3ZUC
 | | Structure of CBM3b of major scaffoldin subunit ScaA from Acetivibrio cellulolyticus determined from the crystals grown in the presence of Nickel | | 分子名称: | 1,2-ETHANEDIOL, CALCIUM ION, CELLULOSOMAL SCAFFOLDIN, ... | | 著者 | Yaniv, O, Halfon, Y, Lamed, R, Frolow, F. | | 登録日 | 2011-07-18 | | 公開日 | 2012-01-11 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.001 Å) | | 主引用文献 | Structure of Cbm3B of the Major Scaffoldin Subunit Scaa from Acetivibrio Cellulolyticus
Acta Crystallogr.,Sect.F, 68, 2012
|
|
3ZQW
 | | Structure of CBM3b of major scaffoldin subunit ScaA from Acetivibrio cellulolyticus | | 分子名称: | 1,2-ETHANEDIOL, CALCIUM ION, CELLULOSOMAL SCAFFOLDIN, ... | | 著者 | Yaniv, O, Halfon, Y, Lamed, R, Frolow, F. | | 登録日 | 2011-06-12 | | 公開日 | 2012-01-11 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (1.07 Å) | | 主引用文献 | Structure of Cbm3B of the Major Scaffoldin Subunit Scaa from Acetivibrio Cellulolyticus
Acta Crystallogr.,Sect.F, 68, 2012
|
|
6S9A
 
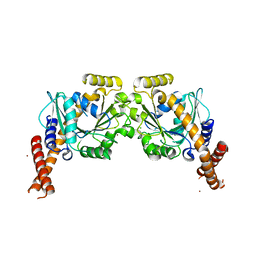 | | Artificial GTPase-BSE dimer of human Dynamin1 | | 分子名称: | CHLORIDE ION, Dynamin-1,Dynamin-1, ZINC ION | | 著者 | Ganichkin, O.M, Vancraenenbroeck, R, Rosenblum, G, Hofmann, H, Daumke, O, Noel, J.K. | | 登録日 | 2019-07-11 | | 公開日 | 2020-08-26 | | 最終更新日 | 2024-01-24 | | 実験手法 | X-RAY DIFFRACTION (1.86 Å) | | 主引用文献 | Quantification and demonstration of the collective constriction-by-ratchet mechanism in the dynamin molecular motor.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
6TEO
 
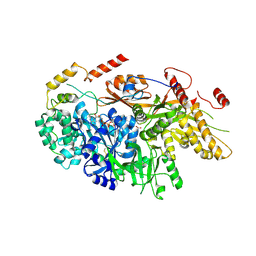 | | Crystal structure of a yeast Snu114-Prp8 complex | | 分子名称: | GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, Pre-mRNA-splicing factor 8, ... | | 著者 | Ganichkin, O, Jia, J, Loll, B, Absmeier, E, Wahl, M.C. | | 登録日 | 2019-11-12 | | 公開日 | 2020-03-18 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (3.1 Å) | | 主引用文献 | A Snu114-GTP-Prp8 module forms a relay station for efficient splicing in yeast.
Nucleic Acids Res., 48, 2020
|
|
7OFV
 
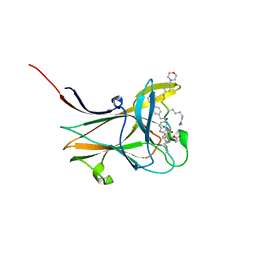 | | NMR-guided design of potent and selective EphA4 agonistic ligands | | 分子名称: | ACETATE ION, EphA4 agonist ligand, Ephrin type-A receptor 4 | | 著者 | Ganichkin, O.M, Craig, T.K, Baggio, C, Pellecchia, M. | | 登録日 | 2021-05-05 | | 公開日 | 2021-08-11 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.43 Å) | | 主引用文献 | NMR-Guided Design of Potent and Selective EphA4 Agonistic Ligands.
J.Med.Chem., 64, 2021
|
|
7S25
 
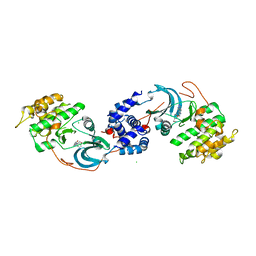 | | ROCK1 IN COMPLEX WITH LIGAND G4998 | | 分子名称: | 2-[3-(methoxymethyl)phenyl]-N-[4-(1H-pyrazol-4-yl)phenyl]acetamide, CHLORIDE ION, Rho-associated protein kinase 1 | | 著者 | Ganichkin, O, Harris, S.F, Steinbacher, S. | | 登録日 | 2021-09-03 | | 公開日 | 2022-10-05 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (2.337 Å) | | 主引用文献 | Chemical space docking enables large-scale structure-based virtual screening to discover ROCK1 kinase inhibitors.
Nat Commun, 13, 2022
|
|
7S26
 
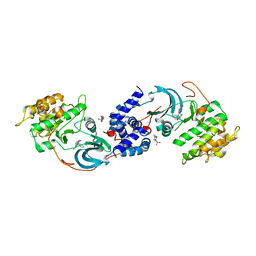 | | ROCK1 IN COMPLEX WITH LIGAND G5018 | | 分子名称: | 2-[methyl(phenyl)amino]-1-[4-(1H-pyrrolo[2,3-b]pyridin-3-yl)-3,6-dihydropyridin-1(2H)-yl]ethan-1-one, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Rho-associated protein kinase 1 | | 著者 | Ganichkin, O, Harris, S.F, Steinbacher, S. | | 登録日 | 2021-09-03 | | 公開日 | 2022-10-05 | | 最終更新日 | 2024-05-22 | | 実験手法 | X-RAY DIFFRACTION (2.744 Å) | | 主引用文献 | Chemical space docking enables large-scale structure-based virtual screening to discover ROCK1 kinase inhibitors.
Nat Commun, 13, 2022
|
|
2V9V
 
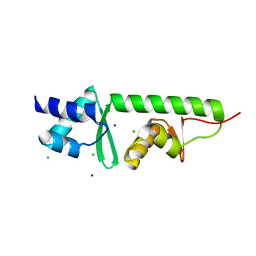 | |
3ZQX
 | | Carbohydrate-binding module CBM3b from the cellulosomal cellobiohydrolase 9A from Clostridium thermocellum | | 分子名称: | CALCIUM ION, CELLULOSE 1,4-BETA-CELLOBIOSIDASE | | 著者 | Yaniv, O, Petkun, S, Shimon, L.J.W, Bayer, E.A, Lamed, R, Frolow, F. | | 登録日 | 2011-06-12 | | 公開日 | 2012-04-25 | | 最終更新日 | 2024-05-01 | | 実験手法 | X-RAY DIFFRACTION (1.04 Å) | | 主引用文献 | A Single Mutation Reforms the Binding Activity of an Adhesion-Deficient Family 3 Carbohydrate-Binding Module
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3ZU8
 | | STRUCTURE OF CBM3B OF MAJOR SCAFFOLDIN SUBUNIT SCAA FROM ACETIVIBRIO CELLULOLYTICUS DETERMINED ON THE NIKEL ABSORPTION EDGE | | 分子名称: | 1,2-ETHANEDIOL, CALCIUM ION, CELLULOSOMAL SCAFFOLDIN, ... | | 著者 | Yaniv, O, Halfon, Y, Lamed, R, Frolow, F. | | 登録日 | 2011-07-17 | | 公開日 | 2012-01-11 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (1.801 Å) | | 主引用文献 | Structure of Cbm3B of the Major Scaffoldin Subunit Scaa from Acetivibrio Cellulolyticus
Acta Crystallogr.,Sect.F, 68, 2012
|
|
4B9P
 
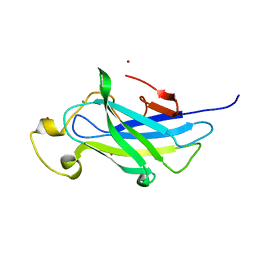 | | Biomass sensoring module from putative Rsgi2 protein of Clostridium thermocellum resemble family 3 carbohydrate-binding module of cellulosome | | 分子名称: | CALCIUM ION, TYPE 3A CELLULOSE-BINDING DOMAIN PROTEIN, ZINC ION | | 著者 | Yaniv, O, Shimon, L.J.W, Bayer, E.A, Lamed, R, Frolow, F. | | 登録日 | 2012-09-06 | | 公開日 | 2013-09-11 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.182 Å) | | 主引用文献 | Fine-Structural Variance of Family 3 Carbohydrate-Binding Modules as Extracellular Biomass-Sensing Components of Clostridium Thermocellum Anti-Sigma(I) Factors.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4B96
 | | Family 3b carbohydrate-binding module from the biomass sensoring system of Clostridium clariflavum | | 分子名称: | CALCIUM ION, CELLULOSE BINDING DOMAIN-CONTAINING PROTEIN, CHLORIDE ION | | 著者 | Yaniv, O, Reddy, Y.H.K, Yoffe, H, Shimon, L.J.W, Bayer, E.A, Lamed, R, Frolow, F. | | 登録日 | 2012-09-02 | | 公開日 | 2013-09-18 | | 最終更新日 | 2024-05-01 | | 実験手法 | X-RAY DIFFRACTION (1.911 Å) | | 主引用文献 | Structure of Cbm3B from the Biomass Sensoring System of Clostridium Clarifalvum
To be Published
|
|
4C8X
 | |
4B97
 | | Biomass sensing modules from putative Rsgi-like proteins of Clostridium thermocellum resemble family 3 carbohydrate-binding module of cellulosome | | 分子名称: | CALCIUM ION, CELLULOSE BINDING DOMAIN-CONTAINING PROTEIN | | 著者 | Yaniv, O, Fichman, G, Shimon, L.J.W, Bayer, E.A, Lamed, R, Frolow, F. | | 登録日 | 2012-09-03 | | 公開日 | 2013-09-11 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.276 Å) | | 主引用文献 | Fine-Structural Variance of Family 3 Carbohydrate-Binding Modules as Extracellular Biomass-Sensing Components of Clostridium Thermocellum Anti-Sigma(I) Factors.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4B9F
 | | High resolution structure for family 3a carbohydrate binding module from the cipA scaffolding of clostridium thermocellum | | 分子名称: | CALCIUM ION, CELLULOSOMAL-SCAFFOLDING PROTEIN A, SULFATE ION | | 著者 | Yaniv, O, Shimon, L.J.W, Bayer, E.A, Lamed, R, Frolow, F. | | 登録日 | 2012-09-04 | | 公開日 | 2012-09-12 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.19 Å) | | 主引用文献 | High Resolution Structure of the Family 3A Carbohydrate-Binding Module from the Mafor Scaffoldin Subunit Cipa of Clostridium Thermocellum
To be Published
|
|
4B9C
 | | Biomass sensoring modules from putative Rsgi-like proteins of Clostridium thermocellum resemble family 3 carbohydrate-binding module of cellulosome | | 分子名称: | CALCIUM ION, TYPE 3A CELLULOSE-BINDING DOMAIN PROTEIN | | 著者 | Yaniv, O, Shimon, L.J.W, Bayer, E.A, Lamed, R, Frolow, F. | | 登録日 | 2012-09-04 | | 公開日 | 2013-09-11 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.171 Å) | | 主引用文献 | Fine-Structural Variance of Family 3 Carbohydrate-Binding Modules as Extracellular Biomass-Sensing Components of Clostridium Thermocellum Anti-Sigma(I) Factors.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
2YLK
 
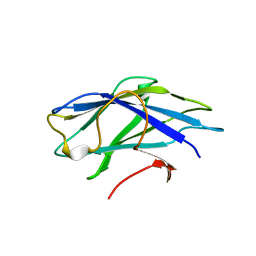 | | Carbohydrate-binding module CBM3b from the cellulosomal cellobiohydrolase 9A from Clostridium thermocellum | | 分子名称: | CELLULOSE 1,4-BETA-CELLOBIOSIDASE | | 著者 | Yaniv, O, Petkun, S, Shimon, L.J.W, Bayer, E.A, Lamed, R, Frolow, F. | | 登録日 | 2011-06-02 | | 公開日 | 2012-04-25 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | A Single Mutation Reforms the Binding Activity of an Adhesion-Deficient Family 3 Carbohydrate-Binding Module
Acta Crystallogr.,Sect.D, 68, 2012
|
|
2XBT
 | | Structure of a scaffoldin carbohydrate-binding module family 3b from the cellulosome of Bacteroides cellulosolvens: Structural diversity and implications for carbohydrate binding | | 分子名称: | CELLULOSOMAL SCAFFOLDIN, NITRATE ION | | 著者 | Yaniv, O, Shimon, L.J.W, Bayer, E.A, Lamed, R, Frolow, F. | | 登録日 | 2010-04-15 | | 公開日 | 2011-04-06 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.832 Å) | | 主引用文献 | Scaffoldin-Borne Family 3B Carbohydrate-Binding Module from the Cellulosome of Bacteroides Cellulosolvens: Structural Diversity and Significance of Calcium for Carbohydrate Binding
Acta Crystallogr.,Sect.D, 67, 2011
|
|
3BC8
 
 | |
3BCA
 
 | |
3BCB
 
 | |
8ERX
 
 | | Structure of chimeric HLA-A*11:01-A*02:01 bound to HIV-1 RT peptide | | 分子名称: | Beta-2-microglobulin, HIV-1 RT, HLA-A*02:01 | | 著者 | Florio, T.J, Ani, O, Young, M.C, Mallik, L, Sgourakis, N.G. | | 登録日 | 2022-10-13 | | 公開日 | 2023-01-25 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (2.07 Å) | | 主引用文献 | Decoupling peptide binding from T cell receptor recognition with engineered chimeric MHC-I molecules.
Front Immunol, 14, 2023
|
|
8ESH
 
 | | Structure of chimeric HLA-A*02:01 bound to CMV peptide | | 分子名称: | Beta-2-microglobulin, CMV peptide, HLA-A*02:01 | | 著者 | Florio, T.J, Ani, O, Young, M.C, Mallik, L, Sgourakis, N.G. | | 登録日 | 2022-10-14 | | 公開日 | 2023-01-25 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (2.72 Å) | | 主引用文献 | Decoupling peptide binding from T cell receptor recognition with engineered chimeric MHC-I molecules.
Front Immunol, 14, 2023
|
|
2POQ
 
 | | Dimeric Dihydrodiol Dehydrogenase complexed with inhibitor, Isoascorbic acid | | 分子名称: | BETA-MERCAPTOETHANOL, Dimeric dihydrodiol dehydrogenase, ISOASCORBIC ACID, ... | | 著者 | Carbone, V, El-Kabbani, O. | | 登録日 | 2007-04-27 | | 公開日 | 2007-07-31 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (2.59 Å) | | 主引用文献 | Structure of monkey dimeric dihydrodiol dehydrogenase in complex with isoascorbic acid.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
3C3U
 
 | | Crystal structure of AKR1C1 in complex with NADP and 3,5-dichlorosalicylic acid | | 分子名称: | 3,5-dichloro-2-hydroxybenzoic acid, Aldo-keto reductase family 1 member C1, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | 著者 | Dhagat, U, El-Kabbani, O. | | 登録日 | 2008-01-28 | | 公開日 | 2008-08-26 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Selectivity determinants of inhibitor binding to human 20alpha-hydroxysteroid dehydrogenase: crystal structure of the enzyme in ternary complex with coenzyme and the potent inhibitor 3,5-dichlorosalicylic acid
J.Med.Chem., 51, 2008
|
|
