3TEA
 
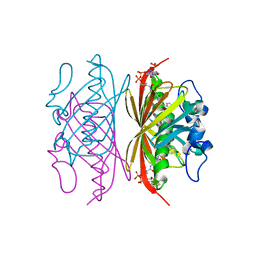 | | Crystal structure of Arthrobacter sp. strain su 4-hydroxybenzoyl CoA thioesterase double mutant Q58E/E73A complexed with 4-hydroxyphenacyl CoA | | Descriptor: | 4-HYDROXYPHENACYL COENZYME A, 4-hydroxybenzoyl-CoA thioesterase | | Authors: | Holden, H.M, Thoden, J.B, Song, F, Zhuang, Z, Trujillo, M, Dunaway-Mariano, D. | | Deposit date: | 2011-08-12 | | Release date: | 2012-08-15 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Catalytic Mechanism of the Hotdog-fold Enzyme Superfamily 4-Hydroxybenzoyl-CoA Thioesterase from Arthrobacter sp. Strain SU.
Biochemistry, 51, 2012
|
|
3GR9
 
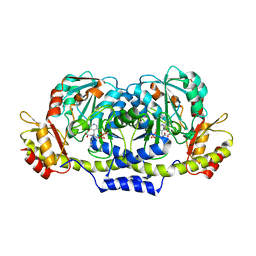 | | Crystal structure of ColD H188K S187N | | Descriptor: | 2-OXOGLUTARIC ACID, ColD | | Authors: | Holden, H.M, Cook, P.D, Kubiak, R.L, Toomey, D.P. | | Deposit date: | 2009-03-25 | | Release date: | 2009-06-16 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Two Site-Directed Mutations Are Required for the Conversion of a Sugar Dehydratase into an Aminotransferase.
Biochemistry, 48, 2009
|
|
4HMZ
 
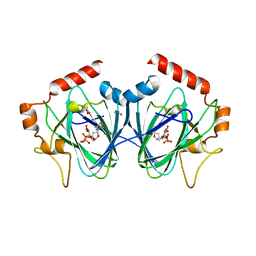 | | Crystal Structure of ChmJ, a 3'-monoepimerase from Streptomyces bikiniensis in complex with dTDP-quinovose | | Descriptor: | 1,2-ETHANEDIOL, Putative 3-epimerase in D-allose pathway, [(2R,3S,5R)-3-hydroxy-5-(5-methyl-2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)tetrahydrofuran-2-yl]methyl (2R,3R,4S,5S,6R)-3,4,5-trihydroxy-6-methyltetrahydro-2H-pyran-2-yl dihydrogen diphosphate | | Authors: | Holden, H.M, Kubiak, R.L. | | Deposit date: | 2012-10-18 | | Release date: | 2012-11-21 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Functional Studies on a 3'-Epimerase Involved in the Biosynthesis of dTDP-6-deoxy-d-allose.
Biochemistry, 51, 2012
|
|
4HN0
 
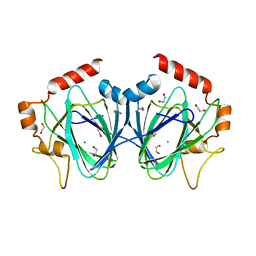 | |
4HN1
 
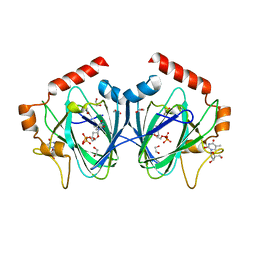 | | Crystal Structure of H60N/Y130F double mutant of ChmJ, a 3'-monoepimerase from Streptomyces bikiniensis in complex with dTDP | | Descriptor: | 1,2-ETHANEDIOL, Putative 3-epimerase in D-allose pathway, THYMIDINE, ... | | Authors: | Holden, H.M, Kubiak, R.L. | | Deposit date: | 2012-10-18 | | Release date: | 2012-11-21 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural and Functional Studies on a 3'-Epimerase Involved in the Biosynthesis of dTDP-6-deoxy-d-allose.
Biochemistry, 51, 2012
|
|
1Z24
 
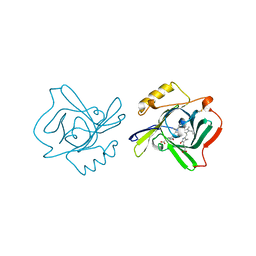 | | The molecular structure of insecticyanin from the tobacco hornworm Manduca sexta L. at 2.6 A resolution. | | Descriptor: | BILIVERDIN IX GAMMA CHROMOPHORE, Insecticyanin A form | | Authors: | Holden, H.M, Rypniewski, W.R, Law, J.H, Rayment, I. | | Deposit date: | 2005-03-07 | | Release date: | 2005-04-05 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The molecular structure of insecticyanin from the tobacco hornworm Manduca sexta L. at 2.6 A resolution.
Embo J., 6, 1987
|
|
3TMN
 
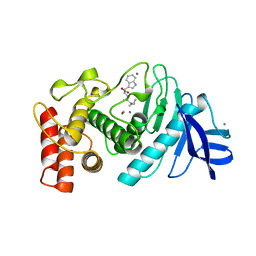 | |
4KCF
 
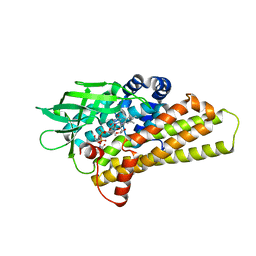 | | X-ray Structure of a KijD3 in Complex with FMN and dTDP-3-amino-2,3,6-trideoxy-4-keto-3-methyl-D-glucose | | Descriptor: | FAD-dependent oxidoreductase, FLAVIN MONONUCLEOTIDE, [(2R,4S,6R)-4-azanyl-4,6-dimethyl-5,5-bis(oxidanyl)oxan-2-yl] [[(2R,3S,5R)-5-[5-methyl-2,4-bis(oxidanylidene)pyrimidin-1-yl]-3-oxidanyl-oxolan-2-yl]methoxy-oxidanyl-phosphoryl] hydrogen phosphate | | Authors: | Holden, H.M, Thoden, J.B, Branch, M.C, Zimmer, A.L, Bruender, N.A. | | Deposit date: | 2013-04-24 | | Release date: | 2013-05-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.098 Å) | | Cite: | Active site architecture of a sugar N-oxygenase.
Biochemistry, 52, 2013
|
|
3FSB
 
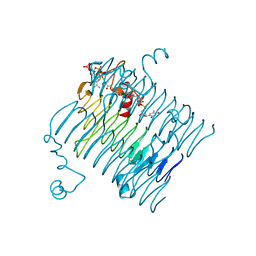 | | Crystal structure of QdtC, the dTDP-3-amino-3,6-dideoxy-D-glucose N-acetyl transferase from Thermoanaerobacterium thermosaccharolyticum in complex with CoA and dTDP-3-amino-quinovose | | Descriptor: | COENZYME A, QdtC, [(3R,4S,5S,6R)-4-amino-3,5-dihydroxy-6-methyloxan-2-yl][hydroxy-[[(2R,3S,5R)-3-hydroxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy]phosphoryl] hydrogen phosphate | | Authors: | Holden, H.M, Thoden, J.B. | | Deposit date: | 2009-01-09 | | Release date: | 2009-02-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural and Functional Studies of QdtC: an N-Acetyltransferase Required for the Biosynthesis of dTDP-3-Acetamido-3,6-Dideoxy-alpha-D-Glucose.
Biochemistry, 48, 2009
|
|
3FSC
 
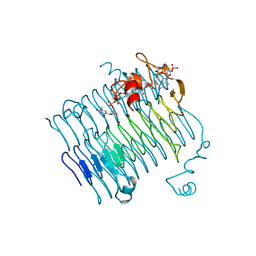 | | Crystal structure of QdtC, the dTDP-3-amino-3,6-dideoxy-D-glucose N-acetyl transferase from Thermoanaerobacterium thermosaccharolyticum in complex with CoA and dTDP-3-amino-fucose | | Descriptor: | (3R,4S,5R,6R)-4-amino-3,5-dihydroxy-6-methyloxan-2-yl][hydroxy-[[(2R,3S,5R)-3-hydroxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy]phosphoryl] hydrogen phosphate, COENZYME A, QdtC | | Authors: | Holden, H.M, Thoden, J.B. | | Deposit date: | 2009-01-09 | | Release date: | 2009-02-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and Functional Studies of QdtC: an N-Acetyltransferase Required for the Biosynthesis of dTDP-3-Acetamido-3,6-Dideoxy-alpha-D-Glucose.
Biochemistry, 48, 2009
|
|
3FS8
 
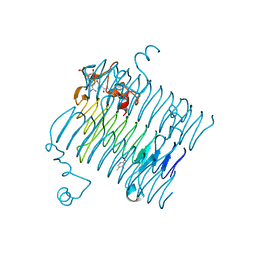 | | Crystal structure of QdtC, the dTDP-3-amino-3,6-dideoxy-D-glucose N-acetyl transferase from Thermoanaerobacterium thermosaccharolyticum in complex with Acetyl-CoA | | Descriptor: | ACETYL COENZYME *A, QdtC, THYMINE | | Authors: | Holden, H.M, Thoden, J.B. | | Deposit date: | 2009-01-09 | | Release date: | 2009-02-17 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural and Functional Studies of QdtC: an N-Acetyltransferase Required for the Biosynthesis of dTDP-3-Acetamido-3,6-Dideoxy-alpha-D-Glucose.
Biochemistry, 48, 2009
|
|
4J7H
 
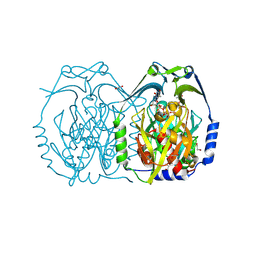 | | Crystal structure of EvaA, a 2,3-dehydratase in complex with dTDP-benzene and dTDP-rhamnose | | Descriptor: | 1,2-ETHANEDIOL, 2'-DEOXY-THYMIDINE-BETA-L-RHAMNOSE, 5'-O-[(S)-hydroxy{[(S)-hydroxy(phenoxy)phosphoryl]oxy}phosphoryl]thymidine, ... | | Authors: | Holden, H.M, Kubiak, R.L, Thoden, J.B. | | Deposit date: | 2013-02-13 | | Release date: | 2013-05-22 | | Last modified: | 2013-12-25 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Structure of EvaA: A Paradigm for Sugar 2,3-Dehydratases.
Biochemistry, 52, 2013
|
|
2OGA
 
 | |
4J7G
 
 | | Crystal structure of EvaA, a 2,3-dehydratase in complex with dTDP-fucose and dTDP-rhamnose | | Descriptor: | 2'-DEOXY-THYMIDINE-BETA-L-RHAMNOSE, EvaA 2,3-dehydratase, [[(2R,3S,5R)-5-[5-methyl-2,4-bis(oxidanylidene)pyrimidin-1-yl]-3-oxidanyl-oxolan-2-yl]methoxy-oxidanyl-phosphoryl] [(2R,3R,4S,5R,6R)-6-methyl-3,4,5-tris(oxidanyl)oxan-2-yl] hydrogen phosphate | | Authors: | Holden, H.M, Kubiak, R.L, Thoden, J.B. | | Deposit date: | 2013-02-13 | | Release date: | 2013-05-22 | | Last modified: | 2013-12-25 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure of EvaA: A Paradigm for Sugar 2,3-Dehydratases.
Biochemistry, 52, 2013
|
|
2OGE
 
 | |
2PA4
 
 | |
3NYT
 
 | | X-ray crystal structure of the WlbE (WpbE) aminotransferase from pseudomonas aeruginosa, mutation K185A, in complex with the PLP external aldimine adduct with UDP-3-amino-2-N-acetyl-glucuronic acid, at 1.3 angstrom resolution | | Descriptor: | (2S,3S,4R,5R,6R)-5-(acetylamino)-6-{[(R)-{[(S)-{[(2R,3S,4R,5R)-5-(2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)-3,4-dihydroxytetrahydrofuran-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}-3-hydroxy-4-{[(1E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]amino}tetrahydro-2H-pyran-2-carboxylic acid (non-preferred name), Aminotransferase WbpE, SODIUM ION | | Authors: | Holden, H.M, Thoden, J.B. | | Deposit date: | 2010-07-15 | | Release date: | 2010-07-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.301 Å) | | Cite: | Structural investigation on WlaRG from Campylobacter jejuni: A sugar aminotransferase.
Protein Sci., 26, 2017
|
|
3NYU
 
 | |
3OA2
 
 | |
3NYS
 
 | |
3OA0
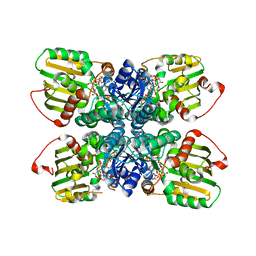 
 | | Crystal structure of the WlbA (WbpB) Dehydrogenase from Thermus thermophilus in complex with NAD and UDP-GlcNAcA | | Descriptor: | (2S,3S,4R,5R,6R)-5-acetamido-6-[[[(2R,3S,4R,5R)-5-(2,4-dioxopyrimidin-1-yl)-3,4-dihydroxy-oxolan-2-yl]methoxy-hydroxy-phosphoryl]oxy-hydroxy-phosphoryl]oxy-3,4-dihydroxy-oxane-2-carboxylic acid, CHLORIDE ION, Lipopolysaccharide biosynthesis protein wbpB, ... | | Authors: | Holden, H.M, Thoden, J.B. | | Deposit date: | 2010-08-04 | | Release date: | 2010-08-18 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Functional Studies of WlbA: A Dehydrogenase Involved in the Biosynthesis of 2,3-Diacetamido-2,3-dideoxy-d-mannuronic Acid .
Biochemistry, 49, 2010
|
|
3O9Z
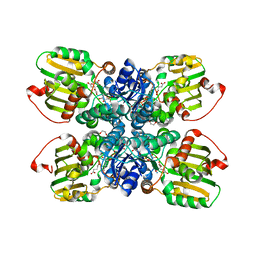 
 | | Crystal structure of the WlbA (WbpB) dehydrogenase from Thermus thermophilus in complex with NAD and alpha-ketoglutarate at 1.45 angstrom resolution | | Descriptor: | 1,2-ETHANEDIOL, 2-OXOGLUTARIC ACID, CHLORIDE ION, ... | | Authors: | Holden, H.M, Thoden, J.B. | | Deposit date: | 2010-08-04 | | Release date: | 2010-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.449 Å) | | Cite: | Structural and Functional Studies of WlbA: A Dehydrogenase Involved in the Biosynthesis of 2,3-Diacetamido-2,3-dideoxy-d-mannuronic Acid .
Biochemistry, 49, 2010
|
|
1BVQ
 
 | |
5K8C
 
 | | X-ray structure of KdnB, 3-deoxy-alpha-D-manno-octulosonate 8-oxidase, from Shewanella oneidensis | | Descriptor: | 1,2-ETHANEDIOL, 3-deoxy-alpha-D-manno-octulosonate 8-oxidase, CHLORIDE ION, ... | | Authors: | Holden, H.M, Thoden, J.B, Zachman-Brockmeyer, T.R. | | Deposit date: | 2016-05-28 | | Release date: | 2016-06-15 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structures of KdnB and KdnA from Shewanella oneidensis: Key Enzymes in the Formation of 8-Amino-3,8-Dideoxy-d-Manno-Octulosonic Acid.
Biochemistry, 55, 2016
|
|
5K8B
 
 | | X-ray structure of KdnA, 8-amino-3,8-dideoxy-alpha-D-manno-octulosonate transaminase, from Shewanella oneidensis in the presence of the external aldimine with PLP and glutamate | | Descriptor: | 8-amino-3,8-dideoxy-alpha-D-manno-octulosonate transaminase, CHLORIDE ION, N-({3-HYDROXY-2-METHYL-5-[(PHOSPHONOOXY)METHYL]PYRIDIN-4-YL}METHYL)-D-GLUTAMIC ACID, ... | | Authors: | Holden, H.M, Thoden, J.B, Zachman-Brockmeyer, T.R. | | Deposit date: | 2016-05-28 | | Release date: | 2016-06-15 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structures of KdnB and KdnA from Shewanella oneidensis: Key Enzymes in the Formation of 8-Amino-3,8-Dideoxy-d-Manno-Octulosonic Acid.
Biochemistry, 55, 2016
|
|
