9YA9
 
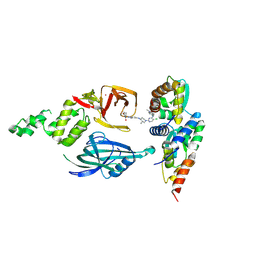 | | Cryo-EM structure of ternary complex BCL6-CRBN-DDB1 with BMS-986458 (local refined), a potent and selective BCL6 ligand directed degrader (LDD) | | Descriptor: | (3R)-3-(6-{[(3R,4R)-1-{5-chloro-4-[(1-methyl-2-oxo-2,3-dihydro-1H-indol-5-yl)amino]pyrimidin-2-yl}-3-methylpiperidin-4-yl]amino}-1-methyl-1H-indazol-3-yl)piperidine-2,6-dione, B-cell lymphoma 6 protein, Protein cereblon, ... | | Authors: | Zhu, J, Fang, W, Pagarigan, B. | | Deposit date: | 2025-09-15 | | Release date: | 2026-02-18 | | Last modified: | 2026-03-11 | | Method: | ELECTRON MICROSCOPY (3.09 Å) | | Cite: | Discovery of BMS-986458, a Potent and Selective B-Cell Lymphoma 6 Protein Ligand-Directed Degrader, for the Treatment of B-Cell Non-Hodgkin Lymphoma.
J.Med.Chem., 69, 2026
|
|
9VIN
 
 | | Ternary complex of TNFR1-DD, TRADD-DD and RIPK1-DD | | Descriptor: | Receptor-interacting serine/threonine-protein kinase 1, Tumor necrosis factor receptor superfamily member 1A, membrane form, ... | | Authors: | Liu, J, Han, Y, Zhao, J, Gao, J, Yuan, J. | | Deposit date: | 2025-06-18 | | Release date: | 2026-04-08 | | Last modified: | 2026-04-15 | | Method: | ELECTRON MICROSCOPY (3.41 Å) | | Cite: | Electric dipole moment drives the dynamics of the TNFR1 complex I signalosome.
Nature, 2026
|
|
9Z9L
 
 | | Q23.MD39 in Complex with Fabs from antibodies CH01 and 35O22 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lin, Z.J, Cui, J, Du, J, Habib, R, Kulp, D, Pallesen, J. | | Deposit date: | 2025-11-18 | | Release date: | 2026-04-22 | | Method: | ELECTRON MICROSCOPY (3.49 Å) | | Cite: | Deep Mining of the Human Antibody Repertoire Identifies Frequent and Genetically Diverse CDRH3 Topologies Targetable by Vaccination
Proc.Natl.Acad.Sci.USA, 2026
|
|
9VGD
 
 | | Helical assembly of TRADD death domain | | Descriptor: | Tumor necrosis factor receptor type 1-associated DEATH domain protein | | Authors: | Liu, J, Han, Y. | | Deposit date: | 2025-06-13 | | Release date: | 2026-04-08 | | Last modified: | 2026-04-15 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Electric dipole moment drives the dynamics of the TNFR1 complex I signalosome.
Nature, 2026
|
|
9YQO
 
 | | AMC008 v4.2 SOSIP Env trimer in complex with PGT121 and VRC01 Fabs | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Envelope glycoprotein gp41, ... | | Authors: | Cui, J, Du, J, Lin, Z, Pallesen, J. | | Deposit date: | 2025-10-15 | | Release date: | 2026-01-21 | | Last modified: | 2026-03-11 | | Method: | ELECTRON MICROSCOPY (3.26 Å) | | Cite: | Conformational landscape of HIV-1 Env from closed to fully open.
Nat Commun, 2026
|
|
9VJL
 
 | | Phosphorylation dependent recognition of RIPK1 by phosphatidylinositol 3,4,5-trisphosphate 5-phosphatase 1 | | Descriptor: | Phosphatidylinositol 3,4,5-trisphosphate 5-phosphatase 1, Receptor-interacting serine/threonine-protein kinase 1 | | Authors: | Liu, J, Liu, Z, Xue, H, Xie, X, Sun, W, Chou, J. | | Deposit date: | 2025-06-20 | | Release date: | 2026-03-25 | | Method: | SOLUTION NMR | | Cite: | Repression of RIPK1 kinase by INPP5D inhibits expression of diverse proinflammatory mediators and late-onset Alzheimer's disease risk factors.
Immunity, 59, 2026
|
|
5FBC
 
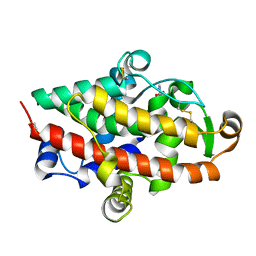 | | S1 nuclease from Aspergillus oryzae in complex with 2'-deoxyadenosine-5'-thio-monophosphate (5'dAMP(S)). | | Descriptor: | 2-DEOXY-ADENOSINE -5'-THIO-MONOPHOSPHATE, 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, ... | | Authors: | Koval, T, Oestergaard, L.H, Dohnalek, J. | | Deposit date: | 2015-12-14 | | Release date: | 2016-12-28 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural and Catalytic Properties of S1 Nuclease from Aspergillus oryzae Responsible for Substrate Recognition, Cleavage, Non-Specificity, and Inhibition.
PLoS ONE, 11, 2016
|
|
5FB9
 
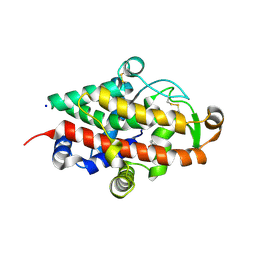 | | S1 nuclease from Aspergillus oryzae with unoccupied active site | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Nuclease S1, ... | | Authors: | Koval, T, Oestergaard, L.H, Dohnalek, J. | | Deposit date: | 2015-12-14 | | Release date: | 2016-12-28 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural and Catalytic Properties of S1 Nuclease from Aspergillus oryzae Responsible for Substrate Recognition, Cleavage, Non-Specificity, and Inhibition.
PLoS ONE, 11, 2016
|
|
5FBF
 
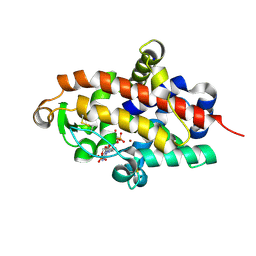 | | S1 nuclease from Aspergillus oryzae in complex with two molecules of 2'-deoxycytidine-5'-monophosphate | | Descriptor: | 2'-DEOXYCYTIDINE-5'-MONOPHOSPHATE, 2-acetamido-2-deoxy-beta-D-glucopyranose, Nuclease S1, ... | | Authors: | Koval, T, Oestergaard, L.H, Dohnalek, J. | | Deposit date: | 2015-12-14 | | Release date: | 2016-12-28 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | Structural and Catalytic Properties of S1 Nuclease from Aspergillus oryzae Responsible for Substrate Recognition, Cleavage, Non-Specificity, and Inhibition.
PLoS ONE, 11, 2016
|
|
9VRC
 
 | |
9VRD
 
 | |
9VRE
 
 | |
9VRB
 
 | |
5G3U
 
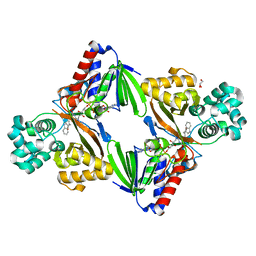 | | The structure of the L-tryptophan oxidase VioA from Chromobacterium violaceum in complex with its inhibitor 2-(1H-indol-3-ylmethyl)prop-2- enoic acid | | Descriptor: | 2-[(1H-indol-3-yl)methyl]prop-2-enoic acid, DIHYDROFLAVINE-ADENINE DINUCLEOTIDE, GLYCEROL, ... | | Authors: | Krausze, J, Rabe, J, Moser, J. | | Deposit date: | 2016-05-01 | | Release date: | 2016-08-03 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.377 Å) | | Cite: | Biosynthesis of Violacein, Structure and Function of l-Tryptophan Oxidase VioA from Chromobacterium violaceum.
J.Biol.Chem., 291, 2016
|
|
5G3S
 
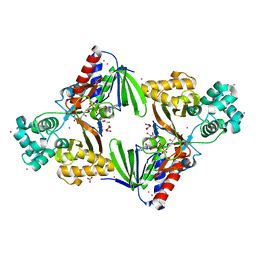 | | The structure of the L-tryptophan oxidase VioA from Chromobacterium violaceum - Samarium derivative | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, DIHYDROFLAVINE-ADENINE DINUCLEOTIDE, ... | | Authors: | Krausze, J, Rabe, J, Moser, J. | | Deposit date: | 2016-05-01 | | Release date: | 2016-08-03 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.076 Å) | | Cite: | Biosynthesis of Violacein: Structure and Function of L-Tryptophan Oxidase Vioa Chromobacterium Violaceum
J.Biol.Chem., 291, 2016
|
|
4WA2
 
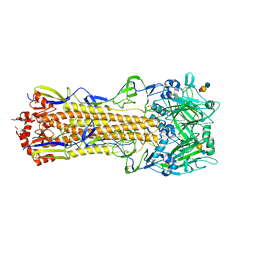 | | The crystal structure of hemagglutinin from a H3N8 influenza virus isolated from New England harbor seals in complex with 3'SLN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin, ... | | Authors: | Yang, H, Villanueva, J.M, Gubareva, L.V, Stevens, J. | | Deposit date: | 2014-08-28 | | Release date: | 2015-01-14 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural and Functional Analysis of Surface Proteins from an A(H3N8) Influenza Virus Isolated from New England Harbor Seals.
J.Virol., 89, 2015
|
|
4WA1
 
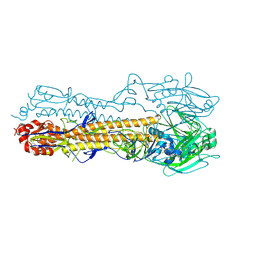 | | The crystal structure of hemagglutinin from a H3N8 influenza virus isolated from New England harbor seals | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin, ... | | Authors: | Yang, H, Villanueva, J.M, Gubareva, L.V, Stevens, J. | | Deposit date: | 2014-08-28 | | Release date: | 2015-01-14 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.898 Å) | | Cite: | Structural and Functional Analysis of Surface Proteins from an A(H3N8) Influenza Virus Isolated from New England Harbor Seals.
J.Virol., 89, 2015
|
|
5FR3
 
 | | X-ray crystal structure of aggregation-resistant protective antigen of Bacillus anthracis (mutant S559L T576E) | | Descriptor: | CALCIUM ION, GLYCEROL, PROTECTIVE ANTIGEN | | Authors: | Ganesan, A, Siekierska, A, Beerten, J, Brams, M, van Durme, J, De Baets, G, van der Kant, R, Gallardo, R, Ramakers, M, Langenberg, T, Wilkinson, H, De Smet, F, Ulens, C, Rousseau, F, Schymkowitz, J. | | Deposit date: | 2015-12-15 | | Release date: | 2016-01-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.935 Å) | | Cite: | Structural Hot Spots for the Solubility of Globular Proteins
Nat.Commun., 7, 2016
|
|
4V6K
 
 | | Structural insights into cognate vs. near-cognate discrimination during decoding. | | Descriptor: | 16S ribosomal RNA, 30S ribosomal protein S10, 30S ribosomal protein S11, ... | | Authors: | Agirrezabala, X, Schreiner, E, Trabuco, L.G, Lei, J, Ortiz-Meoz, R.F, Schulten, K, Green, R, Frank, J. | | Deposit date: | 2011-01-07 | | Release date: | 2014-07-09 | | Last modified: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (8.25 Å) | | Cite: | Structural insights into cognate versus near-cognate discrimination during decoding.
Embo J., 30, 2011
|
|
4V6L
 
 | | Structural insights into cognate vs. near-cognate discrimination during decoding. | | Descriptor: | 16S ribosomal RNA, 30S ribosomal protein S10, 30S ribosomal protein S11, ... | | Authors: | Agirrezabala, X, Schreiner, E, Trabuco, L.G, Lei, J, Ortiz-Meoz, R.F, Schulten, K, Green, R, Frank, J. | | Deposit date: | 2011-01-07 | | Release date: | 2014-07-09 | | Last modified: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (13.2 Å) | | Cite: | Structural insights into cognate versus near-cognate discrimination during decoding.
Embo J., 30, 2011
|
|
4V48
 
 | | Real space refined coordinates of the 30S and 50S subunits fitted into the low resolution cryo-EM map of the initiation-like state of E. coli 70S ribosome | | Descriptor: | 16S RIBOSOMAL RNA, 23S ribosomal RNA, 30S RIBOSOMAL PROTEIN S10, ... | | Authors: | Gao, H, Sengupta, J, Valle, M, Korostelev, A, Eswar, N, Stagg, S.M, Van Roey, P, Agrawal, R.K, Harvey, S.T, Sali, A, Chapman, M.S, Frank, J. | | Deposit date: | 2003-05-06 | | Release date: | 2014-07-09 | | Last modified: | 2024-02-28 | | Method: | ELECTRON MICROSCOPY (11.5 Å) | | Cite: | Study of the structural dynamics of the E. coli 70S ribosome using real space refinement
Cell(Cambridge,Mass.), 113, 2003
|
|
4WLA
 
 | |
4V47
 
 | | Real space refined coordinates of the 30S and 50S subunits fitted into the low resolution cryo-EM map of the EF-G.GTP state of E. coli 70S ribosome | | Descriptor: | 16S RIBOSOMAL RNA, 23S ribosomal RNA, 30S RIBOSOMAL PROTEIN S10, ... | | Authors: | Gao, H, Sengupta, J, Valle, M, Korostelev, A, Eswar, N, Stagg, S.M, Van Roey, P, Agrawal, R.K, Harvey, S.T, Sali, A, Chapman, M.S, Frank, J. | | Deposit date: | 2003-05-06 | | Release date: | 2014-07-09 | | Last modified: | 2024-02-28 | | Method: | ELECTRON MICROSCOPY (12.3 Å) | | Cite: | Study of the structural dynamics of the E. coli 70S ribosome using real space refinement
Cell(Cambridge,Mass.), 113, 2003
|
|
9YQ4
 
 | |
9Y4D
 | |
