5HBJ
 
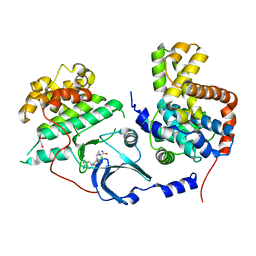 | | CDK8-CYCC IN COMPLEX WITH 8-[2-Amino-3-chloro-5-(1-methyl-1H-indazol-5-yl)-pyridin-4-yl]-2,8-diaza-spiro[4.5]decan-1-one | | Descriptor: | 1,2-ETHANEDIOL, 8-[2-azanyl-3-chloranyl-5-(1-methylindazol-5-yl)pyridin-4-yl]-2,8-diazaspiro[4.5]decan-1-one, Cyclin-C, ... | | Authors: | Musil, D, Blagg, J, Mallinger, A. | | Deposit date: | 2015-12-31 | | Release date: | 2016-02-03 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Discovery of Potent, Selective, and Orally Bioavailable Small-Molecule Modulators of the Mediator Complex-Associated Kinases CDK8 and CDK19.
J.Med.Chem., 59, 2016
|
|
4S3R
 
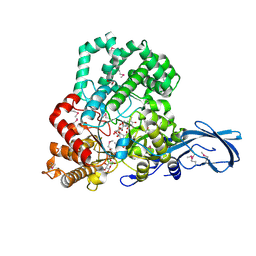 | |
5HCT
 
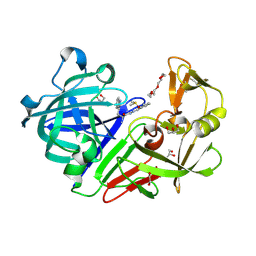 | | Endothiapepsin in complex with biacylhydrazone | | Descriptor: | 1,2-ETHANEDIOL, 2-amino-N'-{3-[(E)-{2-[(2S)-2-amino-3-(1H-indol-3-yl)propanoyl]hydrazinylidene}methyl]benzylidene}-3-(1H-indol-2-yl)propanehydrazide (non-preferred name), ACETATE ION, ... | | Authors: | Radeva, N, Heine, A, Klebe, G. | | Deposit date: | 2016-01-04 | | Release date: | 2016-07-20 | | Method: | X-RAY DIFFRACTION (1.36 Å) | | Cite: | Fragment Linking and Optimization of Inhibitors of the Aspartic Protease Endothiapepsin: Fragment-Based Drug Design Facilitated by Dynamic Combinatorial Chemistry.
Angew.Chem.Int.Ed.Engl., 55, 2016
|
|
6YTS
 
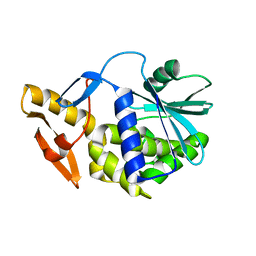 | | Solution NMR structure of type-I ribosome-inactivating protein trichobakin (TBK) | | Descriptor: | Trichobakin | | Authors: | Britikov, V.V, Bocharov, E.V, Britikova, E.V, Le, T.B.T, Phan, C.V, Boyko, K.M, Arseniev, A.S, Usanov, S.A. | | Deposit date: | 2020-04-24 | | Release date: | 2020-05-06 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Structure and dynamics of type-I ribosome-inactivating protein trichobakin (TBK)
To Be Published
|
|
7RZU
 | |
6Z0G
 
 | | Structure of TREM2 transmembrane helix in DPC micelles | | Descriptor: | Triggering receptor expressed on myeloid cells 2 | | Authors: | Steiner, A, Schlepkow, K, Brunner, B, Steiner, H, Haass, C, Hagn, F. | | Deposit date: | 2020-05-08 | | Release date: | 2020-09-16 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | gamma-Secretase cleavage of the Alzheimer risk factor TREM2 is determined by its intrinsic structural dynamics.
Embo J., 39, 2020
|
|
7RZS
 | |
7RZR
 | |
6Y02
 
 | | Thrombin in complex with 13k | | Descriptor: | (2~{S})-1-[(2~{R})-2-azanyl-3-phenyl-propanoyl]-~{N}-[(5-bromanylfuran-2-yl)methyl]pyrrolidine-2-carboxamide, 2-acetamido-2-deoxy-beta-D-glucopyranose, DIMETHYL SULFOXIDE, ... | | Authors: | Sandner, A, Heine, A, Klebe, G, Abazi, N. | | Deposit date: | 2020-02-06 | | Release date: | 2021-02-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Thrombin in complex with 13k
To be published
|
|
8ATH
 | | CRYSTAL STRUCTURE OF LAMP1 IN COMPLEX WITH FAB-B. | | Descriptor: | Fab B Heavy Chain, Fab B Light Chain, Lysosome-associated membrane glycoprotein 1 | | Authors: | Mathieu, M, Dupuy, A. | | Deposit date: | 2022-08-23 | | Release date: | 2023-03-01 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.366 Å) | | Cite: | Deciphering cross-species reactivity of LAMP-1 antibodies using deep mutational epitope mapping and AlphaFold.
Mabs, 15, 2023
|
|
7BOD
 
 | | Bacterial 30S ribosomal subunit assembly complex state M (body domain) | | Descriptor: | 16S rRNA (body domain of 30S subunit), 30S ribosomal protein S11, 30S ribosomal protein S12, ... | | Authors: | Schedlbauer, A, Iturrioz, I, Ochoa-Lizarralde, B, Diercks, T, Lopez-Alonso, J, Kaminishi, T, Capuni, R, Astigarraga, E, Gil-Carton, D, Fucini, P, Connell, S.R. | | Deposit date: | 2021-01-25 | | Release date: | 2021-07-07 | | Last modified: | 2024-04-24 | | Method: | ELECTRON MICROSCOPY (2.88 Å) | | Cite: | A conserved rRNA switch is central to decoding site maturation on the small ribosomal subunit.
Sci Adv, 7, 2021
|
|
6Z0I
 | | Structure of the TREM2 transmembrane helix in complex with DAP12 in DPC micelles | | Descriptor: | Triggering receptor expressed on myeloid cells 2 | | Authors: | Steiner, A, Schlepkow, K, Brunner, B, Steiner, H, Haass, C, Hagn, F. | | Deposit date: | 2020-05-08 | | Release date: | 2020-09-16 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | gamma-Secretase cleavage of the Alzheimer risk factor TREM2 is determined by its intrinsic structural dynamics.
Embo J., 2020
|
|
7RZT
 | |
6Z0H
 
 | | Structure of TREM2 transmembrane helix K186A variant in DPC micelles | | Descriptor: | Triggering receptor expressed on myeloid cells 2 | | Authors: | Steiner, A, Schlepkow, K, Brunner, B, Steiner, H, Haass, C, Hagn, F. | | Deposit date: | 2020-05-08 | | Release date: | 2020-09-16 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | gamma-Secretase cleavage of the Alzheimer risk factor TREM2 is determined by its intrinsic structural dynamics.
Embo J., 39, 2020
|
|
6FVE
 
 | |
6APK
 
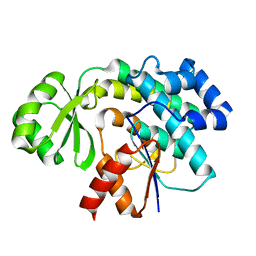 | | Trans-acting transferase from Disorazole synthase solved by serial femtosecond XFEL crystallography | | Descriptor: | DisD protein | | Authors: | Lyubimov, A.Y, Mathews, I.I, Uervivojnangkoorn, M, Khosla, C, Soltis, S.M, Cohen, A.E. | | Deposit date: | 2017-08-17 | | Release date: | 2018-04-04 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The Conformational Flexibility of the Acyltransferase from the Disorazole Polyketide Synthase Is Revealed by an X-ray Free-Electron Laser Using a Room-Temperature Sample Delivery Method for Serial Crystallography.
Biochemistry, 56, 2017
|
|
7RZV
 | |
8SHR
 
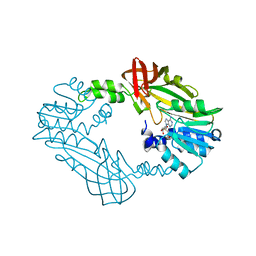 | | Crystal Structure of PRMT3 with Compound YD1-214 | | Descriptor: | 5'-S-[2-(phenylcarbamamido)ethyl]-5'-thioadenosine, Protein arginine N-methyltransferase 3 | | Authors: | Song, X, Dong, A, Deng, Y, Huang, R, Arrowsmith, C.H, Edwards, A.M, Min, J, Structural Genomics Consortium (SGC) | | Deposit date: | 2023-04-14 | | Release date: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Crystal Structure of PRMT3 with Compound YD1-214
To be published
|
|
7RZQ
 | |
7N4F
 
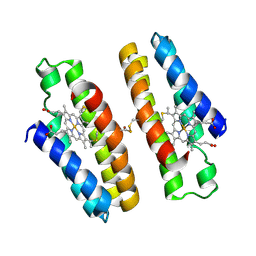 | |
5HEM
 
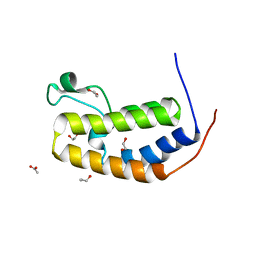 | | Crystal structure of the N-terminus D161Y bromodomain mutant of human BRD2 | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Bromodomain-containing protein 2, ... | | Authors: | Tallant, C, Lori, C, Pasquo, A, Chiaraluce, R, Consalvi, V, Newman, J.A, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Knapp, S. | | Deposit date: | 2016-01-06 | | Release date: | 2016-01-20 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structure of the N-terminus D161Y bromodomain mutant of human BRD2
To Be Published
|
|
6UEK
 
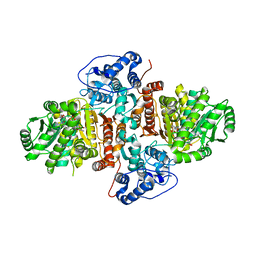 | | Structure of Urocanate Hydratase from Trypanosoma cruzi in complex with NAD+ | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Urocanate hydratase | | Authors: | Boreiko, S, Silva, M, Melo, R.F.P, Silber, A.M, Iulek, J. | | Deposit date: | 2019-09-21 | | Release date: | 2020-01-15 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Structure of Urocanate Hydratase from the protozoan Trypanosoma cruzi.
Int.J.Biol.Macromol., 146, 2019
|
|
7N4G
 
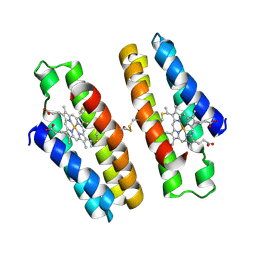 | |
7VJY
 
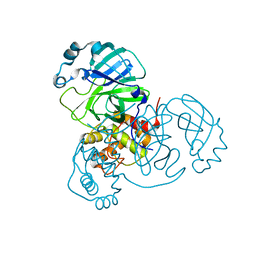 | |
5IAK
 
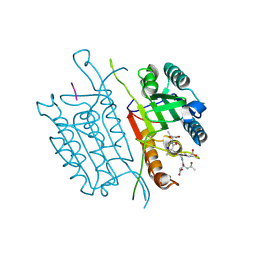 | | Caspase 3 V266S | | Descriptor: | ACE-ASP-GLU-VAL-ASK, Caspase-3, SODIUM ION | | Authors: | Maciag, J.J, Mackenzie, S.H, Tucker, M.B, Schipper, J.L, Swartz, P.D, Clark, A.C. | | Deposit date: | 2016-02-21 | | Release date: | 2016-10-26 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Tunable allosteric library of caspase-3 identifies coupling between conserved water molecules and conformational selection.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
