2O5T
 
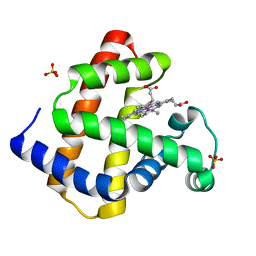 | | Cobalt horse heart myoglobin, oxidized | | Descriptor: | Myoglobin, PROTOPORPHYRIN IX CONTAINING CO, SULFATE ION | | Authors: | Richter-Addo, G.B, Zahran, Z.N, Chooback, L, Copeland, D.M, West, A.H. | | Deposit date: | 2006-12-06 | | Release date: | 2007-10-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structures of manganese- and cobalt-substituted myoglobin in complex with NO and nitrite reveal unusual ligand conformations.
J.Inorg.Biochem., 102, 2008
|
|
4FYR
 
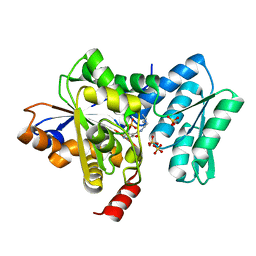
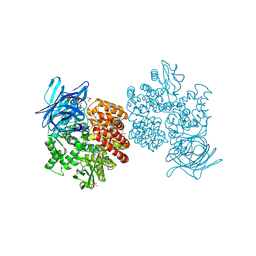 | | Human aminopeptidase N (CD13) in complex with bestatin | | Descriptor: | 2-(3-AMINO-2-HYDROXY-4-PHENYL-BUTYRYLAMINO)-4-METHYL-PENTANOIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Wong, A.H, Rini, J.M. | | Deposit date: | 2012-07-05 | | Release date: | 2012-09-05 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | The X-ray Crystal Structure of Human Aminopeptidase N Reveals a Novel Dimer and the Basis for Peptide Processing.
J.Biol.Chem., 287, 2012
|
|
2OFI
 
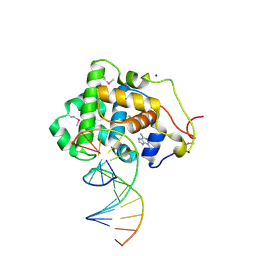 | | Crystal Structure of 3-methyladenine DNA Glycosylase I (TAG) bound to DNA/3mA | | Descriptor: | 3-METHYL-3H-PURIN-6-YLAMINE, 3-methyladenine DNA glycosylase I, constitutive, ... | | Authors: | Metz, A.H, Hollis, T, Eichman, B.F. | | Deposit date: | 2007-01-03 | | Release date: | 2007-05-15 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | DNA damage recognition and repair by 3-methyladenine DNA glycosylase I (TAG).
Embo J., 26, 2007
|
|
2I6O
 
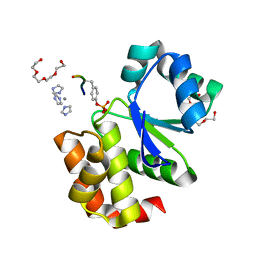 | |
2I80
 
 | | Allosteric inhibition of Staphylococcus aureus D-alanine:D-alanine ligase revealed by crystallographic studies | | Descriptor: | 3-CHLORO-2,2-DIMETHYL-N-[4-(TRIFLUOROMETHYL)PHENYL]PROPANAMIDE, D-alanine-D-alanine ligase | | Authors: | Liu, S, Chang, J.S, Herberg, J.T, Horng, M.-M, Tomich, P.K, Lin, A.H, Marotti, K.R. | | Deposit date: | 2006-08-31 | | Release date: | 2006-09-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Allosteric inhibition of Staphylococcus aureus D-alanine:D-alanine ligase revealed by crystallographic studies.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
2NSS
 
 | |
2NS3
 
 | | Solution structure of ribbon BuIA | | Descriptor: | Alpha-conotoxin BuIA | | Authors: | Jin, A.H, Brandstaetter, H, Nevin, S.T, Tan, C.C, Clark, R.J, Adams, D.J, Alewood, P.F, Craik, D.J, Daly, N.L. | | Deposit date: | 2006-11-03 | | Release date: | 2007-09-25 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Structure of alpha-conotoxin BuIA: influences of disulfide connectivity on structural dynamics
Bmc Struct.Biol., 7, 2007
|
|
2OK9
 
 | | PrTX-I-BPB | | Descriptor: | ISOPROPYL ALCOHOL, Phospholipase A2 homolog 1, p-Bromophenacyl bromide | | Authors: | Marchi-Salvador, D.P, Fernandes, C.A.H, Soares, A.M, Fontes, M.R. | | Deposit date: | 2007-01-16 | | Release date: | 2008-09-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Crystal structure of a phospholipase A(2) homolog complexed with p-bromophenacyl bromide reveals important structural changes associated with the inhibition of myotoxic activity.
Biochim.Biophys.Acta, 1794, 2009
|
|
2OFK
 
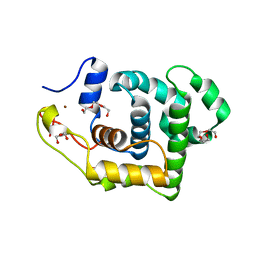 | | Crystal Structure of 3-methyladenine DNA glycosylase I (TAG) | | Descriptor: | 3-methyladenine DNA glycosylase I, constitutive, TRIETHYLENE GLYCOL, ... | | Authors: | Metz, A.H, Hollis, T, Eichman, B.F. | | Deposit date: | 2007-01-03 | | Release date: | 2007-05-15 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | DNA damage recognition and repair by 3-methyladenine DNA glycosylase I (TAG).
Embo J., 26, 2007
|
|
2O5M
 
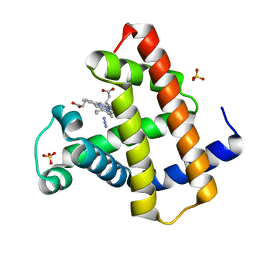 | | Manganese horse heart myoglobin, azide modified | | Descriptor: | AZIDE ION, Myoglobin, PROTOPORPHYRIN IX CONTAINING MN, ... | | Authors: | Richter-Addo, G.B, Zahran, Z.N, Chooback, L, Copeland, D.M, West, A.H. | | Deposit date: | 2006-12-06 | | Release date: | 2007-10-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structures of manganese- and cobalt-substituted myoglobin in complex with NO and nitrite reveal unusual ligand conformations.
J.Inorg.Biochem., 102, 2008
|
|
2I87
 
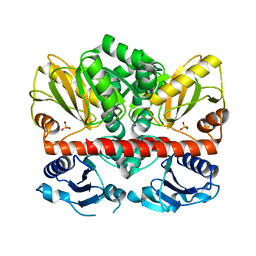 | | Allosteric inhibition of Staphylococcus aureus D-alanine:D-alanine ligase revealed by crystallographic studies | | Descriptor: | D-alanine-D-alanine ligase, SULFATE ION | | Authors: | Liu, S, Chang, J.S, Herberg, J.T, Horng, M, Tomich, P.K, Lin, A.H, Marotti, K.R. | | Deposit date: | 2006-09-01 | | Release date: | 2006-10-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Allosteric inhibition of Staphylococcus aureus D-alanine:D-alanine ligase revealed by crystallographic studies.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
2JPS
 
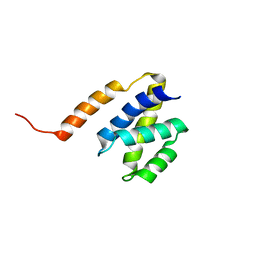 | | NAB2 N-terminal domain | | Descriptor: | Nuclear polyadenylated RNA-binding protein NAB2 | | Authors: | Grant, R, Marshall, N.J, Yang, J, Fasken, M, Kelly, S, Harreman, M.T, Neuhaus, D, Corbett, A.H, Stewart, M. | | Deposit date: | 2007-05-23 | | Release date: | 2008-03-18 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | Structure of the N-terminal Mlp1-binding domain of the Saccharomyces cerevisiae mRNA-binding protein, Nab2
J.Mol.Biol., 376, 2008
|
|
4H27
 
 | |
4FAY
 | | Crystal structure of a trimeric bacterial microcompartment shell protein PduB with glycerol metabolites | | Descriptor: | ACETATE ION, GLYCEROL, Microcompartments protein | | Authors: | Pang, A.H, Prentice, M.B, Pickersgill, R.W. | | Deposit date: | 2012-05-22 | | Release date: | 2012-11-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Substrate channels revealed in the trimeric Lactobacillus reuteri bacterial microcompartment shell protein PduB.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4F2O
 
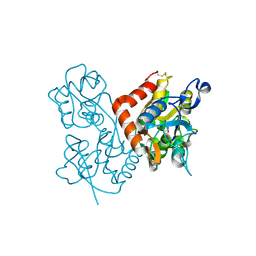 | | Quisqualate bound to the D655A mutant of the ligand binding domain of GluA3 | | Descriptor: | (S)-2-AMINO-3-(3,5-DIOXO-[1,2,4]OXADIAZOLIDIN-2-YL)-PROPIONIC ACID, Glutamate receptor 3, ZINC ION | | Authors: | Ahmed, A.H, Oswald, R.E. | | Deposit date: | 2012-05-08 | | Release date: | 2012-05-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.912 Å) | | Cite: | The loss of an electrostatic contact unique to AMPA receptor ligand binding domain 2 slows channel activation.
Biochemistry, 51, 2012
|
|
4F3G
 
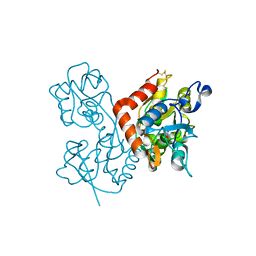 | | Kainate bound to the ligand binding domain of GluA3i | | Descriptor: | 3-(CARBOXYMETHYL)-4-ISOPROPENYLPROLINE, Glutamate receptor 3, ZINC ION | | Authors: | Ahmed, A.H, Oswald, R.E. | | Deposit date: | 2012-05-09 | | Release date: | 2012-05-30 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.064 Å) | | Cite: | The loss of an electrostatic contact unique to AMPA receptor ligand binding domain 2 slows channel activation.
Biochemistry, 51, 2012
|
|
4FA8
 
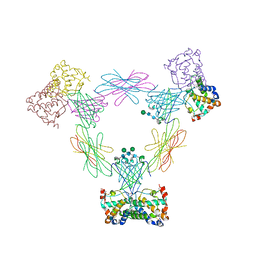 | | Multi-pronged modulation of cytokine signaling | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Macrophage colony-stimulating factor 1, Secreted protein BARF1, ... | | Authors: | He, X, Shim, A.H. | | Deposit date: | 2012-05-21 | | Release date: | 2012-08-29 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Multipronged attenuation of macrophage-colony stimulating factor signaling by Epstein-Barr virus BARF1.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4FM7
 
 | | Crystal Structure of BACE with Compound 14g | | Descriptor: | 4-{[(5R,7S)-1-(3-fluorophenyl)-3,7-dimethyl-2,2-dioxido-2-thia-1,3,8-triazaspiro[4.5]dec-8-yl]methyl}-2-(propan-2-yloxy)phenol, Beta-secretase 1, ZINC ION | | Authors: | Vajdos, F.F, Varghese, A.H. | | Deposit date: | 2012-06-15 | | Release date: | 2012-10-03 | | Last modified: | 2013-06-19 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Spirocyclic sulfamides as beta-secretase 1 (BACE-1) inhibitors for the treatment of Alzheimer's disease: utilization of structure based drug design, WaterMap, and CNS penetration studies to identify centrally efficacious inhibitors.
J.Med.Chem., 55, 2012
|
|
4GXS
 
 | | Ligand binding domain of GluA2 (AMPA/glutamate receptor) bound to (-)-kaitocephalin | | Descriptor: | (5R)-2-[(1S,2R)-2-amino-2-carboxy-1-hydroxyethyl]-5-{(2S)-2-carboxy-2-[(3,5-dichloro-4-hydroxybenzoyl)amino]ethyl}-L-proline, Glutamate receptor 2, ZINC ION | | Authors: | Ahmed, A.H, Oswald, R.E. | | Deposit date: | 2012-09-04 | | Release date: | 2012-10-17 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9634 Å) | | Cite: | The structure of (-)-kaitocephalin bound to the ligand binding domain of the (S)-alpha-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA)/glutamate receptor, GluA2.
J.Biol.Chem., 287, 2012
|
|
2K79
 
 | |
4FM8
 
 | | Crystal Structure of BACE with Compound 12a | | Descriptor: | (5R,7S)-1-(3-fluorophenyl)-3,7-dimethyl-8-[3-(propan-2-yloxy)benzyl]-2-thia-1,3,8-triazaspiro[4.5]decane 2,2-dioxide, 1,2-ETHANEDIOL, Beta-secretase 1, ... | | Authors: | Vajdos, F.F, Varghese, A.H. | | Deposit date: | 2012-06-15 | | Release date: | 2012-10-03 | | Last modified: | 2013-06-19 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Spirocyclic sulfamides as beta-secretase 1 (BACE-1) inhibitors for the treatment of Alzheimer's disease: utilization of structure based drug design, WaterMap, and CNS penetration studies to identify centrally efficacious inhibitors.
J.Med.Chem., 55, 2012
|
|
2KIC
 
 | | n-NafY. N-terminal domain of NafY | | Descriptor: | Nitrogenase gamma subunit | | Authors: | Phillips, A.H, Hernandez, J.A, Erbil, K, Pelton, J.G, Wemmer, D.E, Rubio, L.M. | | Deposit date: | 2009-05-01 | | Release date: | 2010-12-08 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Biological activity and solution structure of the apo-dinitrogenase binding domain of NafY
J.Biol.Chem., 2010
|
|
2K6A
 
 | | Solution structure of EAS D15 truncation mutant | | Descriptor: | Hydrophobin | | Authors: | Kwan, A.H. | | Deposit date: | 2008-07-07 | | Release date: | 2008-08-19 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | The Cys3-Cys4 loop of the hydrophobin EAS is not required for rodlet formation and surface activity.
J.Mol.Biol., 382, 2008
|
|
2K71
 
 | | Structure and dynamics of a DNA GNRA hairpin solved vy high-sensitivity NMR with two independent converging methods, simulated annealing (DYANA) and mesoscopic molecular modelling (BCE/AMBER) | | Descriptor: | 5'-D(*DGP*DCP*DGP*DAP*DAP*DAP*DGP*DC)-3' | | Authors: | Santini, G.P.H, Cognet, J.A.H, Xu, D, Singarapu, K.K, Herve du Penhoat, C.L.M. | | Deposit date: | 2008-07-29 | | Release date: | 2009-07-21 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Nucleic acid folding determined by mesoscale modeling and NMR spectroscopy: solution structure of d(GCGAAAGC).
J.Phys.Chem.B, 113, 2009
|
|
2I6M
 
 | |
