2MCQ
 
 | | NMR structure of a BolA-like hypothetical protein RP812 from Rickettsia prowazekii, Seattle structural genomics center for infectious disease (SSGCID) | | Descriptor: | Uncharacterized protein RP812 | | Authors: | Chen, Y, Barnwal, R, Yang, F, Varani, G, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2013-08-22 | | Release date: | 2013-10-09 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | NMR structure of a BolA-like hypothetical protein RP812 from Rickettsia prowazekii, Seattle structural genomics center for infectious disease (SSGCID)
To be Published
|
|
2L4B
 
 | |
6CKQ
 
 | |
4G7F
 
 | |
3TE8
 
 | |
3IEF
 
 | |
5TQJ
 
 | |
4W5K
 
 | |
4WHX
 | | X-ray Crystal Structure of an Amino Acid Aminotransferase from Burkholderia pseudomallei Bound to the Co-factor Pyridoxal Phosphate | | Descriptor: | 1,2-ETHANEDIOL, ALANINE, Branched-chain-amino-acid transaminase | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID), Fairman, J.W, Dranow, D.M, Taylor, B.M, Lorimer, D, Edwards, T.E. | | Deposit date: | 2014-09-23 | | Release date: | 2014-10-08 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | X-ray Crystal Structure of an Amino Acid Aminotransferase from Burkholderia pseudomallei Bound to the Co-factor Pyridoxal Phosphate
to be published
|
|
4WJM
 
 | |
4WGJ
 
 | |
5SCO
 
 | |
5SD3
 
 | |
5SCV
 
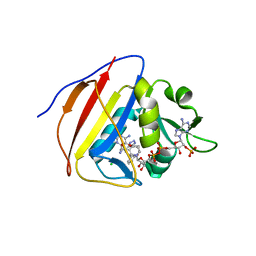 | |
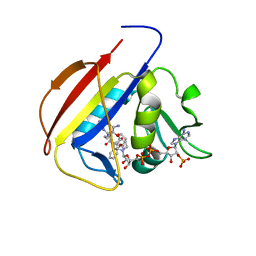
5SCR
 
 | |
4Q04
 
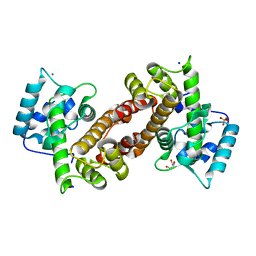 | |
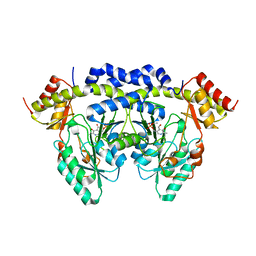
4QGR
 
 | |
6XR5
 
 | |
6XK2
 
 | |
6U3L
 
 | |
6PZJ
 | |
6TZ8
 
 | |
4XDQ
 
 | |
4EX4
 
 | |
6CXM
 
 | |
