1XFD
 
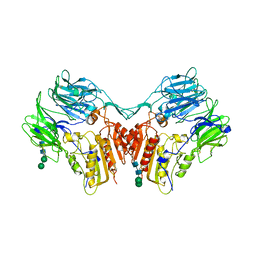 | | Structure of a human A-type Potassium Channel Accelerating factor DPPX, a member of the dipeptidyl aminopeptidase family | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Dipeptidyl aminopeptidase-like protein 6, alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Strop, P, Bankovich, A.J, Hansen, K.C, Garcia, K.C, Brunger, A.T. | | Deposit date: | 2004-09-14 | | Release date: | 2004-10-26 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure of a human A-type potassium channel interacting protein DPPX, a member of the dipeptidyl aminopeptidase family
J.Mol.Biol., 343, 2004
|
|
4QQP
 
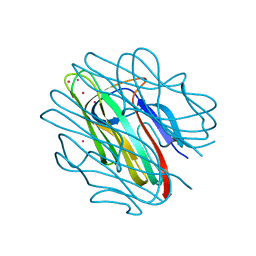 | | Crystal structure of C1QL3 mutant D207A | | Descriptor: | CADMIUM ION, Complement C1q-like protein 3 | | Authors: | Ressl, S, Brunger, A.T. | | Deposit date: | 2014-06-27 | | Release date: | 2015-04-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.464 Å) | | Cite: | Structures of C1q-like Proteins Reveal Unique Features among the C1q/TNF Superfamily.
Structure, 23, 2015
|
|
3EZW
 
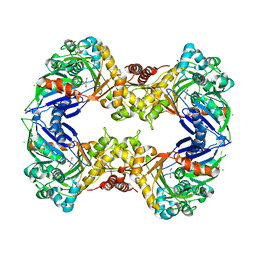 | | Crystal Structure of a Hyperactive Escherichia coli Glycerol Kinase Mutant Gly230 --> Asp Obtained Using Microfluidic Crystallization Devices | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, GLYCEROL, ... | | Authors: | Anderson, M.J, DeLaBarre, B, Dunten, P, Brunger, A.T, Quake, S.R. | | Deposit date: | 2008-10-23 | | Release date: | 2008-11-04 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of a hyperactive Escherichia coli glycerol kinase mutant Gly230 --> Asp obtained using microfluidic crystallization devices.
Biochemistry, 46, 2007
|
|
3J99
 
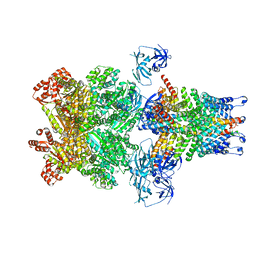 | | Structure of 20S supercomplex determined by single particle cryoelectron microscopy (State IIIb) | | Descriptor: | Alpha-soluble NSF attachment protein, Synaptosomal-associated protein 25, Syntaxin-1A, ... | | Authors: | Zhao, M, Wu, S, Cheng, Y, Brunger, A.T. | | Deposit date: | 2014-12-05 | | Release date: | 2015-01-28 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (8.2 Å) | | Cite: | Mechanistic insights into the recycling machine of the SNARE complex.
Nature, 518, 2015
|
|
3J96
 
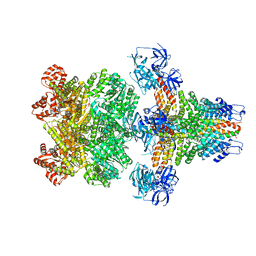 | | Structure of 20S supercomplex determined by single particle cryoelectron microscopy (State I) | | Descriptor: | Alpha-soluble NSF attachment protein, Synaptosomal-associated protein 25, Syntaxin-1A, ... | | Authors: | Zhao, M, Wu, S, Cheng, Y, Brunger, A.T. | | Deposit date: | 2014-12-05 | | Release date: | 2015-01-28 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (7.6 Å) | | Cite: | Mechanistic insights into the recycling machine of the SNARE complex.
Nature, 518, 2015
|
|
3J97
 
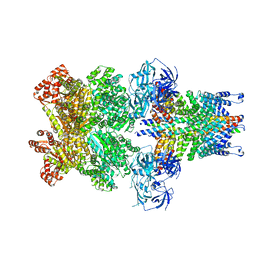 | | Structure of 20S supercomplex determined by single particle cryoelectron microscopy (State II) | | Descriptor: | Alpha-soluble NSF attachment protein, Synaptosomal-associated protein 25, Syntaxin-1A, ... | | Authors: | Zhao, M, Wu, S, Cheng, Y, Brunger, A.T. | | Deposit date: | 2014-12-05 | | Release date: | 2015-01-28 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (7.8 Å) | | Cite: | Mechanistic insights into the recycling machine of the SNARE complex.
Nature, 518, 2015
|
|
3J98
 
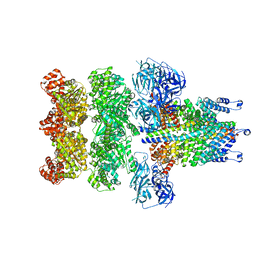 | | Structure of 20S supercomplex determined by single particle cryoelectron microscopy (State IIIa) | | Descriptor: | Alpha-soluble NSF attachment protein, Synaptosomal-associated protein 25, Syntaxin-1A, ... | | Authors: | Zhao, M, Wu, S, Cheng, Y, Brunger, A.T. | | Deposit date: | 2014-12-05 | | Release date: | 2015-01-28 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (8.4 Å) | | Cite: | Mechanistic insights into the recycling machine of the SNARE complex.
Nature, 518, 2015
|
|
3HN8
 
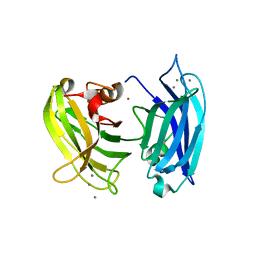 | | Crystal structure of synaptotagmin | | Descriptor: | CALCIUM ION, Synaptotagmin-3, ZINC ION | | Authors: | Strop, P, Vrljic, M, Ernst, J, Brunger, A.T. | | Deposit date: | 2009-05-30 | | Release date: | 2010-02-23 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Molecular mechanism of the synaptotagmin-SNARE interaction in Ca2+-triggered vesicle fusion.
Nat.Struct.Mol.Biol., 17, 2010
|
|
1N0Y
 
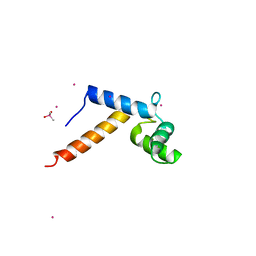 | | Crystal Structure of Pb-bound Calmodulin | | Descriptor: | ACETATE ION, CACODYLATE ION, Calmodulin, ... | | Authors: | Wilson, M.A, Brunger, A.T. | | Deposit date: | 2002-10-15 | | Release date: | 2003-09-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Domain flexibility in the 1.75 A resolution structure of Pb2+-calmodulin.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1XTG
 
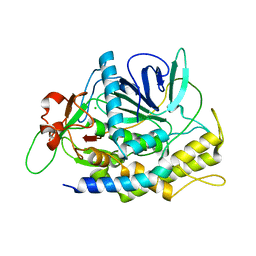 | |
1XTF
 
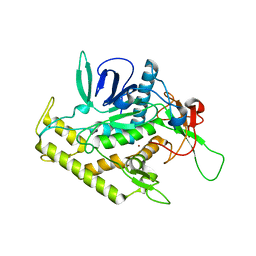 | |
1KWA
 
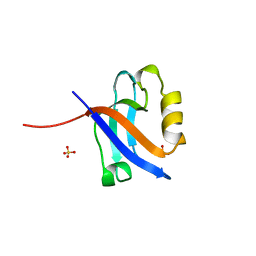 | | HUMAN CASK/LIN-2 PDZ DOMAIN | | Descriptor: | HCASK/LIN-2 PROTEIN, SULFATE ION | | Authors: | Daniels, D.L, Cohen, A.R, Anderson, J.M, Brunger, A.T. | | Deposit date: | 1998-01-16 | | Release date: | 1998-05-27 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Crystal structure of the hCASK PDZ domain reveals the structural basis of class II PDZ domain target recognition
Nat.Struct.Biol., 5, 1998
|
|
1UAD
 
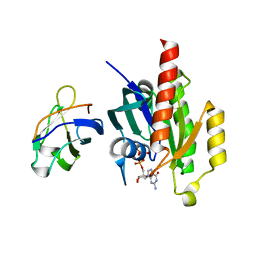 | | Crystal structure of the RalA-GppNHp-Sec5 Ral-binding domain complex | | Descriptor: | Exocyst complex component Sec5, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Fukai, S, Matern, H.T, Scheller, R.H, Brunger, A.T. | | Deposit date: | 2003-03-09 | | Release date: | 2003-07-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural basis of the interaction between RalA and Sec5, a subunit of the Sec6/8 complex
EMBO J., 22, 2003
|
|
3J94
 
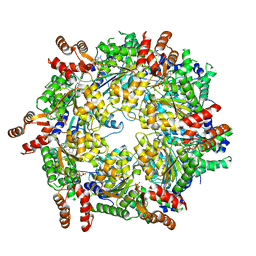 | | Structure of ATP-bound N-ethylmaleimide sensitive factor determined by single particle cryoelectron microscopy | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Vesicle-fusing ATPase | | Authors: | Zhao, M, Wu, S, Cheng, Y, Brunger, A.T. | | Deposit date: | 2014-12-05 | | Release date: | 2015-01-28 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Mechanistic insights into the recycling machine of the SNARE complex.
Nature, 518, 2015
|
|
3FSH
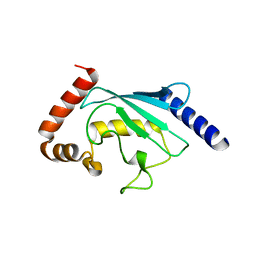 
 | |
4QQO
 
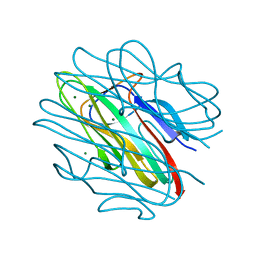 | | Crystal structure of C1QL3 mutant D205A | | Descriptor: | Complement C1q-like protein 3, MAGNESIUM ION | | Authors: | Ressl, S, Brunger, A.T. | | Deposit date: | 2014-06-27 | | Release date: | 2015-04-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.031 Å) | | Cite: | Structures of C1q-like Proteins Reveal Unique Features among the C1q/TNF Superfamily.
Structure, 23, 2015
|
|
4QQL
 
 | | Crystal structure of C1QL3 in P1 space group | | Descriptor: | Complement C1q-like protein 3, MAGNESIUM ION | | Authors: | Ressl, S, Brunger, A.T. | | Deposit date: | 2014-06-27 | | Release date: | 2015-04-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Structures of C1q-like Proteins Reveal Unique Features among the C1q/TNF Superfamily.
Structure, 23, 2015
|
|
3J95
 
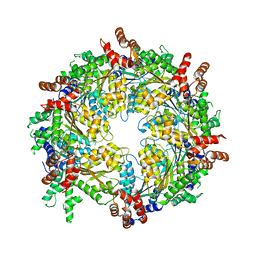 | | Structure of ADP-bound N-ethylmaleimide sensitive factor determined by single particle cryoelectron microscopy | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Vesicle-fusing ATPase | | Authors: | Zhao, M, Wu, S, Cheng, Y, Brunger, A.T. | | Deposit date: | 2014-12-05 | | Release date: | 2015-01-28 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (7.6 Å) | | Cite: | Mechanistic insights into the recycling machine of the SNARE complex.
Nature, 518, 2015
|
|
1NSF
 
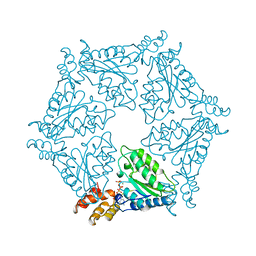 | | D2 HEXAMERIZATION DOMAIN OF N-ETHYLMALEIMIDE SENSITIVE FACTOR (NSF) | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, N-ETHYLMALEIMIDE SENSITIVE FACTOR | | Authors: | Yu, R.C, Hanson, P.I, Jahn, R, Brunger, A.T. | | Deposit date: | 1998-06-26 | | Release date: | 1998-11-25 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of the ATP-dependent oligomerization domain of N-ethylmaleimide sensitive factor complexed with ATP.
Nat.Struct.Biol., 5, 1998
|
|
1OEL
 
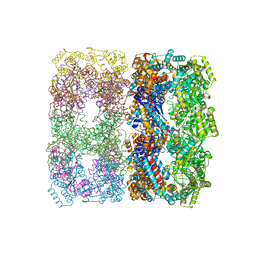 | |
1YTT
 
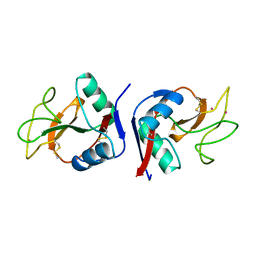 | | YB SUBSTITUTED SUBTILISIN FRAGMENT OF MANNOSE BINDING PROTEIN-A (SUB-MBP-A), MAD STRUCTURE AT 110K | | Descriptor: | MANNOSE-BINDING PROTEIN A, YTTERBIUM (III) ION | | Authors: | Burling, F.T, Weis, W.I, Flaherty, K.M, Brunger, A.T. | | Deposit date: | 1995-11-09 | | Release date: | 1996-06-10 | | Last modified: | 2019-11-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Direct observation of protein solvation and discrete disorder with experimental crystallographic phases.
Science, 271, 1996
|
|
1GTO
 
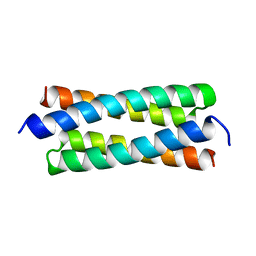 | |
2BBG
 
 | |
1SFC
 
 | | NEURONAL SYNAPTIC FUSION COMPLEX | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, PROTEIN (SNAP-25B), PROTEIN (SYNAPTOBREVIN 2), ... | | Authors: | Sutton, R.B, Brunger, A.T. | | Deposit date: | 1998-08-24 | | Release date: | 1998-10-28 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of a SNARE complex involved in synaptic exocytosis at 2.4 A resolution.
Nature, 395, 1998
|
|
1SX3
 
 | | GroEL14-(ATPgammaS)14 | | Descriptor: | MAGNESIUM ION, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, POTASSIUM ION, ... | | Authors: | Chaudhry, C, Horwich, A.L, Brunger, A.T, Adams, P.D. | | Deposit date: | 2004-03-30 | | Release date: | 2005-03-01 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Exploring the structural dynamics of the E.coli chaperonin GroEL using translation-libration-screw crystallographic refinement of intermediate states.
J.Mol.Biol., 342, 2004
|
|
