3WAQ
 
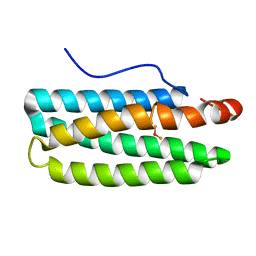 | | Hemerythrin-like domain of DcrH I119E mutant (met) | | Descriptor: | Hemerythrin-like domain protein DcrH, MU-OXO-DIIRON | | Authors: | Okamoto, Y, Onoda, A, Sugimoto, H, Takano, Y, Hirota, S, Kurtz Jr, D.M, Shiro, Y, Hayashi, T. | | Deposit date: | 2013-05-07 | | Release date: | 2014-03-19 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure, exogenous ligand binding, and redox properties of an engineered diiron active site in a bacterial hemerythrin
Inorg.Chem., 52, 2013
|
|
2EKT
 
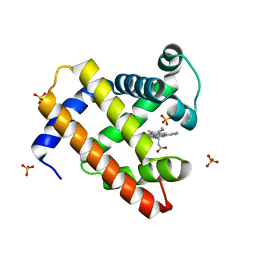 | | Crystal structure of myoglobin reconstituted with 6-methyl-6-depropionatehemin | | Descriptor: | 6-METHY-6-DEPROPIONATEHEMIN, Myoglobin, SULFATE ION | | Authors: | Harada, K, Makino, M, Sugimoto, H, Hirota, S, Matsuo, T, Shiro, Y, Hisaeda, Y, Hayashi, T. | | Deposit date: | 2007-03-25 | | Release date: | 2007-08-14 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Structure and ligand binding properties of myoglobins reconstituted with monodepropionated heme: functional role of each heme propionate side chain
Biochemistry, 46, 2007
|
|
2EKU
 
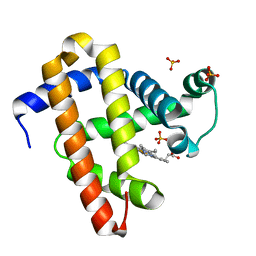 | | Crystal structure of myoglobin reconstituted with 7-methyl-7-depropionatehemin | | Descriptor: | 7-METHYL-7-DEPROPIONATEHEMIN, Myoglobin, SULFATE ION | | Authors: | Harada, K, Makino, M, Sugimoto, H, Hirota, S, Matsuo, T, Shiro, Y, Hisaeda, Y, Hayashi, T. | | Deposit date: | 2007-03-25 | | Release date: | 2007-08-14 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structure and ligand binding properties of myoglobins reconstituted with monodepropionated heme: functional role of each heme propionate side chain
Biochemistry, 46, 2007
|
|
5X94
 
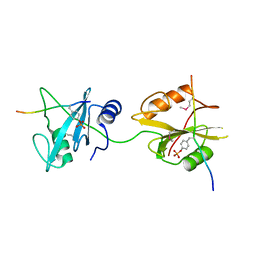 | | Crystal structure of SHP2_SH2-CagA EPIYA_D peptide complex | | Descriptor: | Cag pathogenicity island protein, Tyrosine-protein phosphatase non-receptor type 11 | | Authors: | Senda, M, Senda, T. | | Deposit date: | 2017-03-05 | | Release date: | 2017-09-13 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Differential Mechanisms for SHP2 Binding and Activation Are Exploited by Geographically Distinct Helicobacter pylori CagA Oncoproteins.
Cell Rep, 20, 2017
|
|
5X7B
 
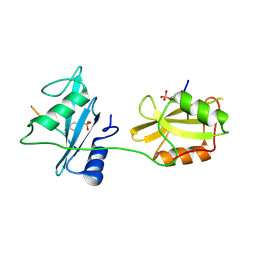 | | Crystal structure of SHP2_SH2-CagA EPIYA_C peptide complex | | Descriptor: | CagA, Tyrosine-protein phosphatase non-receptor type 11 | | Authors: | Senda, M, Senda, T. | | Deposit date: | 2017-02-24 | | Release date: | 2017-09-13 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Differential Mechanisms for SHP2 Binding and Activation Are Exploited by Geographically Distinct Helicobacter pylori CagA Oncoproteins.
Cell Rep, 20, 2017
|
|
3QHE
 
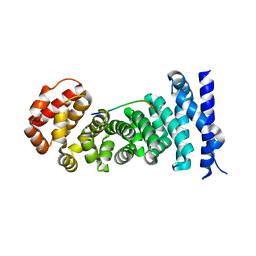 | | Crystal structure of the complex between the armadillo repeat domain of adenomatous polyposis coli and the tyrosine-rich domain of Sam68 | | Descriptor: | Adenomatous polyposis coli protein, KH domain-containing, RNA-binding, ... | | Authors: | Morishita, E.C.J, Murayama, K, Kato-Murayama, M, Ishizuku-Katsura, Y, Tomabechi, Y, Terada, T, Handa, N, Shirouzu, M, Akiyama, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2011-01-25 | | Release date: | 2011-11-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structures of the armadillo repeat domain of adenomatous polyposis coli and its complex with the tyrosine-rich domain of sam68
Structure, 19, 2011
|
|
6Z4R
 
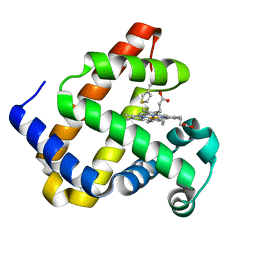 | |
6Z4T
 
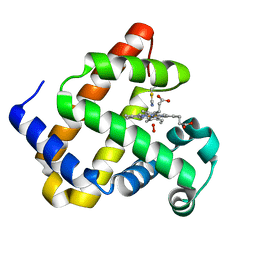 | |
5B35
 
 | | Serial Femtosecond Crystallography (SFX) of Ground State Bacteriorhodopsin Crystallized from Bicelles Determined Using 7-keV X-ray Free Electron Laser (XFEL) at SACLA | | Descriptor: | (3R,5S,7R,8R,9S,10S,12S,13R,14S,17R)-10,13-dimethyl-17-[(2R)-pentan-2-yl]-2,3,4,5,6,7,8,9,11,12,14,15,16,17-tetradecahydro-1H-cyclopenta[a]phenanthrene-3,7,12-triol, Bacteriorhodopsin, DECANE, ... | | Authors: | Mizohata, E, Nakane, T, Suzuki, M. | | Deposit date: | 2016-02-10 | | Release date: | 2016-11-09 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Membrane protein structure determination by SAD, SIR, or SIRAS phasing in serial femtosecond crystallography using an iododetergent
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
5B34
 
 | | Serial Femtosecond Crystallography (SFX) of Ground State Bacteriorhodopsin Crystallized from Bicelles in Complex with Iodine-labeled Detergent HAD13a Determined Using 7-keV X-ray Free Electron Laser (XFEL) at SACLA | | Descriptor: | 2,4,6-tris(iodanyl)-5-(octanoylamino)benzene-1,3-dicarboxylic acid, Bacteriorhodopsin, DECANE, ... | | Authors: | Mizohata, E, Nakane, T. | | Deposit date: | 2016-02-10 | | Release date: | 2016-11-09 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Membrane protein structure determination by SAD, SIR, or SIRAS phasing in serial femtosecond crystallography using an iododetergent
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
1QQY
 
 | |
1EL1
 
 | |
4J6D
 
 | |
4J6B
 
 | |
4J6C
 
 | |
4JBT
 
 | |
3AU3
 
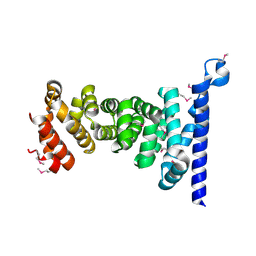 | | Crystal structure of armadillo repeat domain of APC | | Descriptor: | Adenomatous polyposis coli protein | | Authors: | Murayama, K, Kato-Murayama, M, Terada, T, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2011-01-28 | | Release date: | 2011-11-02 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structures of the armadillo repeat domain of adenomatous polyposis coli and its complex with the tyrosine-rich domain of sam68
Structure, 19, 2011
|
|
3ELL
 
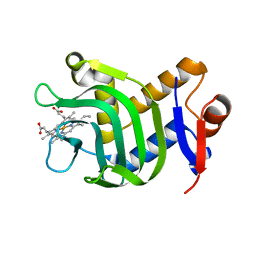 | | Structure of the hemophore from Pseudomonas aeruginosa (HasAp) | | Descriptor: | HasAp (Heme acquisition protein HasAp), PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Schonbrunn, E, Rivera, M. | | Deposit date: | 2008-09-22 | | Release date: | 2009-01-20 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural characterization of the hemophore HasAp from Pseudomonas aeruginosa: NMR spectroscopy reveals protein-protein interactions between Holo-HasAp and hemoglobin.
Biochemistry, 48, 2009
|
|
2ZB9
 
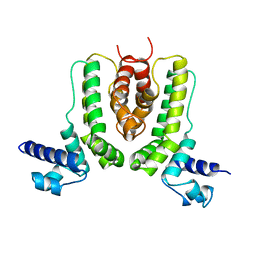 | | Crystal structure of TetR family transcription regulator SCO0332 | | Descriptor: | Putative transcriptional regulator | | Authors: | Okada, U, Kondo, K, Watanabe, N, Yao, M, Tanaka, I. | | Deposit date: | 2007-10-16 | | Release date: | 2008-01-29 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural and functional analysis of the TetR-family transcriptional regulator SCO0332 from Streptomyces coelicolor
ACTA CRYSTALLOGR.,SECT.D, 64, 2008
|
|
2ZCX
 
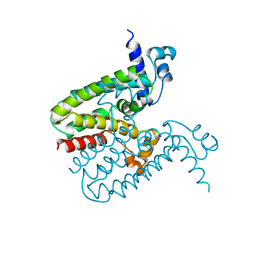 | |
2ZDS
 
 | | Crystal Structure of SCO6571 from Streptomyces coelicolor A3(2) | | Descriptor: | Putative DNA-binding protein | | Authors: | Begum, P, Gao, Y.G, Sakai, N, Yao, M, Watanabe, N, Tanaka, I. | | Deposit date: | 2007-11-27 | | Release date: | 2008-12-02 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of SCO6571 from Streptomyces coelicolor A3(2).
Protein Pept.Lett., 15, 2008
|
|
8AUT
 
 | | WelO5* L221A bound to Zn(II), Cl, 2-oxoglutarate, and 12-epi-hapalindole C | | Descriptor: | 2-OXOGLUTARIC ACID, 3-[(1~{S},2~{R},3~{S},6~{S})-3-ethenyl-2-isocyano-3-methyl-6-prop-1-en-2-yl-cyclohexyl]-1~{H}-indole, CHLORIDE ION, ... | | Authors: | Buller, R, Hueppi, S, Voss, M, Schaub, D. | | Deposit date: | 2022-08-25 | | Release date: | 2022-11-02 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.685 Å) | | Cite: | Enzyme engineering enables inversion of substrate stereopreference of the halogenase WelO5*
Chemcatchem, 2022
|
|
