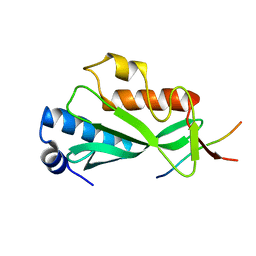
3OBQ
 
 | |
3OBX
 | |
3MHV
 
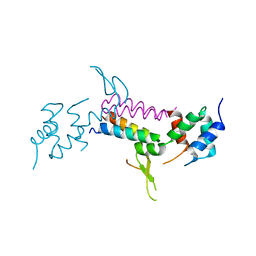 | | Crystal Structure of Vps4 and Vta1 | | Descriptor: | Vacuolar protein sorting-associated protein 4, Vacuolar protein sorting-associated protein VTA1 | | Authors: | Yang, D, Hurley, J.H. | | Deposit date: | 2010-04-09 | | Release date: | 2010-10-06 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural role of the Vps4-Vta1 interface in ESCRT-III recycling
Structure, 18, 2010
|
|
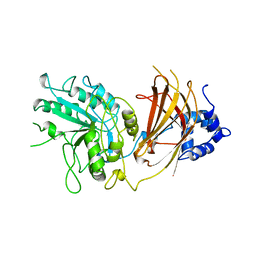
1QAS
 
 | |
3OBS
 
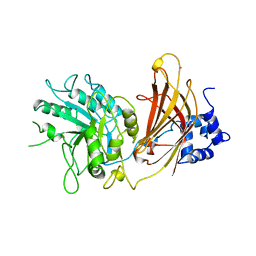
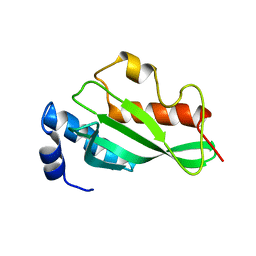 | | Crystal structure of Tsg101 UEV domain | | Descriptor: | Tumor susceptibility gene 101 protein | | Authors: | Im, Y.J, Hurley, J.H. | | Deposit date: | 2010-08-09 | | Release date: | 2010-12-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystallographic and Functional Analysis of the ESCRT-I /HIV-1 Gag PTAP Interaction.
Structure, 18, 2010
|
|
1QAT
 
 | |
3PFQ
 
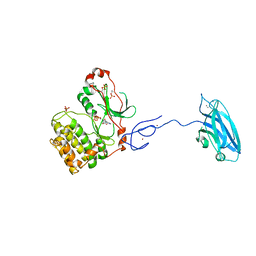 | | Crystal Structure and Allosteric Activation of Protein Kinase C beta II | | Descriptor: | CALCIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, Protein kinase C beta type, ... | | Authors: | Leonard, T.A, Rozycki, B, Saidi, L.F, Hummer, G, Hurley, J.H. | | Deposit date: | 2010-10-28 | | Release date: | 2011-02-02 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (4 Å) | | Cite: | Crystal Structure and Allosteric Activation of Protein Kinase C beta II
Cell(Cambridge,Mass.), 144, 2011
|
|
2P22
 | | Structure of the Yeast ESCRT-I Heterotetramer Core | | Descriptor: | Hypothetical 12.0 kDa protein in ADE3-SER2 intergenic region, Protein SRN2, SULFATE ION, ... | | Authors: | Kostelansky, M.S, Hurley, J.H. | | Deposit date: | 2007-03-06 | | Release date: | 2007-06-05 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Molecular architecture and functional model of the complete yeast ESCRT-I heterotetramer.
Cell(Cambridge,Mass.), 129, 2007
|
|
2OJQ
 
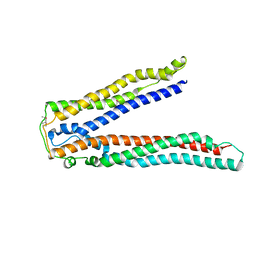 | | Crystal structure of Alix V domain | | Descriptor: | Programmed cell death 6-interacting protein | | Authors: | Lee, S, Hurley, J.H. | | Deposit date: | 2007-01-13 | | Release date: | 2007-02-20 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.87 Å) | | Cite: | Structural basis for viral late-domain binding to Alix
Nat.Struct.Mol.Biol., 14, 2007
|
|
7MU2
 
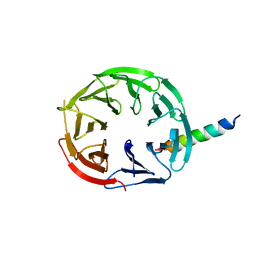 | | Crystal Structure of WIPI2 in complex with W2IR of ATG16L1 | | Descriptor: | Autophagy-related protein 16-1, WD repeat domain phosphoinositide-interacting protein 2 | | Authors: | Strong, L.M, Flower, T.G, Buffalo, C.Z, Hurley, J.H. | | Deposit date: | 2021-05-14 | | Release date: | 2021-05-26 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural basis for membrane recruitment of ATG16L1 by WIPI2 in autophagy.
Elife, 10, 2021
|
|
7MGE
 
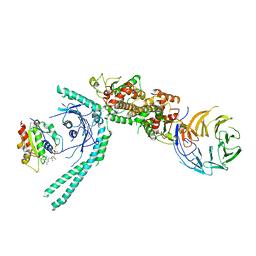 | | Structure of C9orf72:SMCR8:WDR41 in complex with ARF1 | | Descriptor: | ADP-ribosylation factor 1, BERYLLIUM TRIFLUORIDE ION, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Su, M.-Y, Hurley, J.H. | | Deposit date: | 2021-04-12 | | Release date: | 2021-06-23 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (3.94 Å) | | Cite: | Structural basis for the ARF GAP activity and specificity of the C9orf72 complex.
Nat Commun, 12, 2021
|
|
1P4U
 
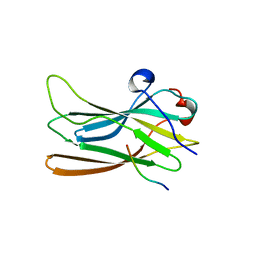 | | CRYSTAL STRUCTURE OF GGA3 GAE DOMAIN IN COMPLEX WITH RABAPTIN-5 PEPTIDE | | Descriptor: | ADP-ribosylation factor binding protein GGA3, Rabaptin-5 | | Authors: | Miller, G.J, Mattera, R, Bonifacino, J.S, Hurley, J.H. | | Deposit date: | 2003-04-24 | | Release date: | 2003-07-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | RECOGNITION OF ACCESSORY PROTEIN MOTIFS BY THE GAMMA-ADAPTIN EAR DOMAIN OF GGA3
Nat.Struct.Biol., 10, 2003
|
|
1OZV
 
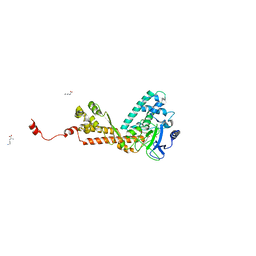 | | Crystal structure of the SET domain of LSMT bound to Lysine and AdoHcy | | Descriptor: | LYSINE, Ribulose-1,5 bisphosphate carboxylase/oxygenase large subunit N-methyltransferase, chloroplast, ... | | Authors: | Trievel, R.C, Flynn, E.M, Houtz, R.L, Hurley, J.H. | | Deposit date: | 2003-04-09 | | Release date: | 2003-07-01 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Mechanism of multiple lysine methylation by the SET domain enzyme Rubisco LSMT
Nat.Struct.Biol., 10, 2003
|
|
1MLV
 
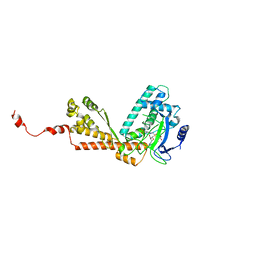 | | Structure and Catalytic Mechanism of a SET Domain Protein Methyltransferase | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Ribulose-1,5 biphosphate carboxylase/oxygenase large subunit N-methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Trievel, R.C, Beach, B.M, Dirk, L.M.A, Houtz, R.L, Hurley, J.H. | | Deposit date: | 2002-08-30 | | Release date: | 2002-10-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure and catalytic mechanism of a SET domain protein methyltransferase.
Cell(Cambridge,Mass.), 111, 2002
|
|
1NWM
 
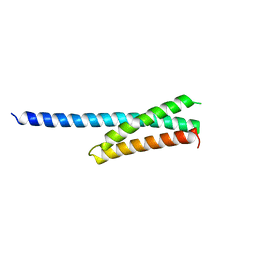 | | GAT domain of human GGA1 | | Descriptor: | ADP-ribosylation factor binding protein GGA1 | | Authors: | Suer, S, Misra, S, Saidi, L.F, Hurley, J.H. | | Deposit date: | 2003-02-06 | | Release date: | 2003-03-25 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of the GAT domain of human GGA1: a syntaxin amino-terminal domain fold in an endosomal trafficking adaptor.
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1PTQ
 
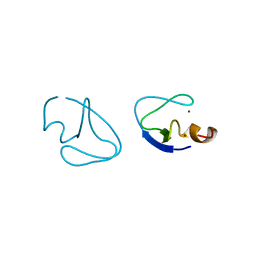 | | PROTEIN KINASE C DELTA CYS2 DOMAIN | | Descriptor: | PROTEIN KINASE C DELTA TYPE, ZINC ION | | Authors: | Zhang, G, Hurley, J.H. | | Deposit date: | 1995-05-11 | | Release date: | 1995-07-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of the cys2 activator-binding domain of protein kinase C delta in complex with phorbol ester.
Cell(Cambridge,Mass.), 81, 1995
|
|
1PTR
 
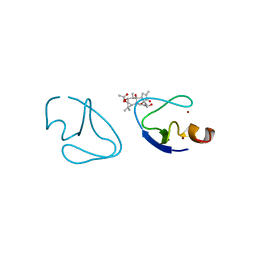 | |
1P0Y
 
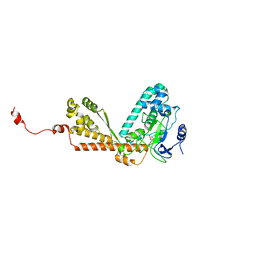 | | Crystal structure of the SET domain of LSMT bound to MeLysine and AdoHcy | | Descriptor: | N-METHYL-LYSINE, Ribulose-1,5 bisphosphate carboxylase/oxygenase large subunit N-methyltransferase, chloroplast, ... | | Authors: | Trievel, R.C, Flynn, E.M, Houtz, R.L, Hurley, J.H. | | Deposit date: | 2003-04-11 | | Release date: | 2003-07-01 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Mechanism of multiple lysine methylation by the SET domain enzyme Rubisco LSMT
Nat.Struct.Biol., 10, 2003
|
|
1JPL
 
 | | GGA3 VHS domain complexed with C-terminal peptide from cation-independent mannose 6-phosphate receptor | | Descriptor: | ADP-RIBOSYLATION FACTOR BINDING PROTEIN GGA3, Cation-Independent Mannose 6-phosphate receptor | | Authors: | Misra, S, Puertollano, R, Bonifacino, J.S, Hurley, J.H. | | Deposit date: | 2001-08-02 | | Release date: | 2002-02-27 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for acidic-cluster-dileucine sorting-signal recognition by VHS domains.
Nature, 415, 2002
|
|
1XA6
 
 | | Crystal Structure of the Human Beta2-Chimaerin | | Descriptor: | Beta2-chimaerin, ZINC ION | | Authors: | Canagarajah, B, Leskow, F.C, Ho, J.Y, Mischak, H, Saidi, L.F, Kazanietz, M.G, Hurley, J.H. | | Deposit date: | 2004-08-25 | | Release date: | 2004-11-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural mechanism for lipid activation of the Rac-specific GAP, beta2-chimaerin.
Cell(Cambridge,Mass.), 119, 2004
|
|
1I9Z
 
 | | CRYSTAL STRUCTURE OF INOSITOL POLYPHOSPHATE 5-PHOSPHATASE DOMAIN (IPP5C) OF SPSYNAPTOJANIN IN COMPLEX WITH INOSITOL (1,4)-BISPHOSPHATE AND CALCIUM ION | | Descriptor: | CALCIUM ION, D-MYO-INOSITOL-1,4-BISPHOSPHATE, PHOSPHATIDYLINOSITOL PHOSPHATE PHOSPHATASE | | Authors: | Tsujishita, Y, Guo, S, Stolz, L, York, J.D, Hurley, J.H. | | Deposit date: | 2001-03-21 | | Release date: | 2001-05-16 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Specificity determinants in phosphoinositide dephosphorylation: crystal structure of an archetypal inositol polyphosphate 5-phosphatase.
Cell(Cambridge,Mass.), 105, 2001
|
|
1ZHW
 
 | | Structure of yeast oxysterol binding protein Osh4 in complex with 20-hydroxycholesterol | | Descriptor: | 20-HYDROXYCHOLESTEROL, KES1 protein, LEAD (II) ION | | Authors: | Im, Y.J, Raychaudhuri, S, Prinz, W.A, Hurley, J.H. | | Deposit date: | 2005-04-26 | | Release date: | 2005-09-06 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural mechanism for sterol sensing and transport by OSBP-related proteins
Nature, 437, 2005
|
|
1ZI7
 
 | | Structure of truncated yeast oxysterol binding protein Osh4 | | Descriptor: | KES1 protein, SULFATE ION | | Authors: | Im, Y.J, Raychaudhuri, S, Prinz, W.A, Hurley, J.H. | | Deposit date: | 2005-04-27 | | Release date: | 2005-09-06 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural mechanism for sterol sensing and transport by OSBP-related proteins
Nature, 437, 2005
|
|
1ZHX
 
 | | Structure of yeast oxysterol binding protein Osh4 in complex with 25-hydroxycholesterol | | Descriptor: | 25-HYDROXYCHOLESTEROL, KES1 protein | | Authors: | Im, Y.J, Raychaudhuri, S, Prinz, W.A, Hurley, J.H. | | Deposit date: | 2005-04-26 | | Release date: | 2005-09-06 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural mechanism for sterol sensing and transport by OSBP-related proteins
Nature, 437, 2005
|
|
1I9Y
 | | CRYSTAL STRUCTURE OF INOSITOL POLYPHOSPHATE 5-PHOSPHATASE DOMAIN (IPP5C) OF SPSYNAPTOJANIN | | Descriptor: | PHOSPHATIDYLINOSITOL PHOSPHATE PHOSPHATASE | | Authors: | Tsujishita, Y, Guo, S, Stolz, L, York, J.D, Hurley, J.H. | | Deposit date: | 2001-03-21 | | Release date: | 2001-05-16 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Specificity determinants in phosphoinositide dephosphorylation: crystal structure of an archetypal inositol polyphosphate 5-phosphatase.
Cell(Cambridge,Mass.), 105, 2001
|
|
