1B2A
 
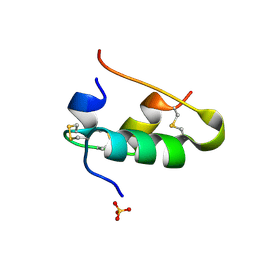 | |
1CO4
 | | SOLUTION STRUCTURE OF A ZINC DOMAIN CONSERVED IN YEAST COPPER-REGULATED TRANSCRIPTION FACTORS | | Descriptor: | PROTEIN (ACTIVATOR OF METALLOTHIONEIN 1), ZINC ION | | Authors: | Turner, R.B, Smith, D.L, Zawrotny, M.E, Summers, M.F, Posewitz, M.C, Winge, D.R. | | Deposit date: | 1999-06-04 | | Release date: | 1999-06-10 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of a zinc domain conserved in yeast copper-regulated transcription factors.
Nat.Struct.Biol., 5, 1998
|
|
1CPH
 | | CONFORMATIONAL CHANGES IN CUBIC INSULIN CRYSTALS IN THE PH RANGE 7-11 | | Descriptor: | 1,2-DICHLOROETHANE, INSULIN (PH 10), SODIUM ION | | Authors: | Gursky, O, Badger, J, Li, Y, Caspar, D.L.D. | | Deposit date: | 1992-10-30 | | Release date: | 1993-01-15 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Conformational changes in cubic insulin crystals in the pH range 7-11.
Biophys.J., 63, 1992
|
|
2LFV
 
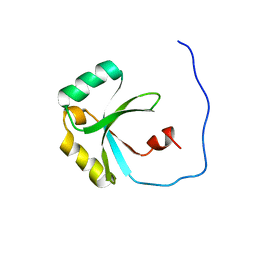 | | Solution Structure of the SPOR domain from E. coli DamX | | Descriptor: | Protein damX | | Authors: | Williams, K.B, Arends, S.J.R, Popham, D.L, Fowler, C.A, Weiss, D.S. | | Deposit date: | 2011-07-15 | | Release date: | 2012-07-18 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Nuclear Magnetic Resonance Solution Structure of the Peptidoglycan-Binding SPOR Domain from Escherichia coli DamX: Insights into Septal Localization.
Biochemistry, 52, 2013
|
|
2LNT
 
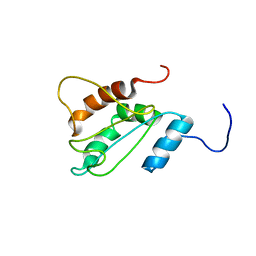 | | Solution structure of E60A mutant AGR2 | | Descriptor: | Anterior gradient protein 2 homolog | | Authors: | Patel, P, Clarke, C.J, Barraclough, D.L, Rudland, P.S, Barraclough, R, Lian, L. | | Deposit date: | 2012-01-04 | | Release date: | 2013-01-09 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Metastasis-promoting anterior gradient 2 protein has a dimeric thioredoxin fold structure and a role in cell adhesion.
J.Mol.Biol., 425, 2013
|
|
2LNS
 
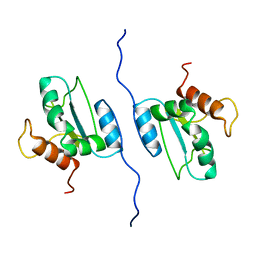 | | Solution structure of AGR2 residues 41-175 | | Descriptor: | Anterior gradient protein 2 homolog | | Authors: | Patel, P, Clarke, C.J, Barraclough, D.L, Rudland, P.S, Barraclough, R, Lian, L. | | Deposit date: | 2012-01-04 | | Release date: | 2013-01-09 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Metastasis-promoting anterior gradient 2 protein has a dimeric thioredoxin fold structure and a role in cell adhesion.
J.Mol.Biol., 425, 2013
|
|
2M5S
 
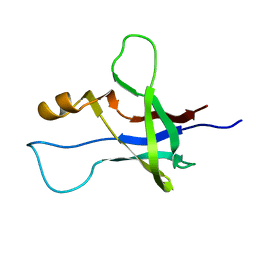 | | High-resolution NMR structure and cryo-EM imaging support multiple functional roles for the accessory I-domain of phage P22 coat protein | | Descriptor: | Coat protein | | Authors: | Rizzo, A.A, Suhanovsky, M.M, Baker, M.L, Fraser, L.C.R, Jones, L.M, Rempel, D.L, Gross, M.L, Chiu, W, Alexandrescu, A.T, Teschke, C.M. | | Deposit date: | 2013-03-05 | | Release date: | 2014-03-05 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Multiple Functional Roles of the Accessory I-Domain of Bacteriophage P22 Coat Protein Revealed by NMR Structure and CryoEM Modeling.
Structure, 22, 2014
|
|
2MGX
 
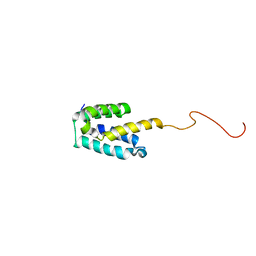 | | NMR structure of SRA1p C-terminal domain | | Descriptor: | Steroid receptor RNA activator 1 | | Authors: | Bilinovich, S.M, Davis, C.M, Morris, D.L, Ray, L.A, Prokop, J.W, Buchan, G.J, Leeper, T.C. | | Deposit date: | 2013-11-10 | | Release date: | 2014-02-12 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The C-Terminal Domain of SRA1p Has a Fold More Similar to PRP18 than to an RRM and Does Not Directly Bind to the SRA1 RNA STR7 Region.
J.Mol.Biol., 426, 2014
|
|
2MZ9
 
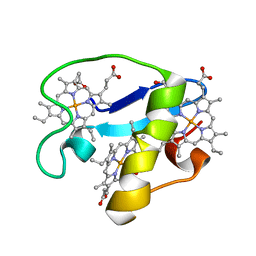 | | Solution structure of oxidized triheme cytochrome PpcA from Geobacter sulfurreducens | | Descriptor: | HEME C, PpcA | | Authors: | Morgado, L, Bruix, M, Pokkuluri, R, Salgueiro, C.A, Turner, D.L. | | Deposit date: | 2015-02-08 | | Release date: | 2016-02-10 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Redox- and pH-linked conformational changes in triheme cytochrome PpcA from Geobacter sulfurreducens.
Biochem. J., 474, 2017
|
|
2NUG
 
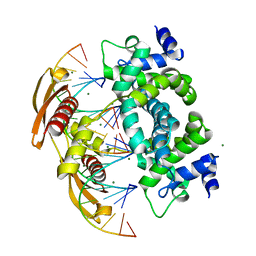 | | Crystal structure of RNase III from Aquifex aeolicus complexed with ds-RNA at 1.7-Angstrom Resolution | | Descriptor: | 5'-R(P*AP*AP*GP*GP*UP*CP*AP*UP*UP*CP*G)-3', 5'-R(P*AP*GP*UP*GP*GP*CP*CP*UP*UP*GP*C)-3', MAGNESIUM ION, ... | | Authors: | Gan, J.H, Shaw, G, Tropea, J.E, Waugh, D.S, Court, D.L, Ji, X. | | Deposit date: | 2006-11-09 | | Release date: | 2007-11-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A stepwise model for double-stranded RNA processing by ribonuclease III.
Mol.Microbiol., 67, 2007
|
|
2NXH
 
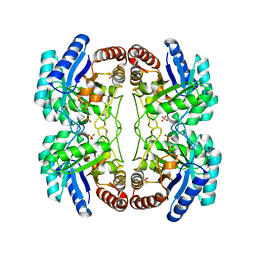 | | Structural and mechanistic changes along an engineered path from metallo to non-metallo KDO8P synthase. | | Descriptor: | 2-dehydro-3-deoxyphosphooctonate aldolase, PHOSPHATE ION, PHOSPHOENOLPYRUVATE | | Authors: | Kona, F, Xu, X, Martin, P, Kuzmic, P, Gatti, D.L. | | Deposit date: | 2006-11-17 | | Release date: | 2007-04-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Structural and Mechanistic Changes along an Engineered Path from Metallo to Nonmetallo 3-Deoxy-d-manno-octulosonate 8-Phosphate Synthases.
Biochemistry, 46, 2007
|
|
2NX3
 
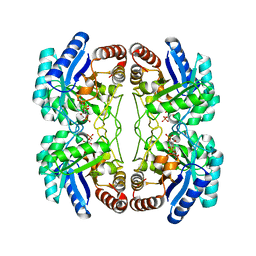 | | Structural and mechanistic changes along an engineered path from metallo to non-metallo KDO8P synthase | | Descriptor: | (2R,4R,5R,6R,7R)-2,4,5,6,7-PENTAHYDROXY-2,8-BIS(PHOSPHONOOXY)OCTANOIC ACID, 2-dehydro-3-deoxyphosphooctonate aldolase, ARABINOSE-5-PHOSPHATE, ... | | Authors: | Kona, F, Xu, X, Martin, P, Kuzmic, P, Gatti, D.L. | | Deposit date: | 2006-11-16 | | Release date: | 2007-04-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural and Mechanistic Changes along an Engineered Path from Metallo to Nonmetallo 3-Deoxy-d-manno-octulosonate 8-Phosphate Synthases.
Biochemistry, 46, 2007
|
|
2O94
 | |
2NWS
 
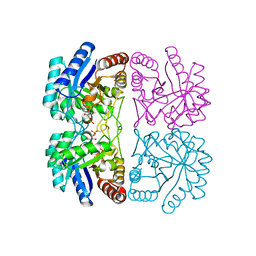 | | Structural and mechanistic changes along an engineered path from metallo to non-metallo KDO8P synthase | | Descriptor: | 2-dehydro-3-deoxyphosphooctonate aldolase, ARABINOSE-5-PHOSPHATE, PHOSPHOENOLPYRUVATE | | Authors: | Kona, F, Xu, X, Martin, P, Kuzmic, P, Gatti, D.L. | | Deposit date: | 2006-11-16 | | Release date: | 2007-04-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and Mechanistic Changes along an Engineered Path from Metallo to Nonmetallo 3-Deoxy-d-manno-octulosonate 8-Phosphate Synthases.
Biochemistry, 46, 2007
|
|
2NUF
 
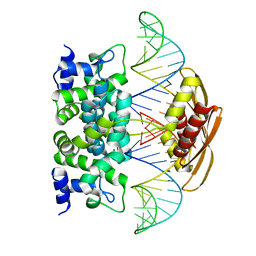 | | Crystal structure of RNase III from Aquifex aeolicus complexed with ds-RNA at 2.5-Angstrom Resolution | | Descriptor: | 28-MER, MAGNESIUM ION, Ribonuclease III | | Authors: | Gan, J.H, Shaw, G, Tropea, J.E, Waugh, D.S, Court, D.L, Ji, X. | | Deposit date: | 2006-11-09 | | Release date: | 2007-11-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A stepwise model for double-stranded RNA processing by ribonuclease III.
Mol.Microbiol., 67, 2007
|
|
2NWR
 
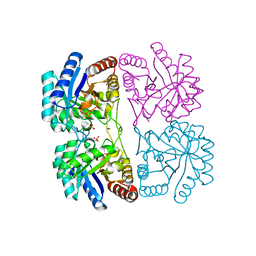 | | Crystal structure of C11N mutant of KDO8P Synthase in complex with PEP | | Descriptor: | 2-dehydro-3-deoxyphosphooctonate aldolase, PHOSPHATE ION, PHOSPHOENOLPYRUVATE | | Authors: | Kona, F, Xu, X, Martin, P, Kuzmic, P, Gatti, D.L. | | Deposit date: | 2006-11-16 | | Release date: | 2007-04-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural and Mechanistic Changes along an Engineered Path from Metallo to Nonmetallo 3-Deoxy-d-manno-octulosonate 8-Phosphate Synthases.
Biochemistry, 46, 2007
|
|
2NXI
 
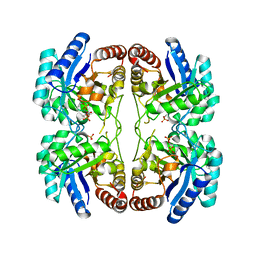 | | Structural and mechanistic changes along an engineered path from metallo to non-metallo KDO8P synthase. | | Descriptor: | 2-dehydro-3-deoxyphosphooctonate aldolase, PHOSPHATE ION, PHOSPHOENOLPYRUVATE | | Authors: | Kona, F, Xu, X, Martin, P, Kuzmic, P, Gatti, D.L. | | Deposit date: | 2006-11-17 | | Release date: | 2007-07-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and mechanistic changes along an engineered path from metallo to nonmetallo 3-deoxy-D-manno-octulosonate 8-phosphate synthases
Biochemistry, 46, 2007
|
|
2O93
 
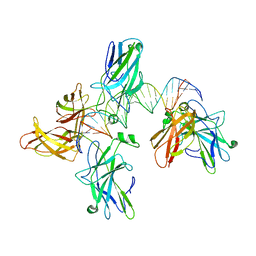 | |
2NX1
 
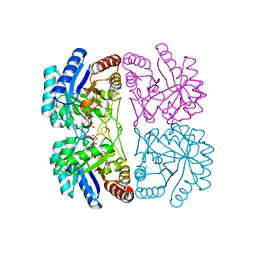 | | Structural and mechanistic changes along an engineered path from metallo to non-metallo KDO8P synthase | | Descriptor: | 2-dehydro-3-deoxyphosphooctonate aldolase, 5-O-phosphono-beta-D-ribofuranose, PHOSPHOENOLPYRUVATE | | Authors: | Kona, F, Xu, X, Martin, P, Kuzmic, P, Gatti, D.L. | | Deposit date: | 2006-11-16 | | Release date: | 2007-04-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and Mechanistic Changes along an Engineered Path from Metallo to Nonmetallo 3-Deoxy-d-manno-octulosonate 8-Phosphate Synthases.
Biochemistry, 46, 2007
|
|
2NUE
 
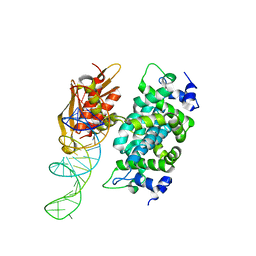 | | Crystal structure of RNase III from Aquifex aeolicus complexed with ds-RNA at 2.9-Angstrom Resolution | | Descriptor: | 46-MER, Ribonuclease III | | Authors: | Gan, J.H, Shaw, G, Tropea, J.E, Waugh, D.S, Court, D.L, Ji, X. | | Deposit date: | 2006-11-09 | | Release date: | 2007-11-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | A stepwise model for double-stranded RNA processing by ribonuclease III.
Mol.Microbiol., 67, 2007
|
|
2NXG
 
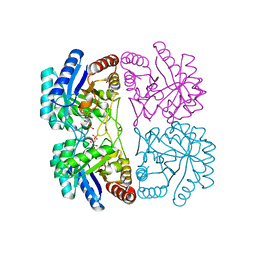 | | Structural and mechanistic changes along an engineered path from metallo to non-metallo KDO8P synthase. | | Descriptor: | (2R,4R,5R,6R,7R)-2,4,5,6,7-PENTAHYDROXY-2,8-BIS(PHOSPHONOOXY)OCTANOIC ACID, 2-dehydro-3-deoxyphosphooctonate aldolase, ARABINOSE-5-PHOSPHATE, ... | | Authors: | Kona, F, Xu, X, Martin, P, Kuzmic, P, Gatti, D.L. | | Deposit date: | 2006-11-17 | | Release date: | 2007-04-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural and Mechanistic Changes along an Engineered Path from Metallo to Nonmetallo 3-Deoxy-d-manno-octulosonate 8-Phosphate Synthases.
Biochemistry, 46, 2007
|
|
2NUL
 
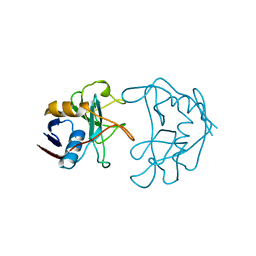 | | PEPTIDYLPROLYL ISOMERASE FROM E. COLI | | Descriptor: | PEPTIDYLPROLYL ISOMERASE | | Authors: | Edwards, K.J, Ollis, D.L. | | Deposit date: | 1996-11-14 | | Release date: | 1997-11-19 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of cytoplasmic Escherichia coli peptidyl-prolyl isomerase: evidence for decreased mobility of loops upon complexation.
J.Mol.Biol., 271, 1997
|
|
2OMB
 
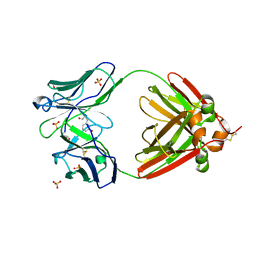 | | Bence Jones KWR Protein- Immunoglobulin Light Chain Dimer, P3(1)21 Crystal Form | | Descriptor: | Bence Jones KWR Protein - Immunoglobulin Light Chain, PHENOL, SULFATE ION | | Authors: | Makino, D.L, Larson, S.B, McPherson, A. | | Deposit date: | 2007-01-21 | | Release date: | 2007-07-17 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Bence Jones KWR protein structures determined by X-ray crystallography.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2OLD
 
 | | Bence Jones KWR Protein- Immunoglobulin Light Chain Dimer, P3(2)21 Crystal Form | | Descriptor: | Bence Jones KWR Protein - Immunoglobulin Light Chain, PHENOL, PHOSPHATE ION | | Authors: | Makino, D.L, Larson, S.B, McPherson, A. | | Deposit date: | 2007-01-19 | | Release date: | 2007-07-17 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Bence Jones KWR protein structures determined by X-ray crystallography.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2OMN
 
 | | Bence Jones KWR Protein- Immunoglobulin Light Chain Dimer, P4(3)2(1)2 Crystal Form | | Descriptor: | Bence Jones KWR Protein - Immunoglobulin Light Chain, PHENOL, SULFATE ION | | Authors: | Makino, D.L, Larson, S.B, McPherson, A. | | Deposit date: | 2007-01-22 | | Release date: | 2007-07-17 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Bence Jones KWR protein structures determined by X-ray crystallography.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
