2J1R
 
 | |
1TCR
 
 | | MURINE T-CELL ANTIGEN RECEPTOR 2C CLONE | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Garcia, K.C, Degano, M, Stanfield, R.L, Wilson, I.A. | | Deposit date: | 1996-09-12 | | Release date: | 1997-03-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | An alphabeta T cell receptor structure at 2.5 A and its orientation in the TCR-MHC complex.
Science, 274, 1996
|
|
8IK4
 
 | |
5KW7
 
 | | T. danielli thaumatin at 278K, Data set 4 | | Descriptor: | L(+)-TARTARIC ACID, Thaumatin-1 | | Authors: | Russi, S, Gonzalez, A, Kenner, L.R, Keedy, D.A, Fraser, J.S. | | Deposit date: | 2016-07-15 | | Release date: | 2016-08-10 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Conformational variation of proteins at room temperature is not dominated by radiation damage.
J Synchrotron Radiat, 24, 2017
|
|
7T7D
 
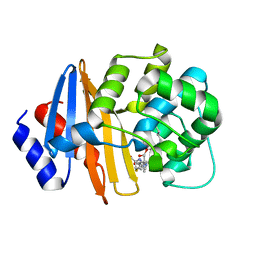 | | MA-1-206-OXA-23 30s complex | | Descriptor: | (2R,4S)-2-(1,3-dihydroxypropan-2-yl)-4-{[(3R,5R)-5-(dimethylcarbamoyl)pyrrolidin-3-yl]sulfanyl}-3,4-dihydro-2H-pyrrole-5-carboxylic acid, Beta-lactamase OXA-23 | | Authors: | Smith, C.A, Stewart, N.K, Vakulenko, S.B. | | Deposit date: | 2021-12-15 | | Release date: | 2022-05-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | C6 Hydroxymethyl-Substituted Carbapenem MA-1-206 Inhibits the Major Acinetobacter baumannii Carbapenemase OXA-23 by Impeding Deacylation.
Mbio, 13, 2022
|
|
7T7E
 
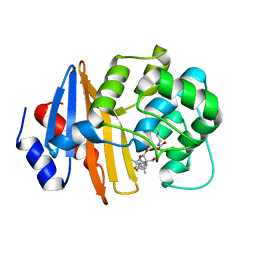 | | MA-1-206-OXA-23 3 minute complex | | Descriptor: | (2R,4S)-2-(1,3-dihydroxypropan-2-yl)-4-{[(3R,5R)-5-(dimethylcarbamoyl)pyrrolidin-3-yl]sulfanyl}-3,4-dihydro-2H-pyrrole-5-carboxylic acid, Beta-lactamase OXA-23 | | Authors: | Smith, C.A, Stewart, N.K, Vakulenko, S.B. | | Deposit date: | 2021-12-15 | | Release date: | 2022-05-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | C6 Hydroxymethyl-Substituted Carbapenem MA-1-206 Inhibits the Major Acinetobacter baumannii Carbapenemase OXA-23 by Impeding Deacylation.
Mbio, 13, 2022
|
|
3EMW
 
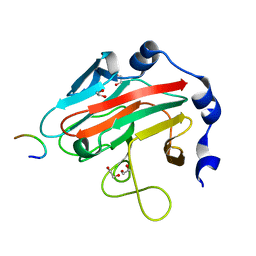 | | Crystal Structure of human splA/ryanodine receptor domain and SOCS box containing 2 (SPSB2) in complex with a 20-residue VASA peptide | | Descriptor: | 1,2-ETHANEDIOL, Peptide (VASA), SPRY domain-containing SOCS box protein 2 | | Authors: | Filippakopoulos, P, Sharpe, T, Keates, T, Murray, J.W, Savitsky, P, Roos, A.K, Pike, A.C.W, Von Delft, F, Arrowsmith, C.H, Edwards, A.M, Weigelt, J, Bountra, C, Knapp, S, Bullock, A, Structural Genomics Consortium (SGC) | | Deposit date: | 2008-09-25 | | Release date: | 2008-10-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for Par-4 recognition by the SPRY domain- and SOCS box-containing proteins SPSB1, SPSB2, and SPSB4.
J.Mol.Biol., 401, 2010
|
|
1THU
 
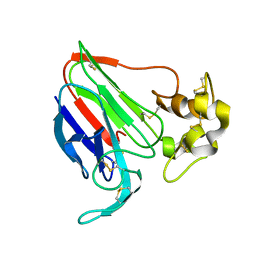 | | THE STRUCTURES OF THREE CRYSTAL FORMS OF THE SWEET PROTEIN THAUMATIN | | Descriptor: | THAUMATIN ISOFORM B | | Authors: | Ko, T.-P, Day, J, Greenwood, A, McPherson, A. | | Deposit date: | 1994-06-10 | | Release date: | 1994-12-20 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structures of three crystal forms of the sweet protein thaumatin.
Acta Crystallogr.,Sect.D, 50, 1994
|
|
4RI1
 
 | |
3ET6
 
 | | The crystal structure of the catalytic domain of a eukaryotic guanylate cyclase | | Descriptor: | PHOSPHATE ION, Soluble guanylyl cyclase beta | | Authors: | Winger, J.A, Derbyshire, E.R, Lamers, M.H, Marletta, M.A, Kuriyan, J. | | Deposit date: | 2008-10-07 | | Release date: | 2008-10-14 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | The crystal structure of the catalytic domain of a eukaryotic guanylate cyclase.
Bmc Struct.Biol., 8, 2008
|
|
8IK8
 
 | |
2WEK
 | | Crystal structure of the human MGC45594 gene product in complex with diclofenac | | Descriptor: | 2-[2,6-DICHLOROPHENYL)AMINO]BENZENEACETIC ACID, CHLORIDE ION, GLYCEROL, ... | | Authors: | Shafqat, N, Yue, W.W, Ugochukwu, E, Niesen, F, Smee, C, Arrowsmith, C, Weigelt, J, Edwards, A, Bountra, C, Oppermann, U. | | Deposit date: | 2009-03-31 | | Release date: | 2009-12-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of the Human Mgc45594 Gene Product in Complex with Diclofenac
To be Published
|
|
7T7K
 
 | | Structure of SPAC806.04c protein from fission yeast bound to Co2+ | | Descriptor: | CHLORIDE ION, COBALT (II) ION, Damage-control phosphatase SPAC806.04c, ... | | Authors: | Jacewicz, A, Sanchez, A.M, Shuman, S. | | Deposit date: | 2021-12-15 | | Release date: | 2022-06-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Fission yeast Duf89 and Duf8901 are cobalt/nickel-dependent phosphatase-pyrophosphatases that act via a covalent aspartyl-phosphate intermediate.
J.Biol.Chem., 298, 2022
|
|
3EVC
 
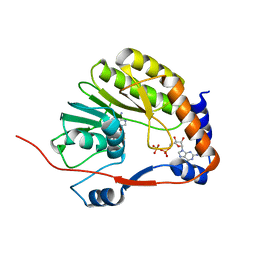 | |
1SY7
 
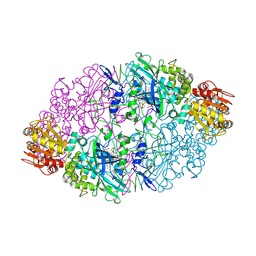 | | Crystal structure of the catalase-1 from Neurospora crassa, native structure at 1.75A resolution. | | Descriptor: | CIS-HEME D HYDROXYCHLORIN GAMMA-SPIROLACTONE, Catalase 1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Diaz, A, Horjales, E, Rudino-Pinera, E, Arreola, R, Hansberg, W. | | Deposit date: | 2004-04-01 | | Release date: | 2004-10-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Unusual Cys-Tyr covalent bond in a large catalase
J.Mol.Biol., 342, 2004
|
|
2WCG
 
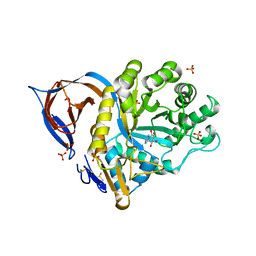 | | X-ray structure of acid-beta-glucosidase with N-octyl(cyclic guanidine)-nojirimycin in the active site | | Descriptor: | CHLORIDE ION, GLUCOSYLCERAMIDASE, N-[(3E,5R,6R,7S,8R,8AR)-5,6,7,8-TETRAHYDROXYHEXAHYDROIMIDAZO[1,5-A]PYRIDIN-3(2H)-YLIDENE]OCTAN-1-AMINIUM, ... | | Authors: | Brumshtein, B, Aguilar, M, Garcia-Moreno, M.I, Mellet, C.O, Garcia-Fernandez, J.M, Silman, I, Shaaltiel, Y, Aviezer, D, Sussman, J.L, Futerman, A.H. | | Deposit date: | 2009-03-12 | | Release date: | 2009-11-24 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | 6-Amino-6-Deoxy-5,6-Di-N-(N'-Octyliminomethylidene)Nojirimycin: Synthesis, Biological Evaluation, and Crystal Structure in Complex with Acid Beta-Glucosidase.
Chembiochem, 10, 2009
|
|
8A4D
 
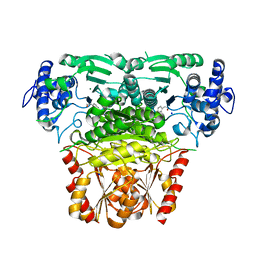 | |
1SZG
 
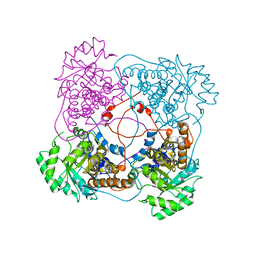 | | A198G:L230A flavocytochrome b2 with sulfite bound | | Descriptor: | Cytochrome b2, mitochondrial, N-SULFO-FLAVIN MONONUCLEOTIDE | | Authors: | Mowat, C.G, Wehenkel, A, Green, A.J, Walkinshaw, M.D, Reid, G.A, Chapman, S.K. | | Deposit date: | 2004-04-05 | | Release date: | 2004-07-27 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Altered Substrate Specificity in Flavocytochrome b(2): Structural Insights into the Mechanism of l-Lactate Dehydrogenation
Biochemistry, 43, 2004
|
|
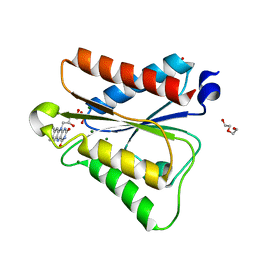
6PUQ
 
 | |
4RJD
 
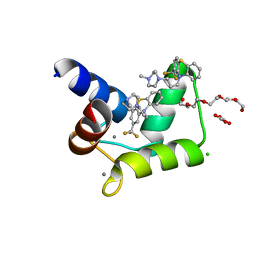 | | TFP bound in alternate orientations to calcium-saturated Calmodulin C-Domains | | Descriptor: | 10-[3-(4-METHYL-PIPERAZIN-1-YL)-PROPYL]-2-TRIFLUOROMETHYL-10H-PHENOTHIAZINE, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Feldkamp, M.D, Gakhar, L, Pandey, N, Shea, M.A. | | Deposit date: | 2014-10-08 | | Release date: | 2015-08-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Opposing orientations of the anti-psychotic drug trifluoperazine selected by alternate conformations of M144 in calmodulin.
Proteins, 83, 2015
|
|
3EQV
 
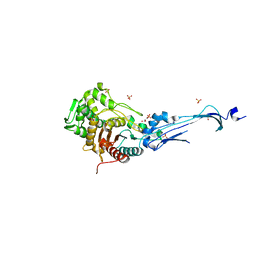 | | Crystal structure of penicillin-binding protein 2 from Neisseria gonorrhoeae containing four mutations associated with penicillin resistance | | Descriptor: | GLYCEROL, Penicillin-binding protein 2, SULFATE ION | | Authors: | Powell, A.J, Deacon, A.M, Nicholas, R.A, Davies, C. | | Deposit date: | 2008-10-01 | | Release date: | 2008-10-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structures of Penicillin-binding Protein 2 from Penicillin-susceptible and -resistant Strains of Neisseria gonorrhoeae Reveal an Unexpectedly Subtle Mechanism for Antibiotic Resistance.
J.Biol.Chem., 284, 2009
|
|
3EVA
 
 | |
6WG1
 
 | |
6PXY
 
 | |
8A6O
 
 | |
