4GKL
 
 | |
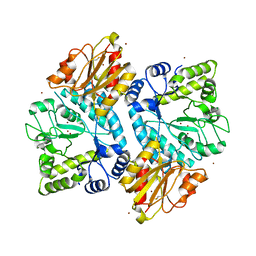
5C0Q
 
 | |
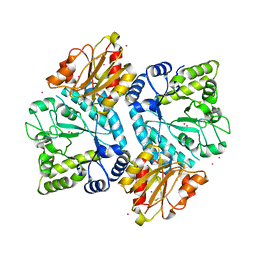
5BZA
 
 | |
4GQZ
 
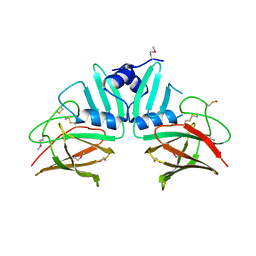 | | Crystal Structure of S.CueP | | Descriptor: | CHLORIDE ION, Putative periplasmic or exported protein | | Authors: | Ha, N.C, Yoon, B.Y. | | Deposit date: | 2012-08-24 | | Release date: | 2013-08-14 | | Last modified: | 2013-10-23 | | Method: | X-RAY DIFFRACTION (1.799 Å) | | Cite: | Structure of the periplasmic copper-binding protein CueP from Salmonella enterica serovar Typhimurium
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4QA8
 
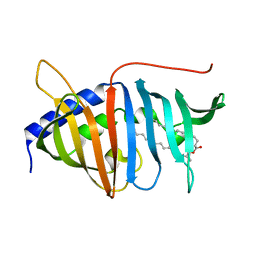 | | Crystal structure of LprF from Mycobacterium bovis | | Descriptor: | (2R)-2-(dodecanoyloxy)propyl (4E,6E,8E,10E,12E)-pentadeca-4,6,8,10,12-pentaenoate, Putative lipoprotein LprF | | Authors: | Ha, N.C, Jiao, L, Kim, J.S. | | Deposit date: | 2014-05-02 | | Release date: | 2014-10-22 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Crystal structure and functional implications of LprF from Mycobacterium tuberculosis and M. bovis
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4I5Q
 
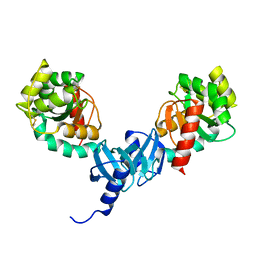 | | Crystal structure and catalytic mechanism for peroplasmic disulfide-bond isomerase DsbC from Salmonella enterica serovar Typhimurium | | Descriptor: | MAGNESIUM ION, Thiol:disulfide interchange protein DsbC | | Authors: | Ha, N.C, Li, J, Kim, J.S, Yoon, B.Y, Yeom, J.H, Lee, K. | | Deposit date: | 2012-11-28 | | Release date: | 2013-10-16 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (1.962 Å) | | Cite: | Crystal structure of the periplasmic disulfide-bond isomerase DsbC from Salmonella enterica serovar Typhimurium and the mechanistic implications.
J.Struct.Biol., 183, 2013
|
|
4ILF
 
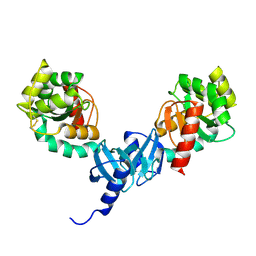 | | Crystal structure of DsbC R125A from Salmonella enterica serovar Typhimurium | | Descriptor: | Thiol:disulfide interchange protein DsbC | | Authors: | Ha, N.C, Li, J, Kim, J.S, Yoon, B.Y, Yeom, J.H, Lee, K. | | Deposit date: | 2012-12-31 | | Release date: | 2013-10-16 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (1.999 Å) | | Cite: | Crystal structure of the periplasmic disulfide-bond isomerase DsbC from Salmonella enterica serovar Typhimurium and the mechanistic implications.
J.Struct.Biol., 183, 2013
|
|
4ML7
 
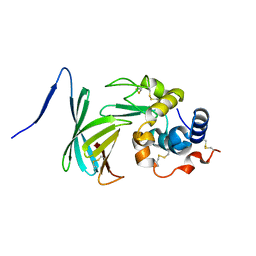 | |
4MIR
 
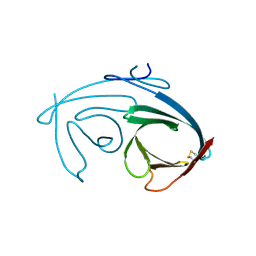 | |
4YK9
 
 | |
2OLG
 
 | |
4Z85
 
 | |
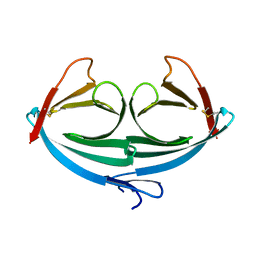
4MIS
 | |
5ZQS
 
 | |
5ZQJ
 
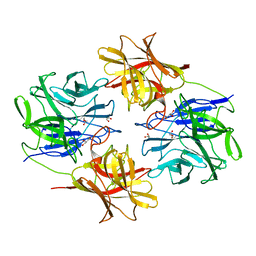 | | Crystal structure of beta-xylosidase from Bacillus pumilus | | Descriptor: | Beta-xylosidase, GLYCEROL | | Authors: | Ha, N.C, Hong, S, Jo, I. | | Deposit date: | 2018-04-19 | | Release date: | 2018-05-30 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Structure-based protein engineering of bacterial beta-xylosidase to increase the production yield of xylobiose from xylose
Biochem. Biophys. Res. Commun., 501, 2018
|
|
5ZQX
 
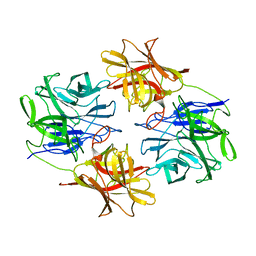 | |
5C22
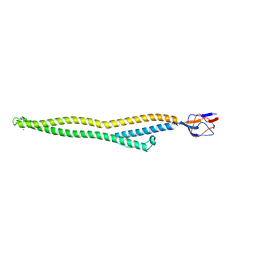 
 | | Crystal structure of Zn-bound HlyD from E. coli | | Descriptor: | Chromosomal hemolysin D, ZINC ION | | Authors: | Ha, N.C, Kim, J.S. | | Deposit date: | 2015-06-15 | | Release date: | 2016-02-17 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.302 Å) | | Cite: | Crystal Structure of a Soluble Fragment of the Membrane Fusion Protein HlyD in a Type I Secretion System of Gram-Negative Bacteria
Structure, 24, 2016
|
|
5C21
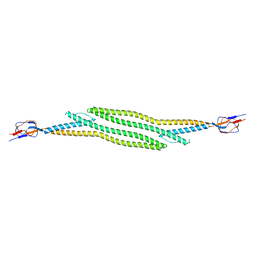 
 | | Crystal structure of native HlyD from E. coli | | Descriptor: | Chromosomal hemolysin D | | Authors: | Ha, N.C, Kim, J.S, Yoon, B.Y. | | Deposit date: | 2015-06-15 | | Release date: | 2016-02-17 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure of a Soluble Fragment of the Membrane Fusion Protein HlyD in a Type I Secretion System of Gram-Negative Bacteria
Structure, 24, 2016
|
|
5C59
 
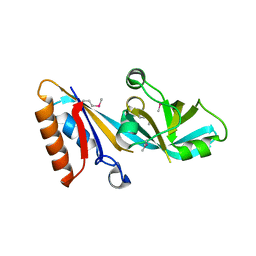 | |
3F6Z
 
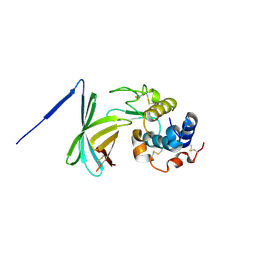 | |
3U95
 
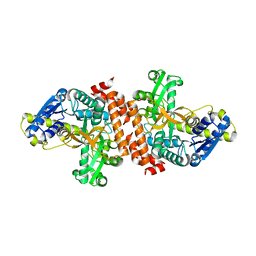 | | Crystal structure of a putative alpha-glucosidase from Thermotoga neapolitana | | Descriptor: | Glycoside hydrolase, family 4, MANGANESE (II) ION | | Authors: | Ha, N.C, Jun, S.Y, Yun, B.Y, Yoon, B.Y, Piao, S. | | Deposit date: | 2011-10-17 | | Release date: | 2012-09-26 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.998 Å) | | Cite: | Crystal structure and thermostability of a putative alpha-glucosidase from Thermotoga neapolitana
Biochem.Biophys.Res.Commun., 416, 2011
|
|
4PQ1
 
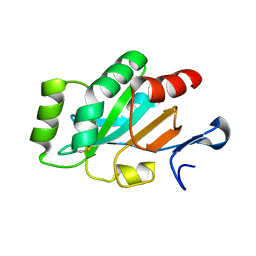 | | Crystal structure and functional implications of a DsbF homologue from Corynebacterium diphtheriae | | Descriptor: | Putative electron transport related protein | | Authors: | Um, S.H, Kim, J.S, Yoon, B.Y, Ha, N.C. | | Deposit date: | 2014-02-28 | | Release date: | 2014-09-10 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (2.097 Å) | | Cite: | Structure of a DsbF homologue from Corynebacterium diphtheriae.
Acta Crystallogr.,Sect.F, 70, 2014
|
|
7YVD
 
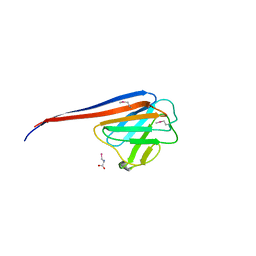 | |
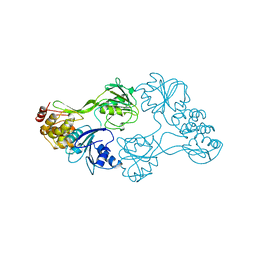
8OHM
 
 | |
4X6G
 
 | |
