5EWP
 
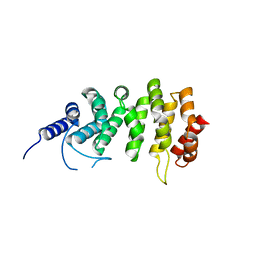 | | ARO (armadillo repeats only protein) from Plasmodium falciparum | | Descriptor: | ARO (armadillo repeats only protein) | | Authors: | Brown, C, Zhang, K, Emery, J, Prusty, D, Wetzel, J, Heincke, D, Gilberger, T, Junop, M. | | Deposit date: | 2015-11-20 | | Release date: | 2016-01-27 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | ARO (armadillo repeats only protein) from Plasmodium falciparum
To Be Published
|
|
6BD0
 
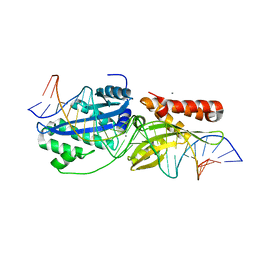 | | I-OnuI K227Y, D236A bound to cognate substrate (pre-cleavage complex) | | Descriptor: | CALCIUM ION, DNA (25-MER), Ribosomal protein 3/homing endonuclease-like protein fusion | | Authors: | Brown, C, Zhang, K, Laforet, M, McMurrough, T.A, Gloor, G.B, Edgell, D.R, Junop, M. | | Deposit date: | 2017-10-20 | | Release date: | 2018-10-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | I-OnuI K227Y, D236A bound to cognate substrate (pre-cleavage complex)
To Be Published
|
|
6BDA
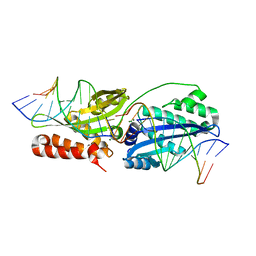 
 | | Wild-type I-OnuI bound to A3G substrate (post-cleavage complex) | | Descriptor: | Cleaved Cognate DNA strand, +11 sense, Cleaved cognate DNA strand, ... | | Authors: | Brown, C, Zhang, K, Laforet, M, McMurrough, T, Gloor, G.B, Edgell, D.R, Junop, M. | | Deposit date: | 2017-10-22 | | Release date: | 2018-10-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Wild-type I-OnuI bound to A3G substrate (post-cleavage complex)
To Be Published
|
|
6BDB
 
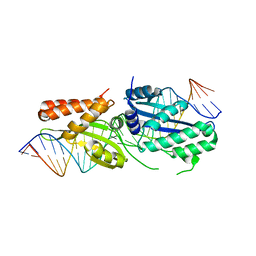 | | I-OnuI K227Y, D236A bound to A3G substrate (pre-cleavage complex) | | Descriptor: | DNA (26-MER), Ribosomal protein 3/homing endonuclease-like protein fusion | | Authors: | Brown, C, Zhang, K, McMurrough, T.A, Laforet, M, Gloor, G.B, Edgell, D.R, Junop, M. | | Deposit date: | 2017-10-22 | | Release date: | 2018-10-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | I-OnuI K227Y, D236A bound to A3G substrate (pre-cleavage complex)
To Be Published
|
|
4F6L
 
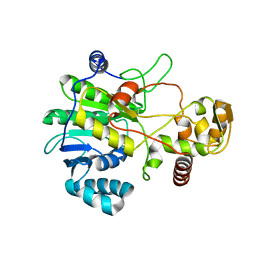 | |
4DZ1
 
 | |
4F6C
 
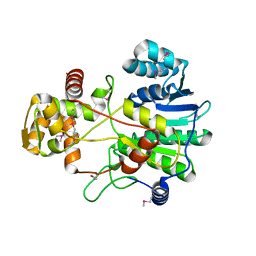 | |
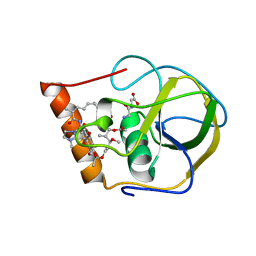
2HW2
 
 | |
6BCH
 
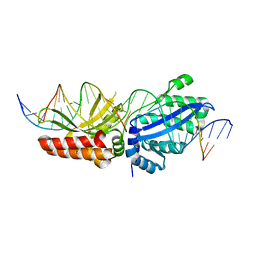 | | I-LtrI E29D bound to cognate substrate (nicked complex) | | Descriptor: | DNA (26-MER), DNA (5'-D(*GP*GP*TP*CP*TP*AP*AP*AP*CP*GP*TP*CP*GP*TP*AP*T)-3'), DNA (5'-D(*TP*AP*GP*GP*AP*GP*CP*AP*TP*TP*T)-3'), ... | | Authors: | Brown, C, Zhang, K, McMurrough, T.A, Gloor, G.B, Edgell, D.R, Junop, M. | | Deposit date: | 2017-10-20 | | Release date: | 2018-10-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | I-LtrI E29D bound to cognate substrate (nicked complex)
To Be Published
|
|
6BDG
 
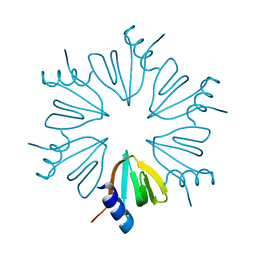 | | HFQ monomer in spacegroup p6 at 1.93 angstrom resolution | | Descriptor: | RNA-binding protein Hfq | | Authors: | Brown, C, Zhang, K, Seo, C, Ellis, M.J, Hanniford, D.B, Junop, M. | | Deposit date: | 2017-10-23 | | Release date: | 2017-11-01 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.964 Å) | | Cite: | HFQ monomer in spacegroup p6 at 1.93 angstrom resolution
To Be Published
|
|
1FU1
 
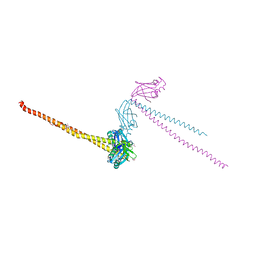 | | CRYSTAL STRUCTURE OF HUMAN XRCC4 | | Descriptor: | ACETIC ACID, DNA REPAIR PROTEIN XRCC4 | | Authors: | Junop, M, Modesti, M, Guarne, A, Gellert, M, Yang, W. | | Deposit date: | 2000-09-13 | | Release date: | 2000-12-11 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the Xrcc4 DNA repair protein and implications for end joining.
EMBO J., 19, 2000
|
|
3EPU
 
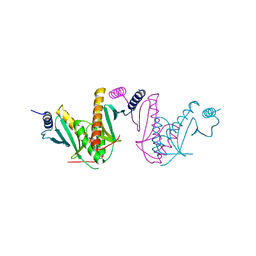 | | Crystal Structure of STM2138, a novel virulence chaperone in Salmonella | | Descriptor: | STM2138 Virulence Chaperone | | Authors: | Zhang, K, Andres, S.N, Hannemann, M, Coombes, B, Junop, M. | | Deposit date: | 2008-09-30 | | Release date: | 2009-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural analysis and quantitative proteomic
interactome of a novel virulence chaperone in Salmonella
To be Published
|
|
2I22
 
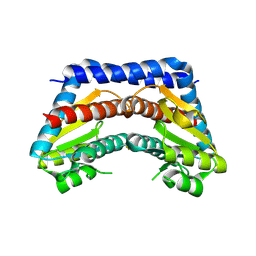 | | Crystal structure of Escherichia coli phosphoheptose isomerase in complex with reaction substrate sedoheptulose 7-phosphate | | Descriptor: | D-ALTRO-HEPT-2-ULOSE 7-PHOSPHATE, Phosphoheptose isomerase | | Authors: | Blakely, K, Zhang, K, DeLeon, G, Wright, G, Junop, M. | | Deposit date: | 2006-08-15 | | Release date: | 2007-08-21 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure and Function of Sedoheptulose-7-phosphate Isomerase, a Critical Enzyme for Lipopolysaccharide Biosynthesis and a Target for Antibiotic Adjuvants
J.Biol.Chem., 283, 2008
|
|
2I2W
 
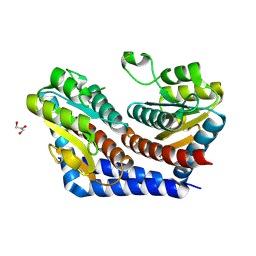 | | Crystal Structure of Escherichia Coli Phosphoheptose Isomerase | | Descriptor: | GLYCEROL, Phosphoheptose isomerase | | Authors: | DeLeon, G, Blakely, K, Zhang, K, Wright, G, Junop, M. | | Deposit date: | 2006-08-17 | | Release date: | 2007-08-21 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure and Function of Sedoheptulose-7-phosphate Isomerase, a Critical Enzyme for Lipopolysaccharide Biosynthesis and a Target for Antibiotic Adjuvants
J.Biol.Chem., 283, 2008
|
|
3II6
 
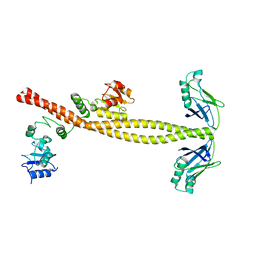 | |
3JYZ
 
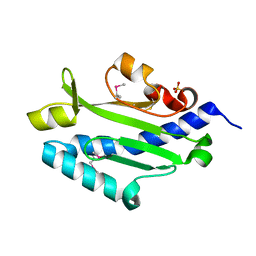 | | Crystal structure of Pseudomonas aeruginosa (strain: Pa110594) typeIV pilin in space group P41212 | | Descriptor: | SULFATE ION, Type IV pilin structural subunit | | Authors: | Nguyen, Y, Jackson, S.G, Aidoo, F, Junop, M.S, Burrows, L.L. | | Deposit date: | 2009-09-22 | | Release date: | 2009-11-24 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural characterization of Novel Pseudomonas aeruginosa type IV pilins.
J.Mol.Biol., 395, 2010
|
|
3JZZ
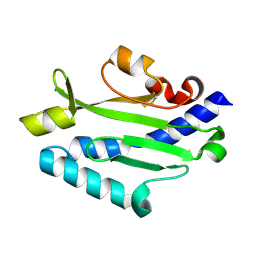 
 | | Crystal structure of Pseudomonas aeruginosa (strain: Pa110594) typeIV pilin in space group P212121 | | Descriptor: | Type IV pilin structural subunit | | Authors: | Nguyen, Y, Jackson, S.G, Aidoo, F, Junop, M.S, Burrows, L.L. | | Deposit date: | 2009-09-24 | | Release date: | 2009-11-24 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.597 Å) | | Cite: | Structural characterization of Novel Pseudomonas aeruginosa type IV pilins.
J.Mol.Biol., 395, 2010
|
|
5CKS
 
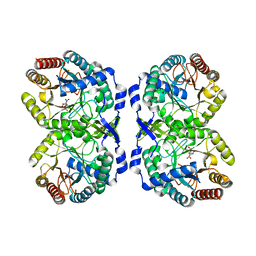 | | DAHP (3-deoxy-D-arabinoheptulosonate-7-phosphate) Synthase in complex with DAHP Oxime. | | Descriptor: | DAHP Oxime, Phospho-2-dehydro-3-deoxyheptonate aldolase, Phe-sensitive, ... | | Authors: | Berti, P, Junop, M, Balachandran, N. | | Deposit date: | 2015-07-15 | | Release date: | 2016-08-10 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.1181 Å) | | Cite: | Potent Inhibition of 3-Deoxy-d-arabinoheptulosonate-7-phosphate (DAHP) Synthase by DAHP Oxime, a Phosphate Group Mimic.
Biochemistry, 55, 2016
|
|
3JZ0
 
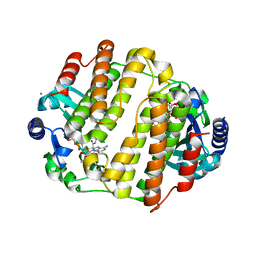 | | LinB complexed with clindamycin and AMPCPP | | Descriptor: | CLINDAMYCIN, DIPHOSPHOMETHYLPHOSPHONIC ACID ADENOSYL ESTER, Lincosamide nucleotidyltransferase, ... | | Authors: | Morar, M, Wright, G.D. | | Deposit date: | 2009-09-22 | | Release date: | 2009-11-03 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure and mechanism of the lincosamide antibiotic adenylyltransferase LinB.
Structure, 17, 2009
|
|
3JYY
 
 | | SeMet LinB complexed with PPi | | Descriptor: | Lincosamide nucleotidyltransferase, MAGNESIUM ION, PYROPHOSPHATE | | Authors: | Morar, M, Wright, G.D. | | Deposit date: | 2009-09-22 | | Release date: | 2009-11-03 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure and mechanism of the lincosamide antibiotic adenylyltransferase LinB.
Structure, 17, 2009
|
|
6BCG
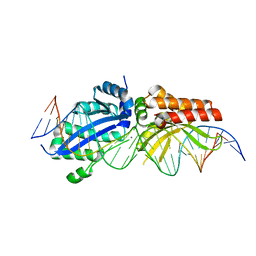 
 | | I-LtrI A28G bound to cognate substrate (pre-cleavage complex) | | Descriptor: | CALCIUM ION, DNA (26-MER), Ribosomal protein 3/homing endonuclease-like fusion protein | | Authors: | Brown, C, Zhang, K, McMurrough, T.A, Gloor, G.B, Edgell, D.R, Junop, M. | | Deposit date: | 2017-10-20 | | Release date: | 2018-10-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Active site residue identity regulates cleavage preference of LAGLIDADG homing endonucleases.
Nucleic Acids Res., 46, 2018
|
|
6BCF
 
 | | I-LtrI G183A bound to cognate substrate (pre-cleavage complex) | | Descriptor: | CALCIUM ION, DNA (26-MER), Ribosomal protein 3/homing endonuclease-like fusion protein | | Authors: | Brown, C, Zhang, K, McMurrough, T.A, Gloor, G.B, Edgell, D.R, Junop, M. | | Deposit date: | 2017-10-20 | | Release date: | 2018-10-24 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.92 Å) | | Cite: | Active site residue identity regulates cleavage preference of LAGLIDADG homing endonucleases.
Nucleic Acids Res., 46, 2018
|
|
6BCT
 
 | | I-LtrI E184D bound to non-cognate C4 substrate (pre-cleavage complex) | | Descriptor: | CALCIUM ION, DNA (26-MER), DNA (27-MER), ... | | Authors: | Brown, C, Zhang, K, McMurrough, T.A, Gloor, G.B, Edgell, D.R, Junop, M. | | Deposit date: | 2017-10-20 | | Release date: | 2018-10-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.73 Å) | | Cite: | Active site residue identity regulates cleavage preference of LAGLIDADG homing endonucleases.
Nucleic Acids Res., 46, 2018
|
|
6BCN
 
 | | I-LtrI E184D bound to cognate substrate (pre-cleavage complex) | | Descriptor: | CALCIUM ION, DNA (26-MER), Ribosomal protein 3/homing endonuclease-like fusion protein | | Authors: | Brown, C, Zhang, K, McMurrough, T.A, Gloor, G.B, Edgell, D.R, Junop, M. | | Deposit date: | 2017-10-20 | | Release date: | 2018-10-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Active site residue identity regulates cleavage preference of LAGLIDADG homing endonucleases.
Nucleic Acids Res., 46, 2018
|
|
6BCI
 
 | | Wild-type I-LtrI bound to non-cognate C4 substrate (pre-cleavage complex) | | Descriptor: | CALCIUM ION, DNA (27-MER), Ribosomal protein 3/homing endonuclease-like fusion protein | | Authors: | Brown, C, Zhang, K, McMurrough, T.A, Gloor, G.B, Edgell, D.R, Junop, M. | | Deposit date: | 2017-10-20 | | Release date: | 2018-10-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Active site residue identity regulates cleavage preference of LAGLIDADG homing endonucleases.
Nucleic Acids Res., 46, 2018
|
|
