5DHX
 
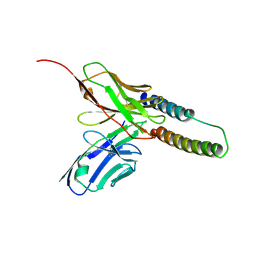 | | HIV-1 Rev NTD dimers with variable crossing angles | | Descriptor: | Anti-Rev Antibody Fab single-chain variable fragment, light chain,Anti-Rev Antibody Fab single-chain variable fragment, heavy chain, ... | | Authors: | DiMattia, M.A, Watts, N.R, Wingfield, P.T, Grimes, J.M, Stuart, D.I, Steven, A.C. | | Deposit date: | 2015-08-31 | | Release date: | 2016-06-22 | | Last modified: | 2017-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | The Structure of HIV-1 Rev Filaments Suggests a Bilateral Model for Rev-RRE Assembly.
Structure, 24, 2016
|
|
2GYS
 
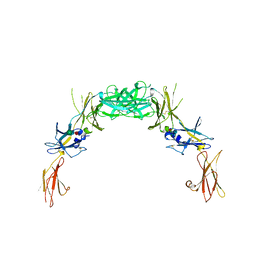 | | 2.7 A structure of the extracellular domains of the human beta common receptor involved in IL-3, IL-5, and GM-CSF signalling | | Descriptor: | Cytokine receptor common beta chain, alpha-L-fucopyranose-(1-3)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)]2-acetamido-2-deoxy-beta-D-glucopyranose, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-3)]2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Carr, P.D, Conlan, F, Ford, S, Ollis, D.L, Young, I.G. | | Deposit date: | 2006-05-09 | | Release date: | 2006-06-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | An improved resolution structure of the human beta common receptor involved in IL-3, IL-5 and GM-CSF signalling which gives better definition of the high-affinity binding epitope.
Acta Crystallogr.,Sect.F, 62, 2006
|
|
2GZ9
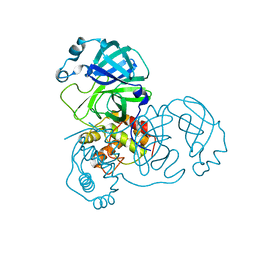 
 | |
2GUU
 
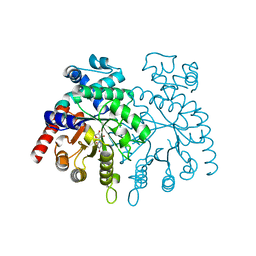 | | crystal structure of Plasmodium vivax orotidine 5-monophosphate decarboxylase with 6-aza-UMP bound | | Descriptor: | 6-AZA URIDINE 5'-MONOPHOSPHATE, ODcase | | Authors: | Dong, A, Lew, J, Zhao, Y, Sundararajan, E, Wasney, G, Vedadi, M, Koeieradzki, I, Edwards, A.M, Arrowsmith, C.H, Weigelt, J, Sundstrom, M, Pai, E.F, Kotra, L, Bochkarev, A, Hui, R, Amani, M, Structural Genomics Consortium (SGC) | | Deposit date: | 2006-05-01 | | Release date: | 2006-05-16 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Crystal structure of Plasmodium vivax orotidine 5-monophosphate decarboxylase with 6-aza-UMP bound
To be Published
|
|
2H1J
 
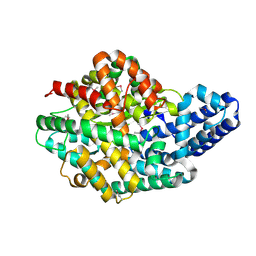 | | 3.1 A X-ray structure of putative Oligoendopeptidase F: Crystals grown by microfluidic seeding | | Descriptor: | Oligoendopeptidase F, ZINC ION | | Authors: | Gerdts, C.J, Tereshko, V, Dementieva, I, Collart, F, Joachimiak, A, Kossiakoff, A, Ismagilov, R.F, Accelerated Technologies Center for Gene to 3D Structure (ATCG3D) | | Deposit date: | 2006-05-16 | | Release date: | 2006-06-13 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Time-Controlled Microfluidic Seeding in nL-Volume Droplets To Separate Nucleation and Growth Stages of Protein Crystallization.
Angew.Chem.Int.Ed.Engl., 45, 2006
|
|
2BF0
 
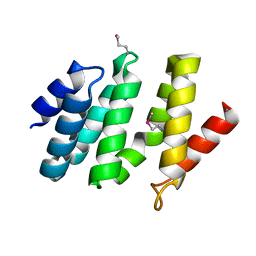 | | crystal structure of the rpr of pcf11 | | Descriptor: | CALCIUM ION, PCF11 | | Authors: | Noble, C.G, Hollingworth, D, Martin, S.R, Adeniran, V.E, Smerdon, S.J, Kelly, G, Taylor, I.A, Ramos, A. | | Deposit date: | 2004-12-02 | | Release date: | 2005-01-18 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Key Features of the Interaction between Pcf11 Cid and RNA Polymerase II Ctd.
Nat.Struct.Mol.Biol., 12, 2005
|
|
2C1N
 | | Molecular basis for the recognition of phosphorylated and phosphoacetylated histone H3 by 14-3-3 | | Descriptor: | 14-3-3 PROTEIN ZETA/DELTA, HISTONE H3 ACETYLPHOSPHOPEPTIDE | | Authors: | Welburn, J.P.I, Macdonald, N, Noble, M.E.M, Nguyen, A, Yaffe, M.B, Clynes, D, Moggs, J.G, Orphanides, G, Thomson, S, Edmunds, J.W, Clayton, A.L, Endicott, J.A, Mahadevan, L.C. | | Deposit date: | 2005-09-16 | | Release date: | 2005-11-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Molecular Basis for the Recognition of Phosphorylated and Phosphoacetylated Histone H3 by 14-3-3.
Mol.Cell, 20, 2005
|
|
2KER
 | | alpha-amylase inhibitor Parvulustat (Z-2685) from Streptomyces parvulus | | Descriptor: | Alpha-amylase inhibitor Z-2685 | | Authors: | Rehm, S, Han, S, Hassani, I, Sokocevic, A, Jonker, H.R.A, Engels, J.W, Schwalbe, H. | | Deposit date: | 2009-02-02 | | Release date: | 2009-02-17 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | The high resolution NMR structure of parvulustat (Z-2685) from Streptomyces parvulus FH-1641: comparison with tendamistat from Streptomyces tendae 4158
Chembiochem, 10, 2009
|
|
3P1E
 
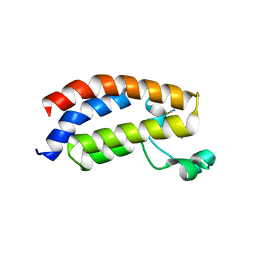 | | Crystal structure of the bromodomain of human CREBBP in complex with dimethyl sulfoxide (DMSO) | | Descriptor: | CREB-binding protein, DIMETHYL SULFOXIDE, POTASSIUM ION, ... | | Authors: | Filippakopoulos, P, Picaud, S, Feletar, I, Fedorov, O, Muniz, J, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Weigelt, J, Bountra, C, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-09-30 | | Release date: | 2010-11-24 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of the bromodomain of human CREBBP in complex with dimethyl sulfoxide (DMSO)
To be Published
|
|
5DQV
 
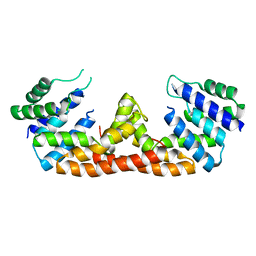 | | The crystal structure of Bacillus subtilis YpgQ | | Descriptor: | NICKEL (II) ION, Uncharacterized protein | | Authors: | Jeon, Y.J, Song, W.S, Yoon, S.I. | | Deposit date: | 2015-09-15 | | Release date: | 2016-04-27 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and biochemical characterization of bacterial YpgQ protein reveals a metal-dependent nucleotide pyrophosphohydrolase
J.Struct.Biol., 195, 2016
|
|
2C61
 
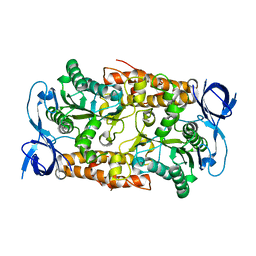 | | Crystal structure of the non-catalytic B subunit of A-type ATPase from M. mazei Go1 | | Descriptor: | A-TYPE ATP SYNTHASE NON-CATALYTIC SUBUNIT B | | Authors: | Schaefer, I, Bailer, S.M, Dueser, M, Boersch, M, Bernal, R.A, Grueber, G, Stock, D. | | Deposit date: | 2005-11-06 | | Release date: | 2006-12-11 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of the Archaeal A1Ao ATP Synthase Subunit B from Methanosarcina Mazei Go1: Implications of Nucleotide-Binding Differences in the Major A1Ao Subunits a and B. Subunits a and B
J.Mol.Biol., 358, 2006
|
|
2KJA
 
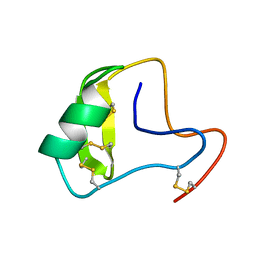 | | Chemical shift assignments, constraints, and coordinates for CN5 scorpion toxin | | Descriptor: | Beta-toxin Cn5 | | Authors: | Prochnicka-Chalufour, A, Corzo, G, Garcia, B.I, Possani, L.D, Delepierre, M. | | Deposit date: | 2009-05-26 | | Release date: | 2009-10-06 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of Cn5, a crustacean toxin found in the venom of the scorpions Centruroides noxius and Centruroides suffusus suffusus.
Biochim.Biophys.Acta, 1794, 2009
|
|
2BKF
 | |
2C4C
 
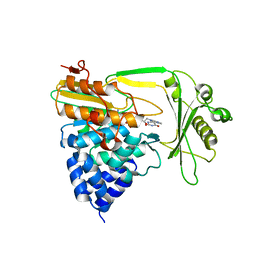 | | Crystal structure of the NADPH-treated monooxygenase domain of MICAL | | Descriptor: | CHLORIDE ION, FLAVIN-ADENINE DINUCLEOTIDE, NEDD9-INTERACTING PROTEIN WITH CALPONIN HOMOLOGY AND LIM DOMAINS | | Authors: | Siebold, C, Berrow, N, Walter, T.S, Harlos, K, Owens, R.J, Terman, J.R, Stuart, D.I, Kolodkin, A.L, Pasterkamp, R.J, Jones, E.Y. | | Deposit date: | 2005-10-18 | | Release date: | 2005-10-26 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | High-Resolution Structure of the Catalytic Region of Mical (Molecule Interacting with Casl), a Multidomain Flavoenzyme-Signaling Molecule.
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
2C5Y
 
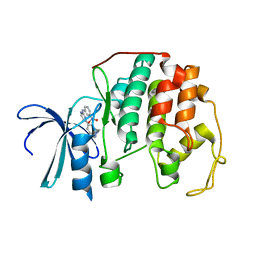 | | DIFFERENTIAL BINDING OF INHIBITORS TO ACTIVE AND INACTIVE CDK2 PROVIDES INSIGHTS FOR DRUG DESIGN | | Descriptor: | CELL DIVISION PROTEIN KINASE 2, HYDROXY(OXO)(3-{[(2Z)-4-[3-(1H-1,2,4-TRIAZOL-1-YLMETHYL)PHENYL]PYRIMIDIN-2(5H)-YLIDENE]AMINO}PHENYL)AMMONIUM | | Authors: | Kontopidis, G, McInnes, C, Pandalaneni, S.R, McNae, I, Gibson, D, Mezna, M, Thomas, M, Wood, G, Wang, S, Walkinshaw, M.D, Fischer, P.M. | | Deposit date: | 2005-11-03 | | Release date: | 2006-03-01 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Differential Binding of Inhibitors to Active and Inactive Cdk2 Provides Insights for Drug Design.
Chem.Biol., 13, 2006
|
|
5DYR
 
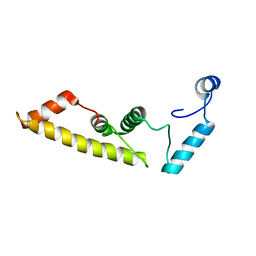 | | Structure of virulence-associated protein D (VapD) from Xylella fastidiosa | | Descriptor: | Virulence-associated protein D | | Authors: | Kochneva, M.V, dos Santos, M.L, dos Santos, C.A, de Souza, A.P, Polikarpov, I, Aparicio, R, Golubev, A.M. | | Deposit date: | 2015-09-25 | | Release date: | 2016-01-27 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure of virulence-associated protein D (VapD) from Xylella fastidiosa
To Be Published
|
|
2BX6
 
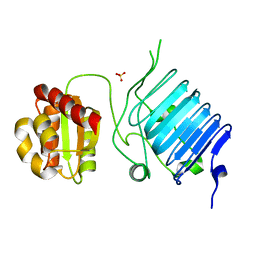 | | Crystal Structure of the human Retinitis Pigmentosa protein 2 (RP2) | | Descriptor: | SULFATE ION, XRP2 PROTEIN | | Authors: | Kuhnel, K, Veltel, S, Schlichting, I, Wittinghofer, A. | | Deposit date: | 2005-07-24 | | Release date: | 2006-01-18 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structure of the Human Retinitis Pigmentosa 2 Protein and its Interaction with Arl3
Structure, 14, 2006
|
|
1AD5
 
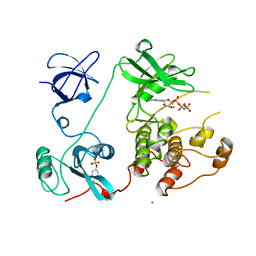 | | SRC FAMILY KINASE HCK-AMP-PNP COMPLEX | | Descriptor: | CALCIUM ION, HAEMATOPOETIC CELL KINASE HCK, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Sicheri, F, Moarefi, I, Kuriyan, J. | | Deposit date: | 1997-02-20 | | Release date: | 1997-05-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of the Src family tyrosine kinase Hck.
Nature, 385, 1997
|
|
2C1G
 
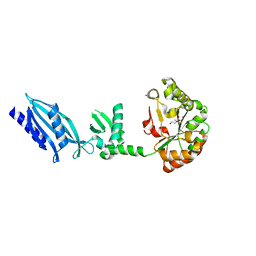 | | Structure of Streptococcus pneumoniae peptidoglycan deacetylase (SpPgdA) | | Descriptor: | ACETATE ION, DI(HYDROXYETHYL)ETHER, PEPTIDOGLYCAN GLCNAC DEACETYLASE, ... | | Authors: | Blair, D.E, Schuttelkopf, A.W, MacRae, J.I, van Aalten, D.M.F. | | Deposit date: | 2005-09-14 | | Release date: | 2005-09-15 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure and metal-dependent mechanism of peptidoglycan deacetylase, a streptococcal virulence factor.
Proc. Natl. Acad. Sci. U.S.A., 102, 2005
|
|
3N6T
 
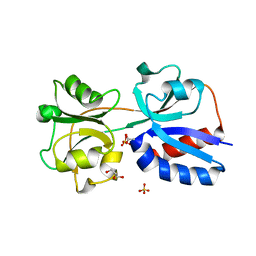 | | Effector binding domain of TsaR | | Descriptor: | ACETATE ION, GLYCEROL, LysR type regulator of tsaMBCD, ... | | Authors: | Monferrer, D, Uson, I. | | Deposit date: | 2010-05-26 | | Release date: | 2011-06-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural studies on the effector binding domain of TsaR
To be Published
|
|
3N7Q
 
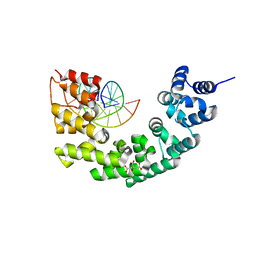 | | Crystal structure of human mitochondrial mTERF fragment (aa 99-399) in complex with a 12-mer DNA encompassing the tRNALeu(UUR) binding sequence | | Descriptor: | CITRIC ACID, DNA (5'-D(*CP*CP*GP*GP*GP*CP*TP*CP*TP*GP*CP*C)-3'), DNA (5'-D(*GP*GP*CP*AP*GP*AP*GP*CP*CP*CP*GP*G)-3'), ... | | Authors: | Jimenez-Menendez, N, Fernandez-Millan, P, Rubio-Cosials, A, Arnan, C, Montoya, J, Jacobs, H.T, Bernado, P, Coll, M, Uson, I, Sola, M. | | Deposit date: | 2010-05-27 | | Release date: | 2010-06-16 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Human mitochondrial mTERF wraps around DNA through a left-handed superhelical tandem repeat.
Nat.Struct.Mol.Biol., 17, 2010
|
|
2K7K
 
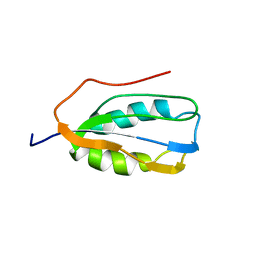 | | Human Acylphosphatase (AcPh) common type | | Descriptor: | Acylphosphatase-1 | | Authors: | Gribenko, A.V, Patel, M.M, Liu, J, McCallum, S.A, Wang, C, Makhatadze, G.I. | | Deposit date: | 2008-08-13 | | Release date: | 2009-02-17 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Rational stabilization of enzymes by computational redesign of surface charge-charge interactions
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
1A8S
 
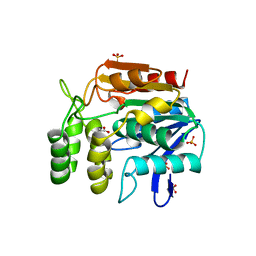 | | CHLOROPEROXIDASE F/PROPIONATE COMPLEX | | Descriptor: | CHLOROPEROXIDASE F, PROPANOIC ACID, SULFATE ION | | Authors: | Hofmann, B, Toelzer, S, Pelletier, I, Altenbuchner, J, Van Pee, K.-H, Hecht, H.-J. | | Deposit date: | 1998-03-27 | | Release date: | 1998-06-17 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural investigation of the cofactor-free chloroperoxidases.
J.Mol.Biol., 279, 1998
|
|
3NGV
 
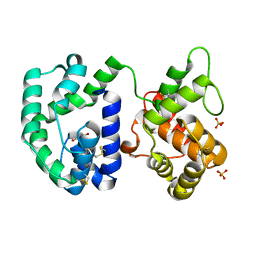 | | Crystal structure of AnSt-D7L1 | | Descriptor: | D7 protein, GLYCEROL, SULFATE ION | | Authors: | Andersen, J.F, Alvarenga, P.H, Francischetti, I.M, Ribeiro, J.M, Calvo, E. | | Deposit date: | 2010-06-13 | | Release date: | 2010-12-29 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | The function and three-dimensional structure of a thromboxane a(2)/cysteinyl leukotriene-binding protein from the saliva of a mosquito vector of the malaria parasite.
Plos Biol., 8, 2010
|
|
3NH8
 | | Crystal structure of murine aminoacylase 3 in complex with N-acetyl-S-1,2-dichlorovinyl-L-cysteine | | Descriptor: | Aspartoacylase-2, CHLORIDE ION, N-acetyl-S-[(1S)-1,2-dichloroethyl]-L-cysteine, ... | | Authors: | Hsieh, J.M, Tsirulnikov, K, Sawaya, M.R, Magilnick, N, Abuladze, N, Kurtz, I, Abramson, J, Pushkin, A. | | Deposit date: | 2010-06-14 | | Release date: | 2010-10-20 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.802 Å) | | Cite: | Structures of aminoacylase 3 in complex with acetylated substrates.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
