3JU8
 
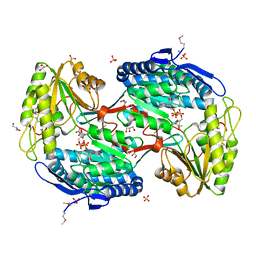 | | Crystal Structure of Succinylglutamic Semialdehyde Dehydrogenase from Pseudomonas aeruginosa. | | Descriptor: | CHLORIDE ION, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Kim, Y, Li, H, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-09-14 | | Release date: | 2009-09-22 | | Last modified: | 2011-11-16 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Crystal Structure of Succinylglutamic Semialdehyde Dehydrogenase from Pseudomonas aeruginosa
To be Published
|
|
3NKL
 
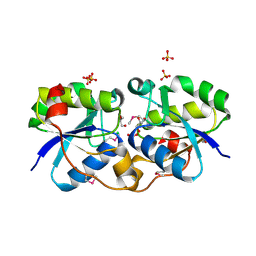 | | Crystal Structure of UDP-D-Quinovosamine 4-Dehydrogenase from Vibrio fischeri | | Descriptor: | ACETIC ACID, CHLORIDE ION, GLYCEROL, ... | | Authors: | Kim, Y, Mack, J, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-06-20 | | Release date: | 2010-08-18 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of UDP-D-Quinovosamine 4-Dehydrogenase from Vibrio fischeri
To be Published, 2010
|
|
3LL7
 
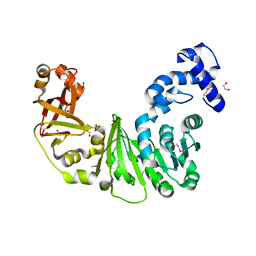 | | Crystal structure of putative methyltransferase PG_1098 from Porphyromonas gingivalis W83 | | Descriptor: | 1,2-ETHANEDIOL, FORMIC ACID, Putative methyltransferase | | Authors: | Chang, C, Chhor, G, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-01-28 | | Release date: | 2010-02-16 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of putative methyltransferase PG_1098 from Porphyromonas gingivalis W83
To be Published
|
|
3LAO
 
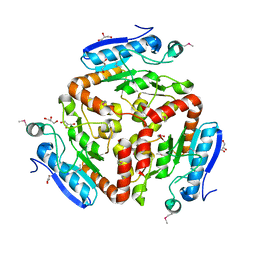 | |
3PVZ
 
 | | UDP-N-acetylglucosamine 4,6-dehydratase from Vibrio fischeri | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, SODIUM ION, THIOCYANATE ION, ... | | Authors: | Osipiuk, J, Marshall, N, Tesar, C, Pearson, L, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-12-07 | | Release date: | 2011-01-12 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | UDP-N-acetylglucosamine 4,6-dehydratase from Vibrio fischeri.
To be Published
|
|
3O5Y
 
 | |
3ISR
 
 | | The Crystal Structure of a Putative Cysteine Protease from Cytophaga hutchinsonii to 1.9A | | Descriptor: | Transglutaminase-like enzymes, putative cysteine protease | | Authors: | Stein, A.J, Bigelow, L, Trevino, D, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-08-27 | | Release date: | 2009-09-08 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Crystal Structure of a Putative Cysteine Protease from Cytophaga hutchinsonii to 1.9A
To be Published
|
|
3IDD
 
 | |
3P9Z
 
 | | Crystal structure of uroporphyrinogen-III synthetase from Helicobacter pylori 26695 | | Descriptor: | MALONATE ION, Uroporphyrinogen III cosynthase (HemD) | | Authors: | Nocek, B, Stein, A, Chhor, G, Fenske, R.J, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-10-18 | | Release date: | 2010-11-03 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of uroporphyrinogen-III synthetase from Helicobacter pylori 26695
To be Published
|
|
3KD8
 
 | |
3MWB
 
 | | The Crystal Structure of Prephenate dehydratase in complex with L-Phe from Arthrobacter aurescens to 2.0A | | Descriptor: | MAGNESIUM ION, PHENYLALANINE, Prephenate dehydratase | | Authors: | Stein, A.J, Chhor, G, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-05-05 | | Release date: | 2010-06-30 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Crystal Structure of Prephenate dehydratase in complex with L-Phe from Arthrobacter aurescens to 2.0A
To be Published
|
|
3HSI
 
 | |
3LEC
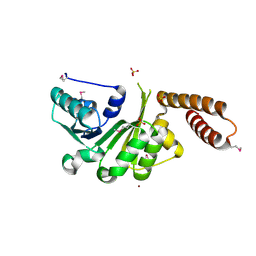 
 | | The Crystal Structure of a protein in the NADB-Rossmann Superfamily from Streptococcus agalactiae to 1.8A | | Descriptor: | NADB-Rossmann Superfamily protein, SULFATE ION, ZINC ION | | Authors: | Stein, A.J, Hatzos, C, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-01-14 | | Release date: | 2010-01-26 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Crystal Structure of a protein in the NADB-Rossmann Superfamily from Streptococcus agalactiae to 1.8A
To be Published
|
|
3LR1
 
 | | The crystal structure of the tungstate ABC transporter from Geobacter sulfurreducens | | Descriptor: | GLYCEROL, TUNGSTEN ION, Tungstate ABC transporter, ... | | Authors: | Zhang, R, Volkart, L, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-02-10 | | Release date: | 2010-03-16 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The crystal structure of the tungstate ABC transporter from Geobacter sulfurreducens
To be Published
|
|
3NE8
 
 | | The crystal structure of a domain from N-acetylmuramoyl-l-alanine amidase of Bartonella henselae str. Houston-1 | | Descriptor: | ACETATE ION, FORMIC ACID, GLYCEROL, ... | | Authors: | Tan, K, Rakowski, E, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-06-08 | | Release date: | 2010-07-14 | | Last modified: | 2012-11-07 | | Method: | X-RAY DIFFRACTION (1.239 Å) | | Cite: | A conformational switch controls cell wall-remodelling enzymes required for bacterial cell division.
Mol.Microbiol., 85, 2012
|
|
