4V36
 
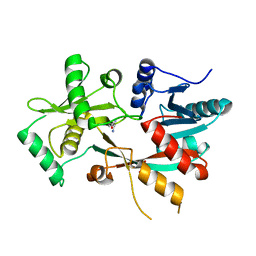 | | The structure of L-PGS from Bacillus licheniformis | | Descriptor: | 2,6-DIAMINO-HEXANOIC ACID AMIDE, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, LYSYL-TRNA-DEPENDENT L-YSYL-PHOSPHATIDYLGYCEROL SYNTHASE | | Authors: | Krausze, J, Hebecker, S, Heinz, D.W, Moser, J. | | Deposit date: | 2014-10-16 | | Release date: | 2015-08-19 | | Last modified: | 2015-09-09 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structures of Two Bacterial Resistance Factors Mediating tRNA-Dependent Aminoacylation of Phosphatidylglycerol with Lysine or Alanine.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
4V34
 
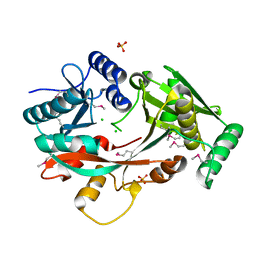 | | The Structure of A-PGS from Pseudomonas aeruginosa (SeMet derivative) | | Descriptor: | ALANYL-TRNA-DEPENDENT L-ALANYL- PHOPHATIDYLGLYCEROL SYNTHASE, CHLORIDE ION, SULFATE ION | | Authors: | Krausze, J, Hebecker, S, Hasenkampf, T, Heinz, D.W, Moser, J. | | Deposit date: | 2014-10-16 | | Release date: | 2015-08-19 | | Last modified: | 2017-07-12 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structures of Two Bacterial Resistance Factors Mediating tRNA-Dependent Aminoacylation of Phosphatidylglycerol with Lysine or Alanine.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
4V35
 
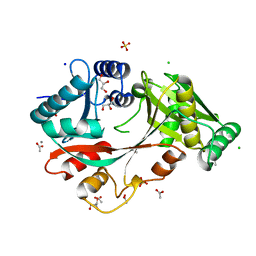 | | The Structure of A-PGS from Pseudomonas aeruginosa | | Descriptor: | ACETATE ION, ALANYL-TRNA-DEPENDENT L-ALANYL- PHOPHATIDYLGLYCEROL SYNTHASE, CALCIUM ION, ... | | Authors: | Krausze, J, Hebecker, S, Hasenkampf, T, Heinz, D.W, Moser, J. | | Deposit date: | 2014-10-16 | | Release date: | 2015-08-19 | | Last modified: | 2015-09-09 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structures of Two Bacterial Resistance Factors Mediating tRNA-Dependent Aminoacylation of Phosphatidylglycerol with Lysine or Alanine.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
5G3U
 
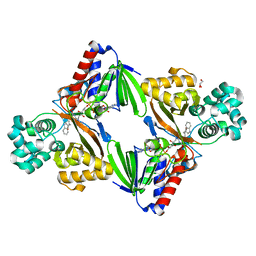 | | The structure of the L-tryptophan oxidase VioA from Chromobacterium violaceum in complex with its inhibitor 2-(1H-indol-3-ylmethyl)prop-2- enoic acid | | Descriptor: | 2-[(1H-indol-3-yl)methyl]prop-2-enoic acid, DIHYDROFLAVINE-ADENINE DINUCLEOTIDE, GLYCEROL, ... | | Authors: | Krausze, J, Rabe, J, Moser, J. | | Deposit date: | 2016-05-01 | | Release date: | 2016-08-03 | | Last modified: | 2019-05-08 | | Method: | X-RAY DIFFRACTION (2.377 Å) | | Cite: | Biosynthesis of Violacein, Structure and Function of l-Tryptophan Oxidase VioA from Chromobacterium violaceum.
J.Biol.Chem., 291, 2016
|
|
5A10
 
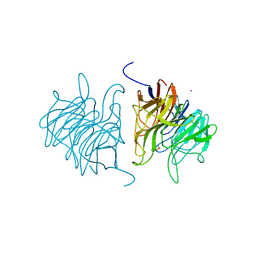 | | The crystal structure of Ta-TFP, a thiocyanate-forming protein involved in glucosinolate breakdown (space group C2) | | Descriptor: | ACETATE ION, SODIUM ION, THIOCYANATE FORMING PROTEIN | | Authors: | Krausze, J, Gumz, F, Wittstock, U. | | Deposit date: | 2015-04-27 | | Release date: | 2015-08-26 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.423 Å) | | Cite: | The Crystal Structure of the Thiocyanate-Forming Protein from Thlaspi Arvense, a Kelch Protein Involved in Glucosinolate Breakdown.
Plant Mol.Biol., 89, 2015
|
|
5A11
 
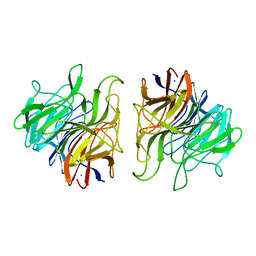 | | The crystal structure of Ta-TFP, a thiocyanate-forming protein involved in glucosinolate breakdown (space group P21) | | Descriptor: | IODIDE ION, THIOCYANATE FORMING PROTEIN | | Authors: | Krausze, J, Gumz, F, Wittstock, U. | | Deposit date: | 2015-04-27 | | Release date: | 2015-08-26 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.47 Å) | | Cite: | The Crystal Structure of the Thiocyanate-Forming Protein from Thlaspi Arvense, a Kelch Protein Involved in Glucosinolate Breakdown.
Plant Mol.Biol., 89, 2015
|
|
5G3S
 
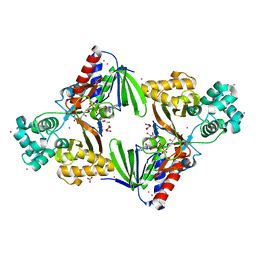 | | The structure of the L-tryptophan oxidase VioA from Chromobacterium violaceum - Samarium derivative | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, DIHYDROFLAVINE-ADENINE DINUCLEOTIDE, ... | | Authors: | Krausze, J, Rabe, J, Moser, J. | | Deposit date: | 2016-05-01 | | Release date: | 2016-08-03 | | Last modified: | 2019-03-06 | | Method: | X-RAY DIFFRACTION (2.076 Å) | | Cite: | Biosynthesis of Violacein: Structure and Function of L-Tryptophan Oxidase Vioa Chromobacterium Violaceum
J.Biol.Chem., 291, 2016
|
|
5G3T
 
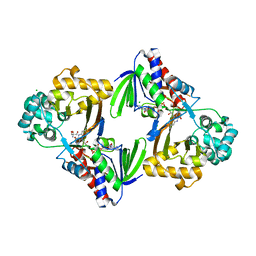 | | The structure of the L-tryptophan oxidase VioA from Chromobacterium violaceum | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, DIHYDROFLAVINE-ADENINE DINUCLEOTIDE, ... | | Authors: | Krausze, J, Rabe, J, Moser, J. | | Deposit date: | 2016-05-01 | | Release date: | 2016-08-03 | | Last modified: | 2019-10-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Biosynthesis of Violacein: Structure and Function of L-Tryptophan Oxidase Vioa Chromobacterium Violaceum
J.Biol.Chem., 291, 2016
|
|
6ROZ
 
 | |
6ROY
 
 | |
6Y01
 
 | |
5G2S
 
 | | Crystal structure of the Mo-insertase domain Cnx1E from Arabidopsis thaliana in complex with molybdate | | Descriptor: | GLYCEROL, MAGNESIUM ION, MOLYBDATE ION, ... | | Authors: | Krausze, J, Probst, C, Kruse, T, Heinz, D.W, Mendel, R.R. | | Deposit date: | 2016-04-13 | | Release date: | 2017-02-15 | | Method: | X-RAY DIFFRACTION (2.838 Å) | | Cite: | Dimerization of the Plant Molybdenum Insertase Cnx1E is Required for Synthesis of the Molybdenum Cofactor.
Biochem.J., 474, 2017
|
|
6Q32
 
 | |
5G2R
 
 | | Crystal structure of the Mo-insertase domain Cnx1E from Arabidopsis thaliana | | Descriptor: | GLYCEROL, MAGNESIUM ION, MOLYBDOPTERIN BIOSYNTHESIS PROTEIN CNX1, ... | | Authors: | Krausze, J, Saha, S, Probst, C, Kruse, T, Heinz, D.W, Mendel, R.R. | | Deposit date: | 2016-04-13 | | Release date: | 2017-02-15 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Dimerization of the Plant Molybdenum Insertase Cnx1E is Required for Synthesis of the Molybdenum Cofactor.
Biochem.J., 474, 2017
|
|
2YNM
 
 | | Structure of the ADPxAlF3-Stabilized Transition State of the Nitrogenase-like Dark-Operative Protochlorophyllide Oxidoreductase Complex from Prochlorococcus marinus with Its Substrate Protochlorophyllide a | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ADENOSINE-5'-DIPHOSPHATE, ALUMINUM FLUORIDE, ... | | Authors: | Krausze, J, Lange, C, Heinz, D.W, Moser, J. | | Deposit date: | 2012-10-16 | | Release date: | 2013-01-30 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of Adp-Aluminium Fluoride-Stabilized Protochlorophyllide Oxidoreductase Complex.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
6RMS
 
 | |
6ETF
 
 | |
6ETH
 
 | |
6ETD
 
 | |
6GB0
 
 | |
6GBC
 
 | |
6GAX
 
 | |
6GBF
 
 | |
6GB4
 
 | |
6GB9
 
 | |
