8PE0
 
 | | X-ray structure of the Thermus thermophilus K167L mutant of the PilF-GSPIIB domain in the c-di-GMP bound state | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), ACETATE ION, SULFATE ION, ... | | Authors: | Neissner, K, Woehnert, J. | | Deposit date: | 2023-06-13 | | Release date: | 2024-06-26 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | X-ray structure of the K167L mutant of PilF-GSPIIB domain in the c-di-GMP bound state
To Be Published
|
|
8PDK
 
 | | X-ray structure of the Thermus thermophilus PilF-GSPIIB domain in the c-di-GMP bound state | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), ACETATE ION, SULFATE ION, ... | | Authors: | Neissner, K, Woehnert, J. | | Deposit date: | 2023-06-12 | | Release date: | 2024-06-26 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | X-ray structure of the PilF-GSPIIB domain in the c-di-GMP bound state
To Be Published
|
|
8PFA
 
 | | X-ray structure of the Thermus thermophilus K167R mutant of the PilF-GSPIIB domain in the c-di-GMP bound state | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), ACETATE ION, SULFATE ION, ... | | Authors: | Neissner, K, Woehnert, J. | | Deposit date: | 2023-06-15 | | Release date: | 2024-06-26 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray structure of the Thermus thermophilus K167R mutant of the PilF-GSPIIB domain in the c-di-GMP bound state
To Be Published
|
|
3U9C
 
 | | Structure of a C-terminal deletion mutant of human protein kinase CK2 catalytic subunit with the ATP-competitive inhibitor resorufin | | Descriptor: | 7-hydroxy-3H-phenoxazin-3-one, CHLORIDE ION, Casein kinase II subunit alpha, ... | | Authors: | Klopffleisch, K, Issinger, O.-G, Niefind, K. | | Deposit date: | 2011-10-18 | | Release date: | 2012-05-30 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Low-density crystal packing of human protein kinase CK2 catalytic subunit in complex with resorufin or other ligands: a tool to study the unique hinge-region plasticity of the enzyme without packing bias.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
8PQU
 
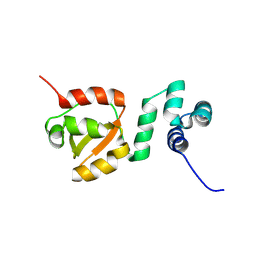 | |
8PKZ
 
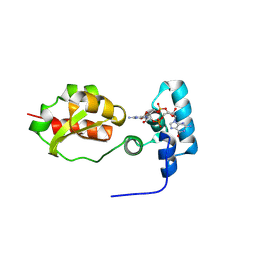 | |
4GU3
 
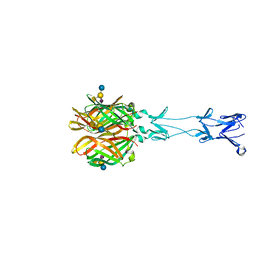 | |
4GU4
 
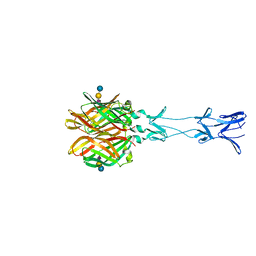 | |
7Z6F
 
 | |
1WAM
 
 | |
1OC2
 
 | |
1V0J
 
 | | Udp-galactopyranose mutase from Mycobacterium tuberculosis | | Descriptor: | BICINE, FLAVIN-ADENINE DINUCLEOTIDE, UDP-GALACTOPYRANOSE MUTASE | | Authors: | Beis, K, Naismith, J.H. | | Deposit date: | 2004-03-30 | | Release date: | 2005-01-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Crystal structures of Mycobacteria tuberculosis and Klebsiella pneumoniae UDP-galactopyranose mutase in the oxidised state and Klebsiella pneumoniae UDP-galactopyranose mutase in the (active) reduced state.
J. Mol. Biol., 348, 2005
|
|
2BI8
 
 | |
2BI7
 
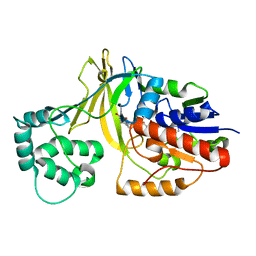 | | udp-galactopyranose mutase from Klebsiella pneumoniae oxidised FAD | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, UDP-GALACTOPYRANOSE MUTASE | | Authors: | Beis, K, Srikannathasan, V, Naismith, J. | | Deposit date: | 2005-01-20 | | Release date: | 2005-05-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structures of Mycobacteria Tuberculosis and Klebsiella Pneumoniae Udp-Galactopyranose Mutase in the Oxidised State and Klebsiella Pneumoniae Udp-Galactopyranose Mutase in the (Active) Reduced State.
J.Mol.Biol., 348, 2005
|
|
1D9X
 
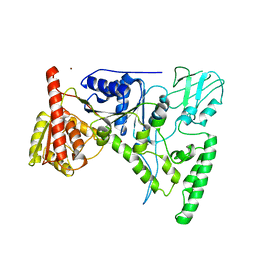 | | CRYSTAL STRUCTURE OF THE DNA REPAIR PROTEIN UVRB | | Descriptor: | EXCINUCLEASE UVRABC COMPONENT UVRB, ZINC ION | | Authors: | Theis, K, Chen, P.J, Skorvaga, M, Van Houten, B, Kisker, C. | | Deposit date: | 1999-10-30 | | Release date: | 2000-05-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of UvrB, a DNA helicase adapted for nucleotide excision repair.
EMBO J., 18, 1999
|
|
1D9Z
 
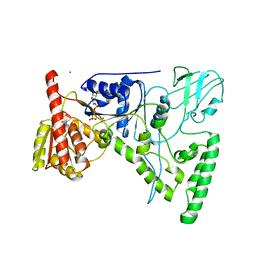 | | CRYSTAL STRUCTURE OF THE DNA REPAIR PROTEIN UVRB IN COMPLEX WITH ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, EXCINUCLEASE UVRABC COMPONENT UVRB, MAGNESIUM ION, ... | | Authors: | Theis, K, Chen, P.J, Skorvaga, M, Van Houten, B, Kisker, C. | | Deposit date: | 1999-10-30 | | Release date: | 2000-05-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Crystal structure of UvrB, a DNA helicase adapted for nucleotide excision repair.
EMBO J., 18, 1999
|
|
3TW5
 
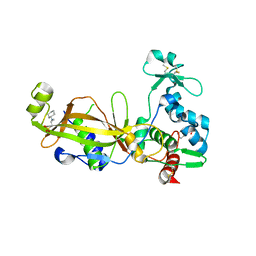 | | Crystal structure of the GP42 transglutaminase from Phytophthora sojae | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, Transglutaminase elicitor | | Authors: | Reiss, K, Kirchner, E, Zocher, G, Stehle, T. | | Deposit date: | 2011-09-21 | | Release date: | 2011-10-12 | | Last modified: | 2020-10-21 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural and Phylogenetic Analyses of the GP42 Transglutaminase from Phytophthora sojae Reveal an Evolutionary Relationship between Oomycetes and Marine Vibrio Bacteria.
J.Biol.Chem., 286, 2011
|
|
6STL
 
 | | Taurine ABC transporter substrate binding protein TauA from E. coli in complex with taurine | | Descriptor: | 2-AMINOETHANESULFONIC ACID, IODIDE ION, Taurine-binding periplasmic protein | | Authors: | Beis, K, Qu, F, Wagner, A, ElOmari, K. | | Deposit date: | 2019-09-10 | | Release date: | 2019-12-18 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Desolvation of the substrate-binding protein TauA dictates ligand specificity for the alkanesulfonate ABC importer TauABC.
Biochem.J., 476, 2019
|
|
6ST1
 
 | |
6ST0
 
 | |
6SSY
 
 | |
6YEP
 
 | |
6YCV
 
 | |
6RCP
 
 | |
6RD3
 
 | |
