6SAE
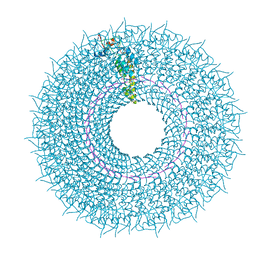 
 | | Cryo-EM structure of TMV in water | | Descriptor: | Capsid protein, MAGNESIUM ION, RNA (5'-R(P*GP*AP*A)-3') | | Authors: | Weis, F, Beckers, M, Sachse, C. | | Deposit date: | 2019-07-16 | | Release date: | 2019-09-18 | | Last modified: | 2019-11-13 | | Method: | ELECTRON MICROSCOPY (1.9 Å) | | Cite: | Elucidation of the viral disassembly switch of tobacco mosaic virus.
Embo Rep., 20, 2019
|
|
6SAG
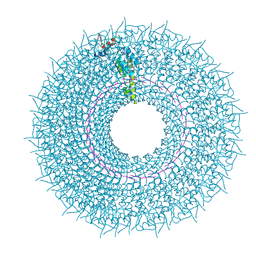 
 | | Cryo-EM structure of TMV with Ca2+ at low pH | | Descriptor: | CALCIUM ION, Capsid protein, MAGNESIUM ION, ... | | Authors: | Weis, F, Beckers, M, Sachse, C. | | Deposit date: | 2019-07-16 | | Release date: | 2019-09-18 | | Last modified: | 2019-11-13 | | Method: | ELECTRON MICROSCOPY (2 Å) | | Cite: | Elucidation of the viral disassembly switch of tobacco mosaic virus.
Embo Rep., 20, 2019
|
|
6TGS
 
 | | AtNBR1-PB1 domain | | Descriptor: | CHLORIDE ION, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Jakobi, A.J, Sachse, C. | | Deposit date: | 2019-11-17 | | Release date: | 2020-02-12 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Structural basis of p62/SQSTM1 helical filaments and their role in cellular cargo uptake.
Nat Commun, 11, 2020
|
|
6TGN
 
 | |
6TGP
 
 | |
6TGY
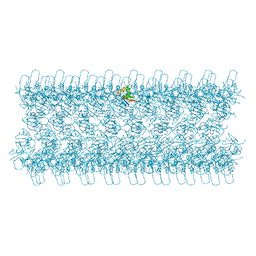 
 | |
6TH3
 
 | |
5OA1
 
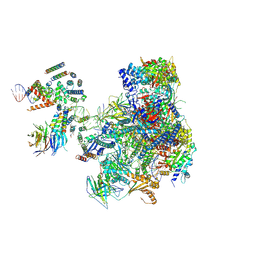 | | RNA polymerase I pre-initiation complex | | Descriptor: | ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA, ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA, ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA, ... | | Authors: | Sadian, Y, Tafur, L, Kosinski, J, Jakobi, A.J, Muller, C.W. | | Deposit date: | 2017-06-20 | | Release date: | 2017-07-26 | | Last modified: | 2018-10-24 | | Method: | ELECTRON MICROSCOPY (4.4 Å) | | Cite: | Structural insights into transcription initiation by yeast RNA polymerase I.
EMBO J., 36, 2017
|
|
6GFJ
 
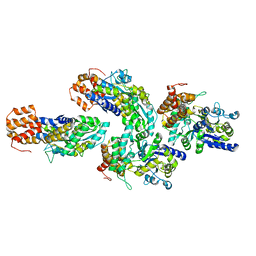 | | Structure of RIP2 CARD domain fused to crystallisable MBP tag | | Descriptor: | Sugar ABC transporter substrate-binding protein,Receptor-interacting serine/threonine-protein kinase 2, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Pellegrini, E, Cusack, S. | | Deposit date: | 2018-04-30 | | Release date: | 2019-03-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | RIP2 filament formation is required for NOD2 dependent NF-kappa B signalling.
Nat Commun, 9, 2018
|
|
5M2N
 
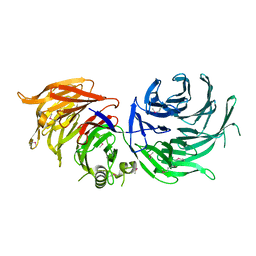 | |
5KC1
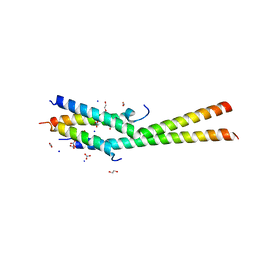 
 | | Structure of the C-terminal dimerization domain of Atg38 | | Descriptor: | 1,2-ETHANEDIOL, AMMONIUM ION, Autophagy-related protein 38, ... | | Authors: | Ohashi, Y, Soler, N, Garcia-Ortegon, M, Zhang, L, Perisic, O, Masson, G.R, Johnson, C.M, Williams, R.J. | | Deposit date: | 2016-06-04 | | Release date: | 2016-10-05 | | Last modified: | 2017-09-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Characterization of Atg38 and NRBF2, a fifth subunit of the autophagic Vps34/PIK3C3 complex.
Autophagy, 12, 2016
|
|
