4XMW
 
 | |
4XO5
 
 | |
4XMV
 
 | |
4XN1
 
 | |
4XNB
 
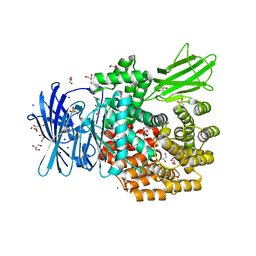 | |
2B3L
 
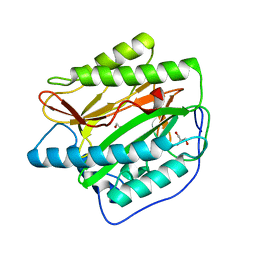 | | Crystal structure of type I human methionine aminopeptidase in the apo form | | Descriptor: | ACETIC ACID, GLYCEROL, Methionine aminopeptidase 1, ... | | Authors: | Addlagatta, A, Hu, X, Liu, J.O, Matthews, B.W. | | Deposit date: | 2005-09-20 | | Release date: | 2005-11-22 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural Basis for the Functional Differences between Type I and Type II Human Methionine Aminopeptidases(,).
Biochemistry, 44, 2005
|
|
2B3K
 
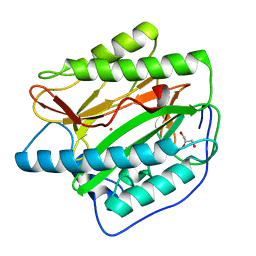 | | Crystal structure of Human Methionine Aminopeptidase Type I in the holo form | | Descriptor: | ACETATE ION, COBALT (II) ION, GLYCEROL, ... | | Authors: | Addlagatta, A, Hu, X, Liu, J.O, Matthews, B.W. | | Deposit date: | 2005-09-20 | | Release date: | 2005-11-22 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural Basis for the Functional Differences between Type I and Type II Human Methionine Aminopeptidases(,).
Biochemistry, 44, 2005
|
|
2B3H
 
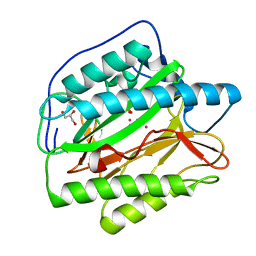 | | Crystal structure of Human Methionine Aminopeptidase Type I with a third cobalt in the active site | | Descriptor: | CHLORIDE ION, COBALT (II) ION, GLYCEROL, ... | | Authors: | Addlagatta, A, Hu, X, Liu, J.O, Matthews, B.W. | | Deposit date: | 2005-09-20 | | Release date: | 2005-11-22 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Structural Basis for the Functional Differences between Type I and Type II Human Methionine Aminopeptidases(,).
Biochemistry, 44, 2005
|
|
2GZ5
 
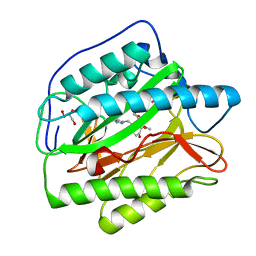 | | Human Type 1 methionine aminopeptidase in complex with ovalicin at 1.1 Ang | | Descriptor: | 3,4-DIHYDROXY-2-METHOXY-4-METHYL-3-[2-METHYL-3-(3-METHYL-BUT-2-ENYL) -OXIRANYL]-CYCLOHEXANONE, COBALT (II) ION, GLYCEROL, ... | | Authors: | Addlagatta, A, Matthews, B.W. | | Deposit date: | 2006-05-10 | | Release date: | 2006-05-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Structure of the angiogenesis inhibitor ovalicin bound to its noncognate target, human Type 1 methionine aminopeptidase.
Protein Sci., 15, 2006
|
|
3B2X
 
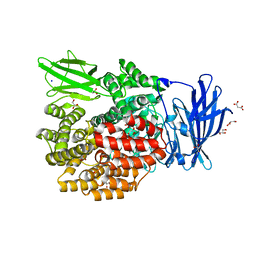 | | Crystal Structure of E. coli Aminopeptidase N in complex with Lysine | | Descriptor: | Aminopeptidase N, GLYCEROL, LYSINE, ... | | Authors: | Addlagatta, A, Gay, L, Matthews, B.W. | | Deposit date: | 2007-10-19 | | Release date: | 2008-05-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural basis for the unusual specificity of Escherichia coli aminopeptidase N.
Biochemistry, 47, 2008
|
|
3PUU
 
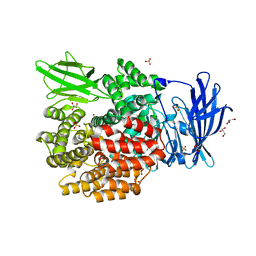 | |
3QJX
 
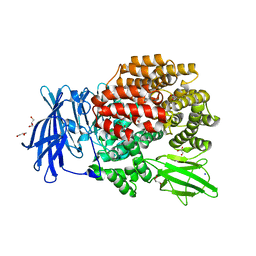 | | Crystal Structure of E. coli Aminopeptidase N in complex with L-Serine | | Descriptor: | Aminopeptidase N, GLYCEROL, MALONATE ION, ... | | Authors: | Addlagatta, A, Gumpena, R, Kishor, C, Ganji, R.J. | | Deposit date: | 2011-01-31 | | Release date: | 2011-11-16 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Discovery of alpha, beta- and alpha, gamma-Diamino Acid Scaffolds for the Inhibition of M1 Family Aminopeptidases
Chemmedchem, 6, 2011
|
|
4IKT
 
 | | Crystal structure of truncated (delta 1-89) human methionine aminopeptidase Type 1 in complex with N1-(5-chloro-6-methyl-2-(pyridin-2-yl)pyrimidin-4-yl)-N2-(5-(trifluoromethyl)pyridin-2-yl)ethane-1,2-diamine | | Descriptor: | COBALT (II) ION, GLYCEROL, Methionine aminopeptidase 1, ... | | Authors: | Addlagatta, A, Kishor, C, Arya, T. | | Deposit date: | 2012-12-28 | | Release date: | 2013-12-11 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Identification, Biochemical and Structural Evaluation of Species-Specific Inhibitors against Type I Methionine Aminopeptidases
J.Med.Chem., 56, 2013
|
|
4IKS
 
 | | Crystal structure of truncated (delta 1-89) human methionine aminopeptidase Type 1 in complex with N1-(5-chloro-6-methyl-2-(pyridin-2-yl)pyrimidin-4-yl)-N2-(6-(trifluoromethyl)pyridin-2-yl)ethane-1,2-diamine | | Descriptor: | COBALT (II) ION, GLYCEROL, Methionine aminopeptidase 1, ... | | Authors: | Addlagatta, A, Kishor, C, Arya, T. | | Deposit date: | 2012-12-28 | | Release date: | 2013-12-11 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Identification, Biochemical and Structural Evaluation of Species-Specific Inhibitors against Type I Methionine Aminopeptidases
J.Med.Chem., 56, 2013
|
|
4IKU
 
 | | Crystal structure of truncated (delta 1-89) human methionine aminopeptidase Type 1 in complex with 2-((5-chloro-6-methyl-2-(pyridin-2-yl)pyrimidin-4-yl)amino)-3-phenylpropanamide | | Descriptor: | COBALT (II) ION, Methionine aminopeptidase 1, Nalpha-[5-chloro-6-methyl-2-(pyridin-2-yl)pyrimidin-4-yl]-D-phenylalaninamide, ... | | Authors: | Addlagatta, A, Kishor, C, Arya, T. | | Deposit date: | 2012-12-28 | | Release date: | 2013-12-11 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Identification, Biochemical and Structural Evaluation of Species-Specific Inhibitors against Type I Methionine Aminopeptidases
J.Med.Chem., 56, 2013
|
|
4IKR
 
 | | Crystal structure of Type 1 human methionine aminopeptidase in complex with 2-(4-(5-chloro-6-methyl-2-(pyridin-2-yl)pyrimidin-4-yl)piperazin-1-yl)ethanol | | Descriptor: | 2-{4-[5-chloro-6-methyl-2-(pyridin-2-yl)pyrimidin-4-yl]piperazin-1-yl}ethanol, COBALT (II) ION, GLYCEROL, ... | | Authors: | Addlagatta, A, Kishor, C, Arya, T. | | Deposit date: | 2012-12-28 | | Release date: | 2013-12-11 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Identification, Biochemical and Structural Evaluation of Species-Specific Inhibitors against Type I Methionine Aminopeptidases
J.Med.Chem., 56, 2013
|
|
3B3B
 
 | |
3B37
 
 | |
3B34
 
 | |
6LH7
 
 | |
6K26
 
 | | Crystal structure of Vibrio cholerae methionine aminopeptidase | | Descriptor: | Methionine aminopeptidase, SODIUM ION | | Authors: | Pillalamarri, V, Addlagatta, A. | | Deposit date: | 2019-05-13 | | Release date: | 2020-05-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Methionine aminopeptidases with short sequence inserts within the catalytic domain are differentially inhibited: Structural and biochemical studies of three proteins from Vibrio spp.
Eur.J.Med.Chem., 209, 2020
|
|
6KSG
 
 | | Vibrio cholerae Methionine Aminopeptidase in holo form | | Descriptor: | GLYCEROL, Methionine aminopeptidase, NICKEL (II) ION, ... | | Authors: | Pillalamarri, V, Addlagatta, A. | | Deposit date: | 2019-08-23 | | Release date: | 2020-08-26 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Methionine aminopeptidases with short sequence inserts within the catalytic domain are differentially inhibited: Structural and biochemical studies of three proteins from Vibrio spp.
Eur.J.Med.Chem., 209, 2020
|
|
4KM3
 
 | |
4X5K
 
 | |
4WXK
 
 | |
