101D
 
 | |
1LEY
 
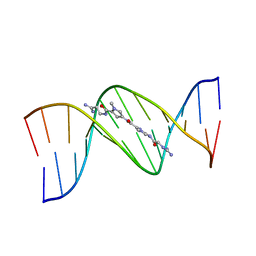 | | STRUCTURE OF A DICATIONIC MONOIMIDAZOLE LEXITROPSIN BOUND TO DNA (ORIENTATION 2) | | Descriptor: | DNA (5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3'), MONOIMIDAZOLE LEXITROPSIN | | Authors: | Goodsell, D.S, Ng, H.L, Kopka, M.L, Lown, J.W, Dickerson, R.E. | | Deposit date: | 1995-10-10 | | Release date: | 1996-04-03 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure of a dicationic monoimidazole lexitropsin bound to DNA.
Biochemistry, 34, 1995
|
|
1LEX
 
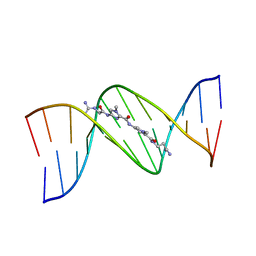 | | STRUCTURE OF A DICATIONIC MONOIMIDAZOLE LEXITROPSIN BOUND TO DNA (ORIENTATION 1) | | Descriptor: | DNA (5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3'), MONOIMIDAZOLE LEXITROPSIN | | Authors: | Goodsell, D.S, Ng, H.L, Kopka, M.L, Lown, J.W, Dickerson, R.E. | | Deposit date: | 1995-10-10 | | Release date: | 1996-04-03 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure of a dicationic monoimidazole lexitropsin bound to DNA.
Biochemistry, 34, 1995
|
|
126D
 
 | |
196D
 
 | |
167D
 | |
274D
 
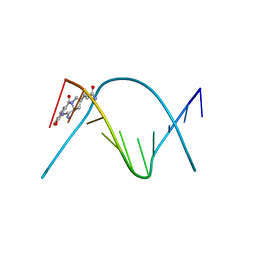 | | CRYSTAL STRUCTURE OF A COVALENT DNA-DRUG ADDUCT: ANTHRAMYCIN BOUND TO C-C-A-A-C-G-T-T-G-G, AND A MOLECULAR EXPLANATION OF SPECIFICITY | | Descriptor: | ANTHRAMYCIN, DNA (5'-D(*CP*CP*AP*AP*CP*GP*TP*TP*(DRUG)GP*G)-3') | | Authors: | Kopka, M.L, Goodsell, D.S, Baikalov, I, Grzeskowiak, K, Cascio, D, Dickerson, R.E. | | Deposit date: | 1994-04-28 | | Release date: | 1994-10-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of a covalent DNA-drug adduct: anthramycin bound to C-C-A-A-C-G-T-T-G-G and a molecular explanation of specificity.
Biochemistry, 33, 1994
|
|
334D
 
 | |
1ZP8
 
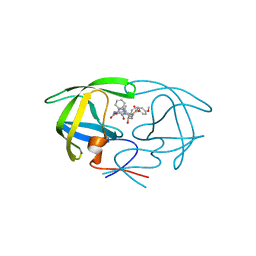 | | HIV Protease with inhibitor AB-2 | | Descriptor: | Pol polyprotein, [1-((1S,2R)-1-BENZYL-2-HYDROXY-3-{ISOBUTYL[(4-METHOXYPHENYL)SULFONYL]AMINO}PROPYL)-1H-1,2,3-TRIAZOL-4-YL]METHYL (1R,2R)-2-HYDROXY-2,3-DIHYDRO-1H-INDEN-1-YLCARBAMATE | | Authors: | Brik, A, Alexandratos, J.N, Elder, J.H, Olson, A.J, Wlodawer, A, Goodsell, D.S, Wong, C.H. | | Deposit date: | 2005-05-16 | | Release date: | 2005-06-21 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | 1,2,3-triazole as a peptide surrogate in the rapid synthesis of HIV-1 protease inhibitors.
Chembiochem, 6, 2005
|
|
1ZPA
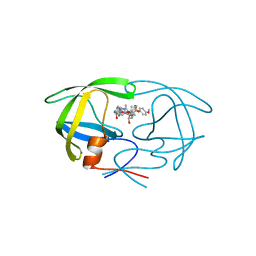 
 | | HIV Protease with Scripps AB-3 Inhibitor | | Descriptor: | Pol polyprotein, TERT-BUTYL 4-[({[1-((1S,2R)-1-BENZYL-2-HYDROXY-3-{ISOBUTYL[(4-METHOXYPHENYL)SULFONYL]AMINO}PROPYL)-1H-1,2,3-TRIAZOL-4-YL]METHYL}AMINO)CARBONYL]BENZYLCARBAMATE | | Authors: | Brik, A, Alexandratos, J, Lin, Y.C, Elder, J.H, Olson, A.J, Wlodawer, A, Goodsell, D.S, Wong, C.H. | | Deposit date: | 2005-05-16 | | Release date: | 2005-05-31 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | 1,2,3-triazole as a peptide surrogate in the rapid synthesis of HIV-1 protease inhibitors
Chembiochem, 6, 2005
|
|
158D
 
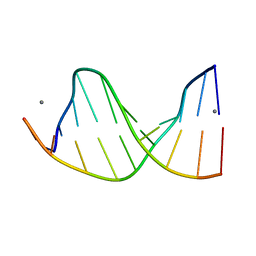 | | CRYSTALLOGRAPHIC ANALYSIS OF C-C-A-A-G-C-T-T-G-G AND ITS IMPLICATIONS FOR BENDING IN B-DNA | | Descriptor: | CALCIUM ION, DNA (5'-D(*CP*CP*AP*AP*GP*CP*TP*TP*GP*G)-3') | | Authors: | Grzeskowiak, K, Goodsell, D.S, Kaczor-Grzeskowiak, M, Cascio, D, Dickerson, R.E. | | Deposit date: | 1994-02-03 | | Release date: | 1994-05-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystallographic analysis of C-C-A-A-G-C-T-T-G-G and its implications for bending in B-DNA.
Biochemistry, 32, 1993
|
|
1D30
 
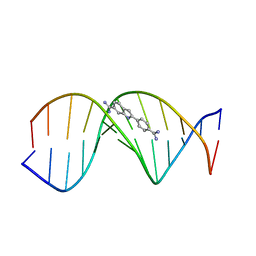 | | THE STRUCTURE OF DAPI BOUND TO DNA | | Descriptor: | 6-AMIDINE-2-(4-AMIDINO-PHENYL)INDOLE, DNA (5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3') | | Authors: | Larsen, T, Goodsell, D.S, Cascio, D, Grzeskowiak, K, Dickerson, R.E. | | Deposit date: | 1991-01-04 | | Release date: | 1992-04-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The structure of DAPI bound to DNA.
J.Biomol.Struct.Dyn., 7, 1989
|
|
8BNA
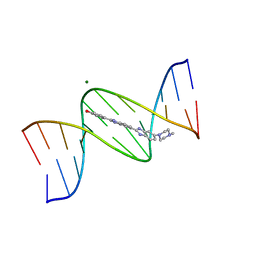 
 | | BINDING OF HOECHST 33258 TO THE MINOR GROOVE OF B-DNA | | Descriptor: | 2'-(4-HYDROXYPHENYL)-5-(4-METHYL-1-PIPERAZINYL)-2,5'-BI-BENZIMIDAZOLE, DNA (5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3'), MAGNESIUM ION | | Authors: | Pjura, P.E, Grzeskowiak, K, Dickerson, R.E. | | Deposit date: | 1986-08-29 | | Release date: | 1987-01-15 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Binding of Hoechst 33258 to the minor groove of B-DNA.
J.Mol.Biol., 197, 1987
|
|
8F2W
 
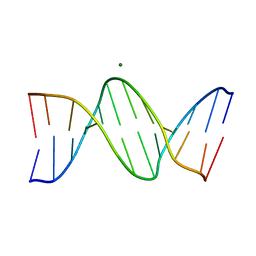 | | Structure of a B-Form Dodecamer: 5'-CGCGAATTCGCG-3 | | Descriptor: | DNA (5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3'), MAGNESIUM ION | | Authors: | Ogbonna, E.N, Wilson, W.D. | | Deposit date: | 2022-11-08 | | Release date: | 2023-02-22 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | X-ray Structure Characterization of the Selective Recognition of AT Base Pair Sequences.
Acs Bio Med Chem Au, 3, 2023
|
|
5W4P
 
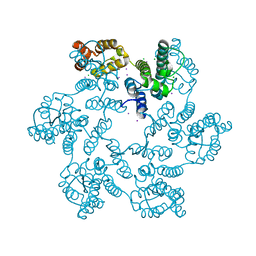 | |
5W4O
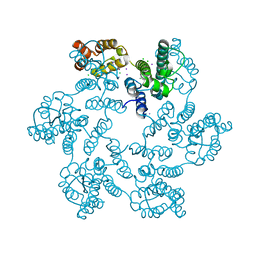 
 | |
5W4Q
 
 | |
1DN9
 
 | |
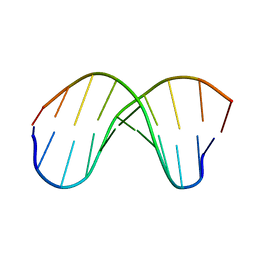
252D
 | |
