8PXP
 | |
2I53
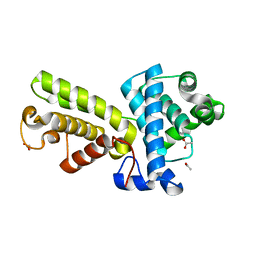 
 | | Crystal structure of Cyclin K | | Descriptor: | ACETATE ION, Cyclin K | | Authors: | Baek, K, Brown, R.S, Birrane, G, Ladias, J.A.A. | | Deposit date: | 2006-08-23 | | Release date: | 2007-01-02 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of Human Cyclin K, a Positive Regulator of Cyclin-dependent Kinase 9.
J.Mol.Biol., 366, 2007
|
|
8RB5
 
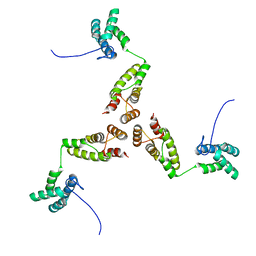 | |
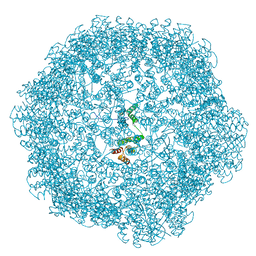
8RB3
 
 | |
8RB7
 
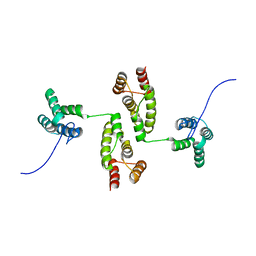 | |
2JGS
 
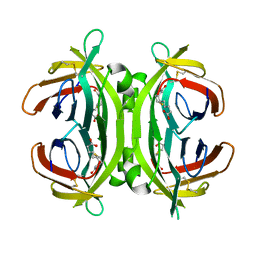 | | Circular permutant of avidin | | Descriptor: | BIOTIN, CIRCULAR PERMUTANT OF AVIDIN | | Authors: | Maatta, J.A.E, Hytonen, V.P, Airenne, T.T, Niskanen, E, Johnson, M.S, Kulomaa, M.S, Nordlund, H.R. | | Deposit date: | 2007-02-14 | | Release date: | 2008-03-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Rational Modification of Ligand-Binding Preference of Avidin by Circular Permutation and Mutagenesis.
Chembiochem, 9, 2008
|
|
2IDH
 
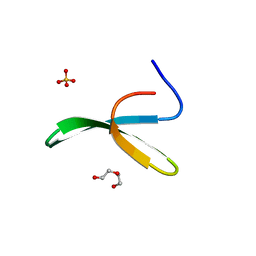 | |
4CDX
 
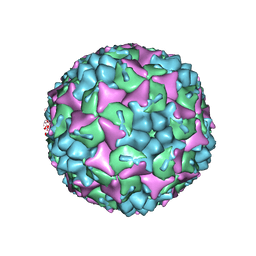 | | Crystal structure of human Enterovirus 71 in complex with the uncoating inhibitor GPP12 | | Descriptor: | 1-(5-((3'-METHYL-[1,1'-BIPHENYL]-4-YL)OXY)PENTYL)-3-(, SODIUM ION, VP1, ... | | Authors: | De Colibus, L, Wang, X, Spyrou, J.A.B, Kelly, J, Ren, J, Grimes, J, Puerstinger, G, Stonehouse, N, Walter, T.S, Hu, Z, Wang, J, Li, X, Peng, W, Rowlands, D, Fry, E.E, Rao, Z, Stuart, D.I. | | Deposit date: | 2013-11-07 | | Release date: | 2014-02-12 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | More-Powerful Virus Inhibitors from Structure-Based Analysis of Hev71 Capsid-Binding Molecules.
Nat.Struct.Mol.Biol., 21, 2014
|
|
4CEY
 
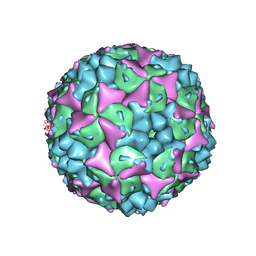 | | Crystal structure of human Enterovirus 71 in complex with the uncoating inhibitor NLD | | Descriptor: | 1-(2-aminopyridin-4-yl)-3-[(3S)-5-{4-[(E)-(ethoxyimino)methyl]phenoxy}-3-methylpentyl]imidazolidin-2-one, SODIUM ION, VP1, ... | | Authors: | De Colibus, L, Wang, X, Spyrou, J.A.B, Kelly, J, Ren, J, Grimes, J, Puerstinger, G, Stonehouse, N, Walter, T.S, Hu, Z, Wang, J, Li, X, Peng, W, Rowlands, D, Fry, E.E, Rao, Z, Stuart, D.I. | | Deposit date: | 2013-11-12 | | Release date: | 2014-02-12 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | More-Powerful Virus Inhibitors from Structure-Based Analysis of Hev71 Capsid-Binding Molecules
Nat.Struct.Mol.Biol., 21, 2014
|
|
6FPP
 
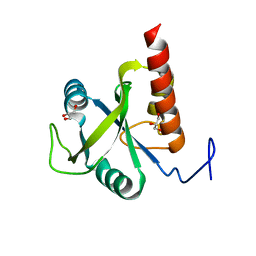 | | Structure of S. pombe Mmi1 | | Descriptor: | GLYCEROL, MAGNESIUM ION, YTH domain-containing protein mmi1 | | Authors: | Stowell, J.A.W, Hill, C.H, Yu, M, Wagstaff, J.L, McLaughlin, S.H, Freund, S.M.V, Passmore, L.A. | | Deposit date: | 2018-02-11 | | Release date: | 2018-05-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | A low-complexity region in the YTH domain protein Mmi1 enhances RNA binding.
J. Biol. Chem., 293, 2018
|
|
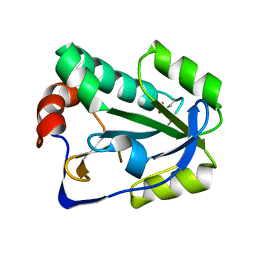
4BPY
 
 | |
8RB4
 
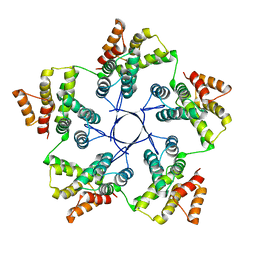 | |
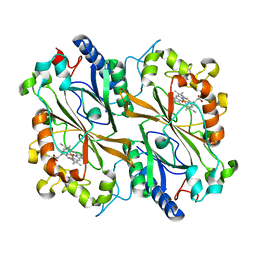
8OLP
 
 | |
2G3W
 
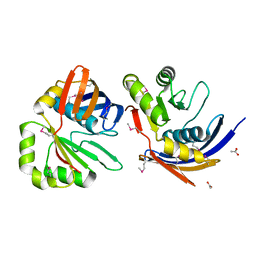 | | The Crystal Structure of YaeQ Protein from Xanthomonas axonopodis pv. citri | | Descriptor: | ACETATE ION, hypothetical protein XAC2396 | | Authors: | Farah, C.S, Guzzo, C.R, Barbosa, J.A.R.G, Nagem, R.A.P. | | Deposit date: | 2006-02-21 | | Release date: | 2007-02-27 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of Xanthomonas axonopodis pv. citri YaeQ reveals a new compact protein fold built around a variation of the PD-(D/E)XK nuclease motif
Proteins, 69, 2007
|
|
2GB8
 
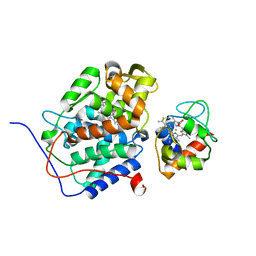 | | Solution structure of the complex between yeast iso-1-cytochrome c and yeast cytochrome c peroxidase | | Descriptor: | Cytochrome c iso-1, Cytochrome c peroxidase, HEME C, ... | | Authors: | Volkov, A.N, Worrall, J.A.R, Ubbink, M. | | Deposit date: | 2006-03-10 | | Release date: | 2006-11-21 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of the complex between cytochrome c and cytochrome c peroxidase determined by paramagnetic NMR.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
8RWY
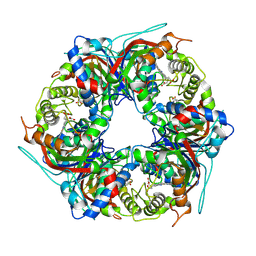 
 | | DtpB hexamer from Streptomyces lividans | | Descriptor: | Dyp-type peroxidase family, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Worrall, J.A.R, Chaplin, A.K, Allport, T. | | Deposit date: | 2024-02-05 | | Release date: | 2024-06-26 | | Method: | ELECTRON MICROSCOPY (3.02 Å) | | Cite: | The oligomeric states of dye-decolorizing peroxidases from Streptomyces lividans and their implications for mechanism of substrate oxidation.
Protein Sci., 33, 2024
|
|
2G81
 
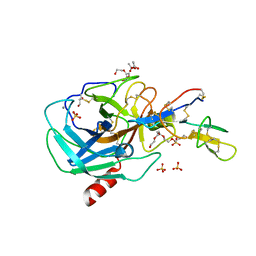 | | Crystal Structure of the Bowman-Birk Inhibitor from Vigna unguiculata Seeds in Complex with Beta-trypsin at 1.55 Angstrons Resolution | | Descriptor: | 1,2-ETHANEDIOL, ACETIC ACID, Bowman-Birk type seed trypsin and chymotrypsin inhibitor, ... | | Authors: | Freitas, S.M, Barbosa, J.A.R.G, Paulino, L.S, Teles, R.C.L, Esteves, G.F, Ventura, M.M. | | Deposit date: | 2006-03-01 | | Release date: | 2007-01-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Crystal Structure of the Bowman-Birk Inhibitor from Vigna unguiculata Seeds in Complex with {beta}-Trypsin at 1.55 A Resolution and Its Structural Properties in Association with Proteinases
Biophys.J., 92, 2007
|
|
6FPQ
 
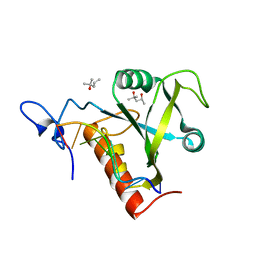 | | Structure of S. pombe Mmi1 in complex with 7-mer RNA | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, RNA (5'-R(*UP*UP*AP*AP*AP*CP*C)-3'), YTH domain-containing protein mmi1 | | Authors: | Stowell, J.A.W, Hill, C.H, Yu, M, Wagstaff, J.L, McLaughlin, S.H, Freund, S.M.V, Passmore, L.A. | | Deposit date: | 2018-02-11 | | Release date: | 2018-05-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | A low-complexity region in the YTH domain protein Mmi1 enhances RNA binding.
J. Biol. Chem., 293, 2018
|
|
2KG2
 
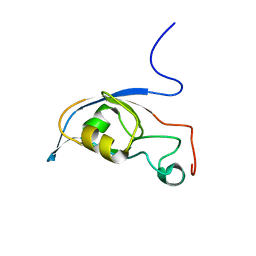 | | Solution structure of a PDZ protein | | Descriptor: | Tax1-binding protein 3 | | Authors: | Durney, M.A, Birrane, G, Anklin, C, Soni, A, Ladias, J.A.A. | | Deposit date: | 2009-03-02 | | Release date: | 2010-01-19 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the human Tax-interacting protein-1.
J.Biomol.Nmr, 45, 2009
|
|
2K71
 
 | | Structure and dynamics of a DNA GNRA hairpin solved vy high-sensitivity NMR with two independent converging methods, simulated annealing (DYANA) and mesoscopic molecular modelling (BCE/AMBER) | | Descriptor: | 5'-D(*DGP*DCP*DGP*DAP*DAP*DAP*DGP*DC)-3' | | Authors: | Santini, G.P.H, Cognet, J.A.H, Xu, D, Singarapu, K.K, Herve du Penhoat, C.L.M. | | Deposit date: | 2008-07-29 | | Release date: | 2009-07-21 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Nucleic acid folding determined by mesoscale modeling and NMR spectroscopy: solution structure of d(GCGAAAGC).
J.Phys.Chem.B, 113, 2009
|
|
8RGS
 
 | | Serial synchrotron in plate room temperature structure of Dye Type Peroxidase Aa | | Descriptor: | Deferrochelatase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Thompson, A.J, Hough, M.A, Williams, L.J, Worrall, J.A.R, Sanchez-Weatherby, J, Mikolajek, H, Sandy, J. | | Deposit date: | 2023-12-14 | | Release date: | 2023-12-27 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Efficient in situ screening of and data collection from microcrystals in crystallization plates.
Acta Crystallogr D Struct Biol, 80, 2024
|
|
8RGY
 
 | | Serial synchrotron in plate room temperature structure of Dye Type Peroxidase Aa, 8 drops merged | | Descriptor: | Deferrochelatase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Thompson, A.J, Hough, M.A, Williams, L.J, Worrall, J.A.R, Sanchez-Weatherby, J, Mikolajek, H, Sandy, J. | | Deposit date: | 2023-12-14 | | Release date: | 2023-12-27 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Efficient in situ screening of and data collection from microcrystals in crystallization plates.
Acta Crystallogr D Struct Biol, 80, 2024
|
|
8RGW
 
 | | Serial synchrotron in plate room temperature structure of Dye Type Peroxidase Aa, 12 drops merged | | Descriptor: | Deferrochelatase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Thompson, A.J, Hough, M.A, Williams, L.J, Worrall, J.A.R, Sanchez-Weatherby, J, Mikolajek, H, Sandy, J. | | Deposit date: | 2023-12-14 | | Release date: | 2023-12-27 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Efficient in situ screening of and data collection from microcrystals in crystallization plates.
Acta Crystallogr D Struct Biol, 80, 2024
|
|
8RGE
 
 | | Serial synchrotron in plate room temperature structure of Lysozyme. | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION | | Authors: | Thompson, A.J, Hough, M.A, Sanchez-Weatherby, J, Williams, L.J, Sandy, J, Worrall, J.A.R. | | Deposit date: | 2023-12-13 | | Release date: | 2023-12-27 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Efficient in situ screening of and data collection from microcrystals in crystallization plates.
Acta Crystallogr D Struct Biol, 80, 2024
|
|
6FPX
 
 | | Structure of S. pombe Mmi1 in complex with 11-mer RNA | | Descriptor: | GLYCEROL, RNA (5'-R(P*UP*UP*UP*AP*AP*AP*CP*CP*UP*A)-3'), YTH domain-containing protein mmi1 | | Authors: | Stowell, J.A.W, Hill, C.H, Yu, M, Wagstaff, J.L, McLaughlin, S.H, Freund, S.M.V, Passmore, L.A. | | Deposit date: | 2018-02-12 | | Release date: | 2018-05-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | A low-complexity region in the YTH domain protein Mmi1 enhances RNA binding.
J. Biol. Chem., 293, 2018
|
|
