5ECA
 
 | |
6TG7
 
 | |
5I11
 
 | |
3M0P
 
 | |
3M0S
 
 | |
8AH4
 
 | |
8AH7
 
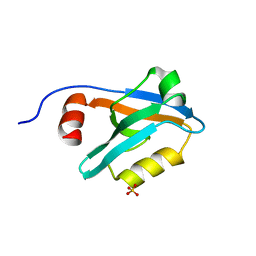 | |
2O88
 | |
7NER
 
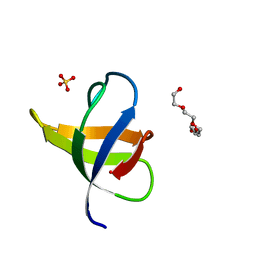 | |
7NET
 
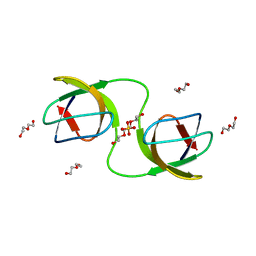 | |
7NES
 | |
1Z9K
 
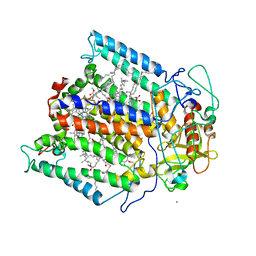 | |
5EC7
 | |
5DK8
 | |
2HKI
 
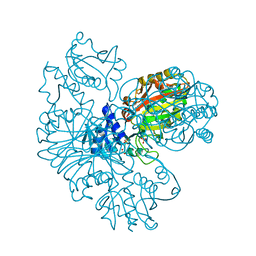 | |
2HDA
 
 | | Yes SH3 domain | | Descriptor: | Proto-oncogene tyrosine-protein kinase Yes, SULFATE ION | | Authors: | Camara-Artigas, A, Luque, I, Ruiz-Sanz, J, Mateo, P.L, Martin-Garcia, J.M. | | Deposit date: | 2006-06-20 | | Release date: | 2007-04-17 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystallographic structure of the SH3 domain of the human c-Yes tyrosine kinase: Loop flexibility and amyloid aggregation.
Febs Lett., 581, 2007
|
|
7ZLX
 
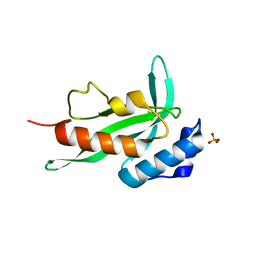 | |
7ZR2
 | | Crystal structure of a chimeric protein mimic of SARS-CoV-2 Spike HR1 in complex with HR2 | | Descriptor: | Spike protein S2', Spike protein S2',Chimeric protein mimic of SARS-CoV-2 Spike HR1 | | Authors: | Camara-Artigas, A, Gavira, J.A, Cano-Munoz, M, Polo-Megias, D, Conejero-Lara, F. | | Deposit date: | 2022-05-03 | | Release date: | 2022-11-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Novel chimeric proteins mimicking SARS-CoV-2 spike epitopes with broad inhibitory activity.
Int.J.Biol.Macromol., 222, 2022
|
|
6QJJ
 
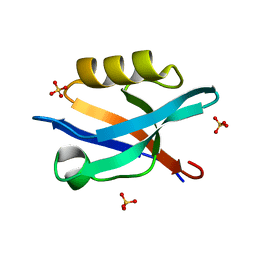 | |
6QJD
 
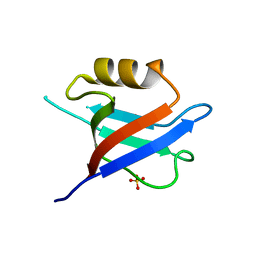 | |
6QJN
 
 | |
6QJI
 
 | |
8AH5
 
 | |
8AH6
 
 | |
8AH8
 | |
