1TD7
 
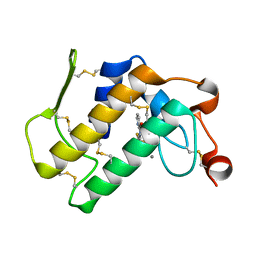 | | Interactions of a specific non-steroidal anti-inflammatory drug (NSAID) with group I phospholipase A2 (PLA2): Crystal structure of the complex formed between PLA2 and niflumic acid at 2.5 A resolution | | Descriptor: | 2-{[3-(TRIFLUOROMETHYL)PHENYL]AMINO}NICOTINIC ACID, CALCIUM ION, Phospholipase A2 isoform 3 | | Authors: | Jabeen, T, Singh, N, Singh, R.K, Sharma, S, Perbandt, M, Betzel, C, Singh, T.P. | | Deposit date: | 2004-05-21 | | Release date: | 2004-06-08 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Non-steroidal anti-inflammatory drugs as potent inhibitors of phospholipase A2: structure of the complex of phospholipase A2 with niflumic acid at 2.5 Angstroms resolution.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
1S6B
 
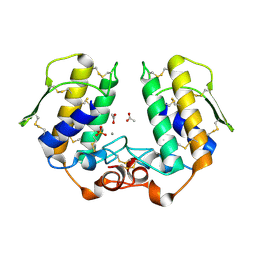 | | X-ray Crystal Structure of a Complex Formed Between Two Homologous Isoforms of Phospholipase A2 from Naja naja sagittifera: Principle of Molecular Association and Inactivation | | Descriptor: | ACETIC ACID, CALCIUM ION, PHOSPHATE ION, ... | | Authors: | Jabeen, T, Sharma, S, Singh, R.K, Kaur, P, Singh, T.P. | | Deposit date: | 2004-01-23 | | Release date: | 2004-02-10 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of a calcium-induced dimer of two isoforms of cobra phospholipase A2 at 1.6 A resolution.
Proteins, 59, 2005
|
|
2X3A
 
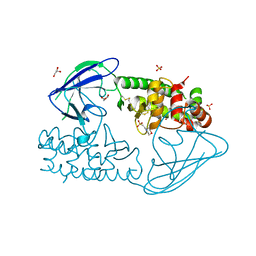 | | AsaP1 inactive mutant E294Q, an extracellular toxic zinc metalloendopeptidase | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, GLYCEROL, SULFATE ION, ... | | Authors: | Bogdanovic, X, Palm, G.J, Singh, R.K, Hinrichs, W. | | Deposit date: | 2010-01-22 | | Release date: | 2011-02-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Evidence of Intramolecular Propeptide Inhibition of the Aspzincin Metalloendopeptidase Asap1.
FEBS Lett., 590, 2016
|
|
2X3B
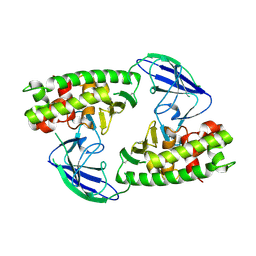 
 | | AsaP1 inactive mutant E294A, an extracellular toxic zinc metalloendopeptidase | | Descriptor: | TOXIC EXTRACELLULAR ENDOPEPTIDASE, ZINC ION | | Authors: | Bogdanovic, X, Palm, G.J, Singh, R.K, Hinrichs, W. | | Deposit date: | 2010-01-22 | | Release date: | 2011-02-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Structural Evidence of Intramolecular Propeptide Inhibition of the Aspzincin Metalloendopeptidase Asap1.
FEBS Lett., 590, 2016
|
|
2X3C
 
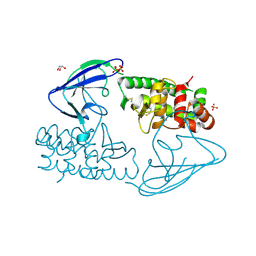 | | AsaP1 inactive mutant E294Q, an extracellular toxic zinc metalloendopeptidase | | Descriptor: | CHLORIDE ION, GLYCEROL, SULFATE ION, ... | | Authors: | Bogdanovic, X, Palm, G.J, Singh, R.K, Hinrichs, W. | | Deposit date: | 2010-01-22 | | Release date: | 2011-02-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Structural Evidence of Intramolecular Propeptide Inhibition of the Aspzincin Metalloendopeptidase Asap1.
FEBS Lett., 590, 2016
|
|
2Y40
 
 | | Structure of Isopropylmalate dehydrogenase from Thermus thermophilus - complex with Mn | | Descriptor: | 3-ISOPROPYLMALATE DEHYDROGENASE, MANGANESE (II) ION | | Authors: | Graczer, E, merlin, A, Singh, R.K, Manikandan, K, Zavodsky, P, Weiss, M.S, Vas, M. | | Deposit date: | 2011-01-04 | | Release date: | 2011-01-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Atomic Level Description of the Domain Closure in a Dimeric Enzyme: Thermus Thermophilus 3-Isopropylmalate Dehydrogenase.
Mol.Biosyst., 7, 2011
|
|
2Y42
 
 | | Structure of Isopropylmalate dehydrogenase from Thermus thermophilus - complex with NADH and Mn | | Descriptor: | 3-ISOPROPYLMALATE DEHYDROGENASE, BICINE, MANGANESE (II) ION, ... | | Authors: | Graczer, E, merlin, A, Singh, R.K, Manikandan, K, Zavodsky, P, Weiss, M.S, Vas, M. | | Deposit date: | 2011-01-04 | | Release date: | 2011-01-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Atomic Level Description of the Domain Closure in a Dimeric Enzyme: Thermus Thermophilus 3-Isopropylmalate Dehydrogenase.
Mol.Biosyst., 7, 2011
|
|
2Y3Z
 
 | | Structure of Isopropylmalate dehydrogenase from Thermus thermophilus - apo enzyme | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 3-ISOPROPYLMALATE DEHYDROGENASE, GLYCEROL, ... | | Authors: | Graczer, E, merlin, A, Singh, R.K, Manikandan, K, Zavodsky, P, Weiss, M.S, Vas, M. | | Deposit date: | 2011-01-04 | | Release date: | 2011-01-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Atomic Level Description of the Domain Closure in a Dimeric Enzyme: Thermus Thermophilus 3-Isopropylmalate Dehydrogenase.
Mol.Biosyst., 7, 2011
|
|
2Y41
 
 | | Structure of Isopropylmalate dehydrogenase from Thermus thermophilus - complex with IPM and MN | | Descriptor: | 3-ISOPROPYLMALATE DEHYDROGENASE, 3-ISOPROPYLMALIC ACID, MANGANESE (II) ION | | Authors: | Graczer, E, merlin, A, Singh, R.K, Manikandan, K, Zavodsky, P, Weiss, M.S, Vas, M. | | Deposit date: | 2011-01-04 | | Release date: | 2011-01-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Atomic Level Description of the Domain Closure in a Dimeric Enzyme: Thermus Thermophilus 3-Isopropylmalate Dehydrogenase.
Mol.Biosyst., 7, 2011
|
|
1OWS
 
 | | Crystal structure of a C49 Phospholipase A2 from Indian cobra reveals carbohydrate binding in the hydrophobic channel | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Phospholipase A2, ZINC ION | | Authors: | Jabeen, T, Jasti, J, Singh, N, Singh, R.K, Sharma, S, Kaur, P, Singh, T.P. | | Deposit date: | 2003-03-30 | | Release date: | 2003-05-20 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of a C49 Phospholipase A2 from Indian cobra reveals carbohydrate binding in the hydrophobic channel
To be Published
|
|
1YXH
 
 | | Crystal structure of a novel phospholipase A2 from Naja naja sagittifera with a strong anticoagulant activity | | Descriptor: | CALCIUM ION, ETHANOL, PHOSPHATE ION, ... | | Authors: | Jabeen, T, Singh, N, Singh, R.K, Sharma, S, Srinivasan, A, Singh, T.P. | | Deposit date: | 2005-02-21 | | Release date: | 2005-05-03 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Crystal structure of a novel phospholipase A(2) from Naja naja sagittifera with a strong anticoagulant activity
Toxicon, 46, 2005
|
|
1XXW
 
 | | Structure of zinc induced heterodimer of two calcium free isoforms of phospholipase A2 from Naja naja sagittifera at 2.7A resolution | | Descriptor: | ACETIC ACID, Phospholipase A2 isoform 1, Phospholipase A2 isoform 2, ... | | Authors: | Jabeen, T, Sharma, S, Singh, N, Singh, R.K, Verma, A.K, Paramasivam, M, Srinivasan, A, Singh, T.P. | | Deposit date: | 2004-11-09 | | Release date: | 2005-03-15 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of the zinc-induced heterodimer of two calcium-free isoforms of phospholipase A2 from Naja naja sagittifera at 2.7 angstroms resolution.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
1Y75
 
 | | A new form of catalytically inactive phospholipase A2 with an unusual disulphide bridge Cys 32- Cys 49 reveals recognition for N-acetylglucosmine | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ZINC ION, phospholipase A2 isoform 5, ... | | Authors: | Jabeen, T, Singh, N, Jasti, J, Singh, R.K, Sharma, S, Perbandt, M, Betzel, C, Kaur, P, Srinivasan, A, Singh, T.P. | | Deposit date: | 2004-12-08 | | Release date: | 2005-05-03 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of a heterodimer of phospholipase A2 from Naja naja sagittifera at 2.3 A resolution reveals the presence of a new PLA2-like protein with a novel cys 32-Cys 49 disulphide bridge with a bound sugar at the substrate-binding site
Proteins, 62, 2006
|
|
1MH2
 
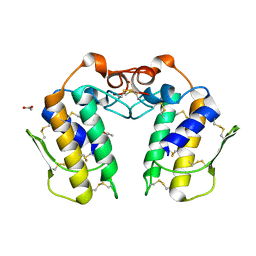 | | Crystal Structure of a Zinc Containing Dimer of Phospholipase A2 from the Venom of Indian Cobra (Naja Naja Sagittifera) | | Descriptor: | ACETIC ACID, PHOSPHOLIPASE A2, ZINC ION | | Authors: | Jabeen, T, Varma, A.K, Paramasivam, M, Singh, N, Singh, R.K, Sharma, S, Srinivasan, A, Singh, T.P. | | Deposit date: | 2002-08-19 | | Release date: | 2003-05-20 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structure of a Zinc Containing Dimer of Phospholipase A2 from the Venom of Indian cobra (Naja Naja Saggittifera)
To be Published
|
|
1MH8
 
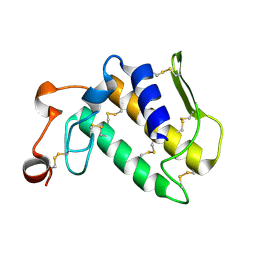 | | Crystal Structure of a Phopholipase A2 Monomer with Isoleucine at Second Position | | Descriptor: | PHOSPHOLIPASE A2 | | Authors: | Jabeen, T, Jasti, J, Singh, N, Singh, R.K, Sharma, S, Singh, T.P. | | Deposit date: | 2002-08-19 | | Release date: | 2003-06-10 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Crystal Structure of a Phospholipase A2 Monomer with Isoleucine at Second Position
To be Published
|
|
5M04
 
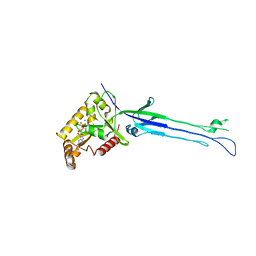 | | Structure of ObgE from Escherichia coli | | Descriptor: | GTPase ObgE/CgtA, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION | | Authors: | Gkekas, S, Singh, R.K, Versees, W. | | Deposit date: | 2016-10-03 | | Release date: | 2017-03-01 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural and biochemical analysis of Escherichia coli ObgE, a central regulator of bacterial persistence.
J. Biol. Chem., 292, 2017
|
|
1MH7
 
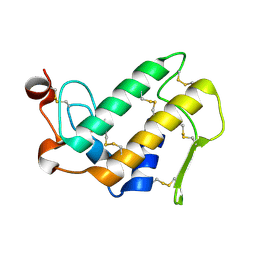 | |
7PU1
 
 | | High resolution X-ray structure of Thermoascus aurantiacus LPMO | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, COPPER (II) ION, ... | | Authors: | Banerjee, S, Frandsen, K.E.H, Singh, R.K, Bjerrum, M.J, Lo Leggio, L. | | Deposit date: | 2021-09-28 | | Release date: | 2022-03-16 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.06 Å) | | Cite: | Protonation State of an Important Histidine from High Resolution Structures of Lytic Polysaccharide Monooxygenases.
Biomolecules, 12, 2022
|
|
3V9I
 
 | |
3V9H
 
 | |
5W46
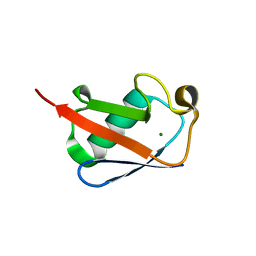 
 | |
4K57
 
 | |
6ZGQ
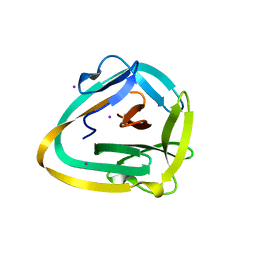 
 | | AceL NrdHF class 3 split intein GSH linked splice inactive variant - C124A, N146A | | Descriptor: | AceL NrdHF-1-1 Intein, IODIDE ION | | Authors: | Hoffmann, S, Mootz, H.D, Kuemmel, D, Singh, R. | | Deposit date: | 2020-06-19 | | Release date: | 2020-09-16 | | Last modified: | 2021-01-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Biochemical and Structural Characterization of an Unusual and Naturally Split Class 3 Intein.
Chembiochem, 22, 2021
|
|
6HLU
 
 | |
7B22
 
 | | Vibrio cholerae ParD2 Antitoxin | | Descriptor: | Antitoxin ParD | | Authors: | Garcia-Rodriguez, G, Loris, R. | | Deposit date: | 2020-11-25 | | Release date: | 2021-10-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.08 Å) | | Cite: | Entropic pressure controls the oligomerization of the Vibrio cholerae ParD2 antitoxin.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
