methionine transferase cocrystallized with ATP | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, PHOSPHATE ION, ... | | Authors: | Jackson, C.J, Tan, L.L, Laurino, P. | | Deposit date: | 2021-02-10 | | Release date: | 2021-09-15 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Substrate Dynamics Contribute to Enzymatic Specificity in Human and Bacterial Methionine Adenosyltransferases.
Jacs Au, 1, 2021
|
|
