4TT0
 | |
4TT1
 
 | |
4ADP
 
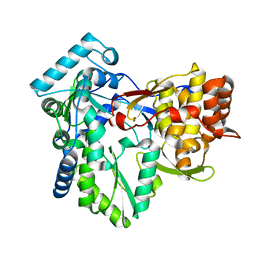 | | HCV-J6 NS5B POLYMERASE V405I MUTANT | | 分子名称: | RNA-DIRECTED RNA POLYMERASE | | 著者 | Scrima, N, Bressanelli, S. | | 登録日 | 2012-01-02 | | 公開日 | 2012-05-02 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.902 Å) | | 主引用文献 | Two Crucial Early Steps in RNA Synthesis by the Hepatitis C Virus Polymerase Involve a Dual Role of Residue 405.
J.Virol., 86, 2012
|
|
2XWH
 
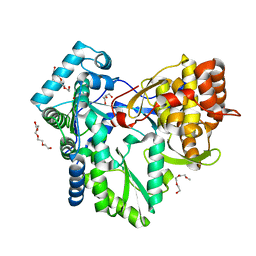 | | HCV-J6 NS5B polymerase structure at 1.8 Angstrom | | 分子名称: | DI(HYDROXYETHYL)ETHER, POLYETHYLENE GLYCOL (N=34), RNA DEPENDENT RNA POLYMERASE | | 著者 | Scrima, N, Bressanelli, S. | | 登録日 | 2010-11-03 | | 公開日 | 2011-01-12 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | A Comprehensive Structure-Function Comparison of Hepatitis C Virus Strains Jfh1 and J6 Polymerases Reveals a Key Residue Stimulating Replication in Cell Culture Across Genotypes.
J.Virol., 85, 2011
|
|
3Q87
 
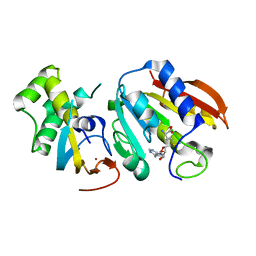 | | Structure of E. cuniculi Mtq2-Trm112 complex responible for the methylation of eRF1 translation termination factor | | 分子名称: | N6 adenine specific DNA methylase, Putative uncharacterized protein ECU08_1170, S-ADENOSYLMETHIONINE, ... | | 著者 | Liger, D, Mora, L, Lazar, N, Figaro, S, Henri, J, Scrima, N, Buckingham, R.H, van Tilbeurgh, H, Heurgue-Hamard, V, Graille, M. | | 登録日 | 2011-01-06 | | 公開日 | 2011-05-18 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (1.997 Å) | | 主引用文献 | Mechanism of activation of methyltransferases involved in translation by the Trm112 'hub' protein
Nucleic Acids Res., 2011
|
|
2B3T
 
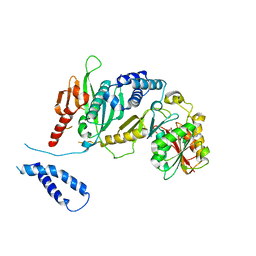 | | Structure of complex between E. coli translation termination factor RF1 and the PrmC methyltransferase | | 分子名称: | Peptide chain release factor 1, Protein methyltransferase hemK, S-ADENOSYL-L-HOMOCYSTEINE | | 著者 | Graille, M, Heurgue-Hamard, V, Champ, S, Mora, L, Scrima, N, Ulryck, N, van Tilbeurgh, H, Buckingham, R.H. | | 登録日 | 2005-09-21 | | 公開日 | 2006-01-24 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (3.1 Å) | | 主引用文献 | Molecular basis for bacterial class I release factor methylation by PrmC
Mol.Cell, 20, 2005
|
|
2J6A
 
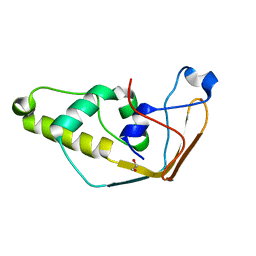 | | Structure of S. cerevisiae Trm112 protein, a methyltransferase activator | | 分子名称: | 1,2-ETHANEDIOL, PROTEIN TRM112, ZINC ION | | 著者 | Heurgue-Hamard, V, Graille, M, Scrima, N, Ulryck, N, Champ, S, Van Tilbeurgh, H, Buckingham, R.H. | | 登録日 | 2006-09-27 | | 公開日 | 2006-09-28 | | 最終更新日 | 2021-03-17 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | The Zinc Finger Protein Ynr046W is Plurifunctional and a Component of the Erf1 Methyltransferase in Yeast.
J.Biol.Chem., 281, 2006
|
|
4AEP
 
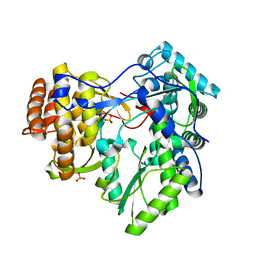 | |
4AEX
 
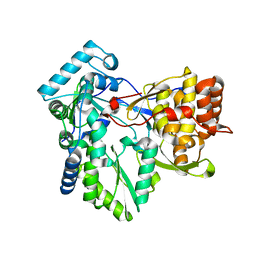 | |
2XYM
 
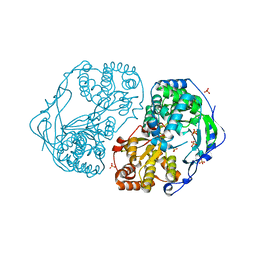 | | HCV-JFH1 NS5B T385A mutant | | 分子名称: | PHOSPHATE ION, RNA-DIRECTED RNA POLYMERASE | | 著者 | Simister, P.C, Caillet-Saguy, C, Bressanelli, S. | | 登録日 | 2010-11-18 | | 公開日 | 2011-01-12 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.774 Å) | | 主引用文献 | A Comprehensive Structure-Function Comparison of Hepatitis C Virus Strains Jfh1 and J6 Polymerases Reveals a Key Residue Stimulating Replication in Cell Culture Across Genotypes.
J.Virol., 85, 2011
|
|
2XXD
 
 | | HCV-JFH1 NS5B polymerase structure at 1.9 angstrom | | 分子名称: | PHOSPHATE ION, RNA-DIRECTED RNA POLYMERASE | | 著者 | Caillet-Saguy, C, Bressanelli, S. | | 登録日 | 2010-11-10 | | 公開日 | 2011-01-12 | | 最終更新日 | 2024-05-01 | | 実験手法 | X-RAY DIFFRACTION (1.881 Å) | | 主引用文献 | A Comprehensive Structure-Function Comparison of Hepatitis C Virus Strains Jfh1 and J6 Polymerases Reveals a Key Residue Stimulating Replication in Cell Culture Across Genotypes.
J.Virol., 85, 2011
|
|
