2W9J
 
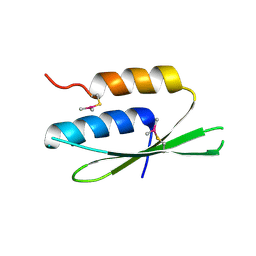 | | The crystal structure of SRP14 from the Schizosaccharomyces pombe signal recognition particle | | Descriptor: | SIGNAL RECOGNITION PARTICLE SUBUNIT SRP14 | | Authors: | Brooks, M.A, Ravelli, R.B.G, McCarthy, A.A, Strub, K, Cusack, S. | | Deposit date: | 2009-01-25 | | Release date: | 2009-02-03 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of Srp14 from the Schizosaccharomyces Pombe Signal Recognition Particle.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
2WFH
 
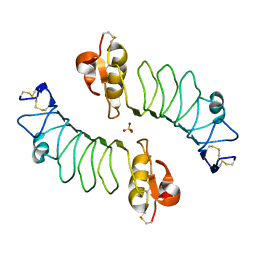 | | The Human Slit 2 Dimerization Domain D4 | | Descriptor: | SLIT HOMOLOG 2 PROTEIN C-PRODUCT, SULFATE ION | | Authors: | Seiradake, E, von Philipsborn, A.C, Henry, M, Fritz, M, Lortat-Jacob, H, Jamin, M, Hemrika, W, Bastmeyer, M, Cusack, S, McCarthy, A.A. | | Deposit date: | 2009-04-06 | | Release date: | 2009-04-21 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure and functional relevance of the Slit2 homodimerization domain.
EMBO Rep., 10, 2009
|
|
2X7A
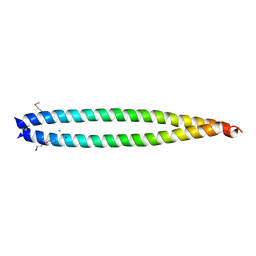 
 | | Structural basis of HIV-1 tethering to membranes by the Bst2-tetherin ectodomain | | Descriptor: | BONE MARROW STROMAL ANTIGEN 2, CHLORIDE ION, GLYCEROL, ... | | Authors: | Natrajan, G, McCarthy, A.A, Weissenhorn, W. | | Deposit date: | 2010-02-25 | | Release date: | 2010-04-28 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.77 Å) | | Cite: | Structural Basis of HIV-1 Tethering to Membranes by the Bst-2/Tetherin Ectodomain.
Cell Host Microbe, 7, 2010
|
|
6ZHB
 
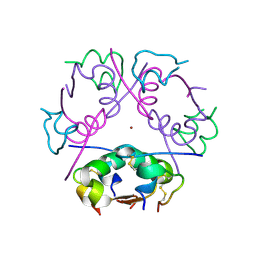 | | 3D electron diffraction structure of bovine insulin | | Descriptor: | Insulin, ZINC ION | | Authors: | Blum, T, Housset, D, Clabbers, M.T.B, van Genderen, E, Bacia-Verloop, M, Zander, U, McCarthy, A.A, Schoehn, G, Ling, W.L, Abrahams, J.P. | | Deposit date: | 2020-06-22 | | Release date: | 2021-01-27 | | Last modified: | 2024-01-24 | | Method: | ELECTRON CRYSTALLOGRAPHY (3.25 Å) | | Cite: | Statistically correcting dynamical electron scattering improves the refinement of protein nanocrystals, including charge refinement of coordinated metals.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
6ZI8
 
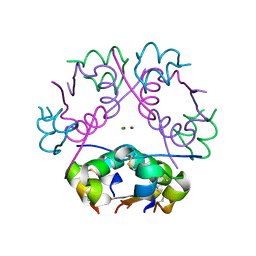 | | X-ray diffraction structure of bovine insulin at 2.3 A resolution | | Descriptor: | CHLORIDE ION, Insulin, ZINC ION | | Authors: | Housset, D, Ling, W.L, Bacia-Verloop, M, Zander, U, McCarthy, A.A, Schoehn, G. | | Deposit date: | 2020-06-25 | | Release date: | 2021-01-20 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Statistically correcting dynamical electron scattering improves the refinement of protein nanocrystals, including charge refinement of coordinated metals.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
2XMW
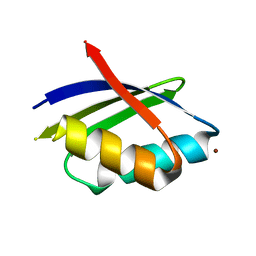 
 | | PacS, N-terminal domain, from Synechocystis PCC6803 | | Descriptor: | CATION-TRANSPORTING ATPASE PACS, COPPER (I) ION | | Authors: | Badarau, A, Firbank, S.J, McCarthy, A.A, Banfield, M.J, Dennison, C. | | Deposit date: | 2010-07-29 | | Release date: | 2010-08-18 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Visualizing the Metal-Binding Versatility of Copper Trafficking Sites .
Biochemistry, 49, 2010
|
|
2XMJ
 
 | | Visualising the Metal-binding Versatility of Copper Trafficking Sites: Atx1 side-to-side (aerobic) | | Descriptor: | CHLORIDE ION, COPPER (II) ION, SODIUM ION, ... | | Authors: | Badarau, A, Firbank, S.J, McCarthy, A.A, Banfield, M.J, Dennison, C. | | Deposit date: | 2010-07-28 | | Release date: | 2010-08-25 | | Last modified: | 2019-05-22 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | Visualizing the Metal-Binding Versatility of Copper Trafficking Sites .
Biochemistry, 49, 2010
|
|
2XMK
 
 | | Visualising the Metal-binding Versatility of Copper Trafficking Sites: Atx1 side-to-side (anaerobic) | | Descriptor: | CHLORIDE ION, COPPER (II) ION, SODIUM ION, ... | | Authors: | Badarau, A, Firbank, S.J, McCarthy, A.A, Banfield, M.J, Dennison, C. | | Deposit date: | 2010-07-28 | | Release date: | 2010-08-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Visualizing the Metal-Binding Versatility of Copper Trafficking Sites .
Biochemistry, 49, 2010
|
|
2XMU
 
 | | Copper chaperone Atx1 from Synechocystis PCC6803 (Cu2 form) | | Descriptor: | COPPER (I) ION, SSR2857 PROTEIN | | Authors: | Badarau, A, Firbank, S.J, McCarthy, A.A, Banfield, M.J, Dennison, C. | | Deposit date: | 2010-07-29 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Visualizing the Metal-Binding Versatility of Copper Trafficking Sites .
Biochemistry, 49, 2010
|
|
2XMT
 
 | | Copper chaperone Atx1 from Synechocystis PCC6803 (Cu1 form) | | Descriptor: | COPPER (I) ION, SSR2857 PROTEIN | | Authors: | Badarau, A, Firbank, S.J, McCarthy, A.A, Banfield, M.J, Dennison, C. | | Deposit date: | 2010-07-29 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Visualizing the Metal-Binding Versatility of Copper Trafficking Sites .
Biochemistry, 49, 2010
|
|
2XMM
 
 | | Visualising the Metal-binding Versatility of Copper Trafficking Sites: H61Y Atx1 side-to-side | | Descriptor: | CHLORIDE ION, COPPER (II) ION, SODIUM ION, ... | | Authors: | Badarau, A, Firbank, S.J, McCarthy, A.A, Banfield, M.J, Dennison, C. | | Deposit date: | 2010-07-28 | | Release date: | 2010-08-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Visualizing the Metal-Binding Versatility of Copper Trafficking Sites .
Biochemistry, 49, 2010
|
|
2XZS
 
 | | Death associated protein kinase 1 residues 1-312 | | Descriptor: | DEATH ASSOCIATED KINASE 1, MAGNESIUM ION | | Authors: | Yumerefendi, H, Mas, P.J, Dordevic, N, McCarthy, A.A, Hart, D.J. | | Deposit date: | 2010-11-29 | | Release date: | 2011-12-07 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Death-Associated Protein Kinase Activity is Regulated by Coupled Calcium/Calmodulin Binding to Two Distinct Sites.
Structure, 24, 2016
|
|
2XMV
 
 | | Copper chaperone Atx1 from Synechocystis PCC6803 (Cu1, trimeric form, His61Tyr mutant) | | Descriptor: | COPPER (I) ION, SSR2857 PROTEIN | | Authors: | Badarau, A, Firbank, S.J, McCarthy, A.A, Banfield, M.J, Dennison, C. | | Deposit date: | 2010-07-29 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Visualizing the Metal-Binding Versatility of Copper Trafficking Sites .
Biochemistry, 49, 2010
|
|
2Y0A
 
 | | Structure of DAPK1 construct residues 1-304 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, DEATH-ASSOCIATED PROTEIN KINASE 1 | | Authors: | Yumerefendi, H, Mas, P.J, Dordevic, N, McCarthy, A.A, Hart, D.J. | | Deposit date: | 2010-12-01 | | Release date: | 2011-12-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Library-Based Construct Screening of Death-Associated Protein Kinase 1 Identifies the Minimal Calmodulin Interaction Region and Autoinhibitory Conformation of the Catalytic Domain
To be Published
|
|
2UUH
 
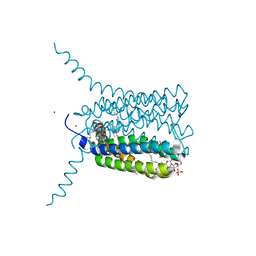 | | Crystal structure of Human Leukotriene C4 Synthase in complex with substrate glutathione | | Descriptor: | DODECYL-ALPHA-D-MALTOSIDE, GLUTATHIONE, LEUKOTRIENE C4 SYNTHASE, ... | | Authors: | Martinez Molina, D, Wetterholm, A, Kohl, A, McCarthy, A.A, Niegowski, D, Ohlson, E, Hammarberg, T, Eshaghi, S, Haeggstrom, J.Z, Nordlund, P. | | Deposit date: | 2007-03-02 | | Release date: | 2007-07-17 | | Last modified: | 2011-12-07 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structural Basis for Synthesis of Inflammatory Mediators by Human Leukotriene C4 Synthase.
Nature, 448, 2007
|
|
2UUI
 
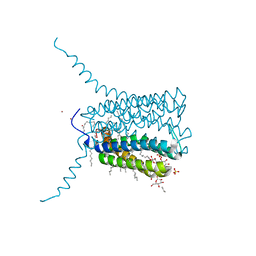 | | Crystal structure of Human Leukotriene C4 Synthase | | Descriptor: | DODECYL-BETA-D-MALTOSIDE, LEUKOTRIENE C4 SYNTHASE, NICKEL (II) ION, ... | | Authors: | Martinez Molina, D, Wetterholm, A, Kohl, A, McCarthy, A.A, Niegowski, D, Ohlson, E, Hammarberg, T, Eshaghi, S, Haeggstrom, J.Z, Nordlund, P. | | Deposit date: | 2007-03-02 | | Release date: | 2007-07-17 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for Synthesis of Inflammatory Mediators by Human Leukotriene C4 Synthase.
Nature, 448, 2007
|
|
6GFN
 
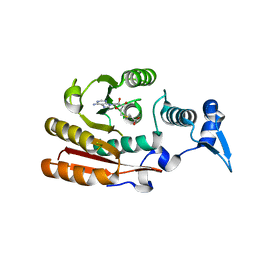 | | METTL16 MTase domain | | Descriptor: | S-ADENOSYL-L-HOMOCYSTEINE, U6 small nuclear RNA (adenine-(43)-N(6))-methyltransferase | | Authors: | Chen, K.M, Mendel, M, Homolka, D, McCarthy, A.A, Pillai, R.S. | | Deposit date: | 2018-05-01 | | Release date: | 2018-09-19 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.86 Å) | | Cite: | Methylation of Structured RNA by the m6A Writer METTL16 Is Essential for Mouse Embryonic Development.
Mol. Cell, 71, 2018
|
|
6GT5
 
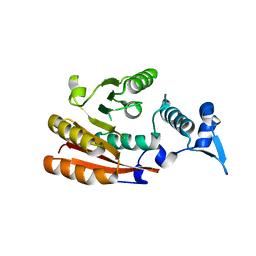 | | METTL16 MTase domain (crystal form 2) | | Descriptor: | U6 small nuclear RNA (adenine-(43)-N(6))-methyltransferase | | Authors: | Chen, K.M, Mendel, M, Homolka, D, McCarthy, A.A, Pillai, R.S. | | Deposit date: | 2018-06-15 | | Release date: | 2018-09-19 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Methylation of Structured RNA by the m6A Writer METTL16 Is Essential for Mouse Embryonic Development.
Mol. Cell, 71, 2018
|
|
6GFK
 
 | | delta-N METTL16 MTase domain | | Descriptor: | S-ADENOSYL-L-HOMOCYSTEINE, SULFATE ION, U6 small nuclear RNA (adenine-(43)-N(6))-methyltransferase | | Authors: | Chen, K.M, Mendel, M, Homolka, D, McCarthy, A.A, Pillai, R.S. | | Deposit date: | 2018-04-30 | | Release date: | 2018-09-19 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Methylation of Structured RNA by the m6A Writer METTL16 Is Essential for Mouse Embryonic Development.
Mol. Cell, 71, 2018
|
|
4A2Q
 
 | | Structure of duck RIG-I tandem CARDs and helicase domain | | Descriptor: | RETINOIC ACID INDUCIBLE PROTEIN I | | Authors: | Kowalinski, E, Lunardi, T, McCarthy, A.A, Cusack, S. | | Deposit date: | 2011-09-28 | | Release date: | 2011-10-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Structural Basis for the Activation of Innate Immune Pattern Recognition Receptor Rig-I by Viral RNA.
Cell(Cambridge,Mass.), 147, 2011
|
|
4A2P
 
 | | Structure of duck RIG-I helicase domain | | Descriptor: | RETINOIC ACID INDUCIBLE PROTEIN I | | Authors: | Kowalinski, E, Lunardi, T, McCarthy, A.A, Cusack, S. | | Deposit date: | 2011-09-28 | | Release date: | 2011-10-19 | | Last modified: | 2019-04-03 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural Basis for the Activation of Innate Immune Pattern Recognition Receptor Rig-I by Viral RNA.
Cell(Cambridge,Mass.), 147, 2011
|
|
4A2V
 
 | | Structure of duck RIG-I C-terminal domain (CTD) | | Descriptor: | GLYCEROL, RETINOIC ACID INDUCIBLE PROTEIN I, ZINC ION | | Authors: | Kowalinski, E, Lunardi, T, McCarthy, A.A, Cusack, S. | | Deposit date: | 2011-09-29 | | Release date: | 2011-10-19 | | Last modified: | 2011-10-26 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | Structural Basis for the Activation of Innate Immune Pattern Recognition Receptor Rig-I by Viral RNA.
Cell(Cambridge,Mass.), 147, 2011
|
|
4A2W
 
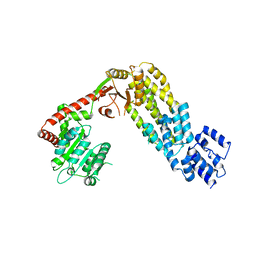 | | Structure of full-length duck RIG-I | | Descriptor: | RETINOIC ACID INDUCIBLE PROTEIN I | | Authors: | Kowalinski, E, Lunardi, T, McCarthy, A.A, Cusack, S. | | Deposit date: | 2011-09-29 | | Release date: | 2011-10-19 | | Last modified: | 2019-03-27 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | Structural Basis for the Activation of Innate Immune Pattern Recognition Receptor Rig-I by Viral RNA.
Cell(Cambridge,Mass.), 147, 2011
|
|
4A2X
 
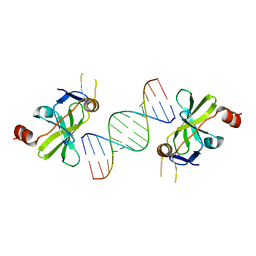 | | Structure of duck RIG-I C-terminal domain (CTD) with 14-mer dSRNA | | Descriptor: | 5'-R(*CP*GP*CP*GP*UP*UP*GP*UP*UP*CP*UP*CP*CP*CP)-3', 5'-R(*GP*GP*GP*AP*GP*AP*AP*CP*AP*AP*CP*GP*CP*GP)-3', RETINOIC ACID INDUCIBLE PROTEIN I, ... | | Authors: | Kowalinski, E, Lunardi, T, McCarthy, A.A, Cusack, S. | | Deposit date: | 2011-09-29 | | Release date: | 2011-10-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (4 Å) | | Cite: | Structural basis for the activation of innate immune pattern-recognition receptor RIG-I by viral RNA.
Cell, 147, 2011
|
|
4A36
 
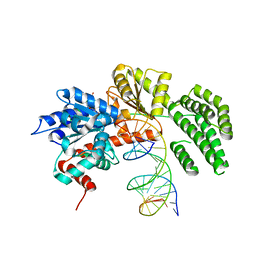 | | Structure of duck RIG-I helicase domain bound to 19-mer dsRNA and ATP transition state analogue | | Descriptor: | 5'-R(*GP*CP*AP*UP*GP*CP*GP*AP*CP*CP*UP*CP*UP*GP *UP*UP*UP*GP*A)-3', 5'-R(*UP*CP*AP*AP*AP*CP*AP*GP*AP*GP*GP*UP*CP*GP *CP*AP*UP*GP*C)-3', ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Kowalinski, E, Lunardi, T, McCarthy, A.A, Cusack, S. | | Deposit date: | 2011-09-30 | | Release date: | 2011-10-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | Structural Basis for the Activation of Innate Immune Pattern Recognition Receptor Rig-I by Viral RNA.
Cell(Cambridge,Mass.), 147, 2011
|
|
