1HCW
 
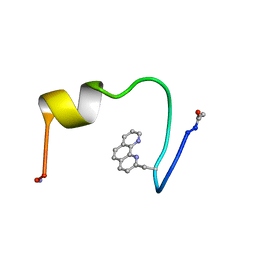 | |
4M9C
 
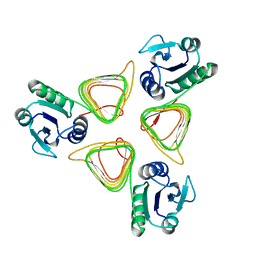 | | WeeI from Acinetobacter baumannii AYE | | Descriptor: | Bacterial transferase hexapeptide (Three repeats) family protein | | Authors: | Morrison, M.J, Imperiali, B. | | Deposit date: | 2013-08-14 | | Release date: | 2013-10-02 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Biochemical analysis and structure determination of bacterial acetyltransferases responsible for the biosynthesis of UDP-N,N'-diacetylbacillosamine.
J.Biol.Chem., 288, 2013
|
|
8G1N
 
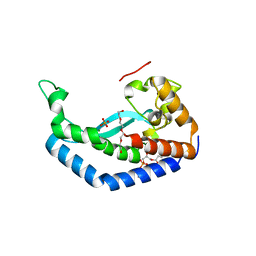 | | Structure of Campylobacter concisus PglC I57M/Q175M Variant with modeled C-terminus | | Descriptor: | DI-PALMITOYL-3-SN-PHOSPHATIDYLETHANOLAMINE, MAGNESIUM ION, N,N'-diacetylbacilliosaminyl-1-phosphate transferase, ... | | Authors: | Dodge, G.J, Ray, L.C, Das, D, Imperiali, B, Allen, K.N. | | Deposit date: | 2023-02-02 | | Release date: | 2023-05-31 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.74 Å) | | Cite: | Co-conserved sequence motifs are predictive of substrate specificity in a family of monotopic phosphoglycosyl transferases.
Protein Sci., 32, 2023
|
|
8T53
 
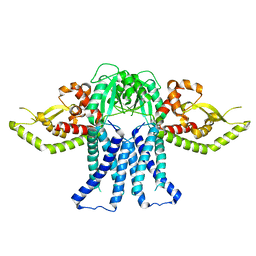 | | S. enterica WbaP in a styrene maleic acid liponanoparticle | | Descriptor: | Undecaprenyl-phosphate galactose phosphotransferase | | Authors: | Dodge, G.J, Imperiali, B. | | Deposit date: | 2023-06-12 | | Release date: | 2024-02-14 | | Last modified: | 2024-02-28 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Mapping the architecture of the initiating phosphoglycosyl transferase from S. enterica O-antigen biosynthesis in a liponanoparticle.
Elife, 12, 2024
|
|
3BSW
 
 | |
3BSY
 
 | |
5TYH
 
 | |
3BSS
 
 | | PglD from Campylobacter jejuni, NCTC 11168, with native substrate | | Descriptor: | Acetyltransferase, UDP-2-acetamido-4-amino-2,4,6-trideoxy-alpha-D-glucopyranose | | Authors: | Olivier, N.B, Imperiali, B. | | Deposit date: | 2007-12-26 | | Release date: | 2008-07-22 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure and catalytic mechanism of PglD from Campylobacter jejuni.
J.Biol.Chem., 283, 2008
|
|
5T2Y
 
 | | Crystal Structure of C. jejuni PglD in complex with 5-methyl-4-(methylamino)-2-phenethylthieno[2,3-d]pyrimidine-6-carboxylic acid | | Descriptor: | 5-methyl-4-(methylamino)-2-(2-phenylethyl)thieno[2,3-d]pyrimidine-6-carboxylic acid, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | De Schutter, J.W, Imperiali, B. | | Deposit date: | 2016-08-24 | | Release date: | 2017-02-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Targeting Bacillosamine Biosynthesis in Bacterial Pathogens: Development of Inhibitors to a Bacterial Amino-Sugar Acetyltransferase from Campylobacter jejuni.
J. Med. Chem., 60, 2017
|
|
3VDZ
 
 | | Tailoring Encodable Lanthanide-Binding Tags as MRI Contrast Agents: xq-dSE3-Ubiquitin at 2.4 Angstroms | | Descriptor: | GADOLINIUM ATOM, SULFATE ION, Ubiquitin-40S ribosomal protein S27a | | Authors: | Daughtry, K.D, Martin, L.J, Surraju, A, Imperiali, B, Allen, K.N. | | Deposit date: | 2012-01-06 | | Release date: | 2012-11-28 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Tailoring encodable lanthanide-binding tags as MRI contrast agents.
Chembiochem, 13, 2012
|
|
1XOF
 
 | | Heterooligomeric Beta Beta Alpha Miniprotein | | Descriptor: | BBAhetT1 | | Authors: | Ali, M.H, Taylor, C.M, Grigoryan, G, Allen, K.N, Imperiali, B, Keating, A.E. | | Deposit date: | 2004-10-06 | | Release date: | 2005-02-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Design of a Heterospecific, Tetrameric, 21-Residue Miniprotein with Mixed alpha/beta Structure.
Structure, 13, 2005
|
|
3POK
 
 | | Interleukin-1-beta LBT L3 Mutant | | Descriptor: | Interleukin-1 beta | | Authors: | Barthelmes, K, Reynolds, A.M, Peisach, E, Jonker, H.R.A, DeNunzio, N.J, Allen, K.N, Imperiali, B, Schwalbe, H. | | Deposit date: | 2010-11-22 | | Release date: | 2011-01-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Engineering encodable lanthanide-binding tags into loop regions of proteins.
J.Am.Chem.Soc., 133, 2011
|
|
8DVW
 
 | | Structure of the Campylobacter concisus glycosyltransferase PglA R203Q | | Descriptor: | N, N'-diacetylbacillosaminyl-diphospho-undecaprenol alpha-1,3-N-acetylgalactosaminyltransferase, URIDINE-DIPHOSPHATE-N-ACETYLGALACTOSAMINE | | Authors: | Vuksanovic, N, Clasman, J.R, Bernstein, H.M, Imperiali, B, Allen, K.N. | | Deposit date: | 2022-07-30 | | Release date: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Specificity determinants revealed by the structure of glycosyltransferase Campylobacter concisus PglA.
Protein Sci., 33, 2024
|
|
8DVZ
 
 | | Structure of the Campylobacter concisus glycosyltransferase PglA R282V variant | | Descriptor: | N, N'-diacetylbacillosaminyl-diphospho-undecaprenol alpha-1,3-N-acetylgalactosaminyltransferase, URIDINE-DIPHOSPHATE-N-ACETYLGALACTOSAMINE | | Authors: | Vuksanovic, N, Clasman, J.R, Bernstein, H.M, Imperiali, B, Allen, K.N. | | Deposit date: | 2022-07-30 | | Release date: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Specificity determinants revealed by the structure of glycosyltransferase Campylobacter concisus PglA.
Protein Sci., 33, 2024
|
|
8DQD
 
 | | Structure of the Campylobacter concisus glycosyltransferase PglA | | Descriptor: | N, N'-diacetylbacillosaminyl-diphospho-undecaprenol alpha-1,3-N-acetylgalactosaminyltransferase, URIDINE-DIPHOSPHATE-N-ACETYLGALACTOSAMINE | | Authors: | Vuksanovic, N, Clasman, J.R, Bernstein, H.M, Imperiali, B, Allen, K.N. | | Deposit date: | 2022-07-18 | | Release date: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Specificity determinants revealed by the structure of glycosyltransferase Campylobacter concisus PglA.
Protein Sci., 33, 2024
|
|
3JXT
 
 | |
3GSL
 
 | |
1TJB
 
 | | Crystal Structure of a High Affinity Lanthanide-Binding Peptide (LBT) | | Descriptor: | CHLORIDE ION, Lanthanide-Binding Peptide, TERBIUM(III) ION | | Authors: | Nitz, M, Sherawat, M, Franz, K.J, Peisach, E, Allen, K.N, Imperiali, B. | | Deposit date: | 2004-06-03 | | Release date: | 2004-08-03 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Origin of the High Affinity of a Chemically Evolved Lanthanide-Binding Peptide
Angew.Chem.Int.Ed.Engl., 43, 2004
|
|
4M99
 
 | |
4M98
 
 | |
1ICL
 | |
8E37
 
 | | Structure of Campylobacter concisus wild-type SeMet PglC | | Descriptor: | N,N'-diacetylbacilliosaminyl-1-phosphate transferase | | Authors: | Vuksanovic, N, Ray, L.C, Imperiali, B, Allen, K.N. | | Deposit date: | 2022-08-16 | | Release date: | 2023-09-06 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3.01 Å) | | Cite: | Synergistic computational and experimental studies of a phosphoglycosyl transferase membrane/ligand ensemble.
J.Biol.Chem., 299, 2023
|
|
1IC9
 | |
1ICO
 
 | |
3NU8
 
 | |
