3NDS
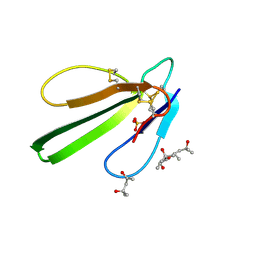 
 | | Crystal structure of engineered Naja Nigricollis toxin alpha | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, SULFATE ION, Short neurotoxin 1 | | Authors: | Stura, E.A, Drevet, P, Gregory, G, Gallopin, M, Vandame, M. | | Deposit date: | 2010-06-08 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Conformational exchange is critical for the productivity of an oxidative folding intermediate with buried free cysteines.
J.Mol.Biol., 403, 2010
|
|
2ERA
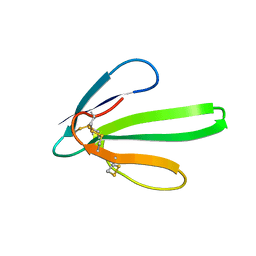 
 | | RECOMBINANT ERABUTOXIN A, S8G MUTANT | | Descriptor: | ERABUTOXIN A | | Authors: | Gaucher, J.F, Menez, R, Arnoux, B, Menez, A, Ducruix, A. | | Deposit date: | 1997-06-25 | | Release date: | 1997-12-31 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | High resolution x-ray analysis of two mutants of a curaremimetic snake toxin
Eur.J.Biochem., 267, 2000
|
|
3ERA
 
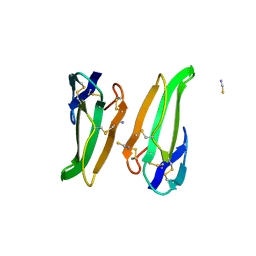 | | RECOMBINANT ERABUTOXIN A (S8T MUTANT) | | Descriptor: | ERABUTOXIN A, THIOCYANATE ION | | Authors: | Gaucher, J.F, Menez, R, Arnoux, B, Menez, A, Ducruix, A. | | Deposit date: | 1997-06-25 | | Release date: | 1997-12-31 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | High resolution x-ray analysis of two mutants of a curaremimetic snake toxin
Eur.J.Biochem., 267, 2000
|
|
3Q4F
 
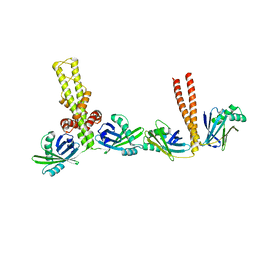 | | Crystal structure of xrcc4/xlf-cernunnos complex | | Descriptor: | DNA repair protein XRCC4, Non-homologous end-joining factor 1 | | Authors: | Ropars, V, Legrand, P, Charbonnier, J.B. | | Deposit date: | 2010-12-23 | | Release date: | 2011-08-03 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (5.5 Å) | | Cite: | Structural characterization of filaments formed by human Xrcc4-Cernunnos/XLF complex involved in nonhomologous DNA end-joining.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
6ERG
 
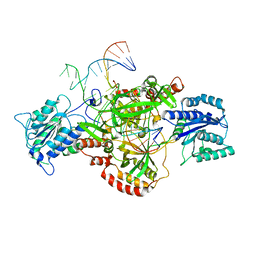 | | Complex of XLF and heterodimer Ku bound to DNA | | Descriptor: | DNA (21-MER), DNA (34-MER), Non-homologous end-joining factor 1, ... | | Authors: | Nemoz, C, Legrand, P, Ropars, V, Charbonnier, J.B. | | Deposit date: | 2017-10-18 | | Release date: | 2018-10-17 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | XLF and APLF bind Ku80 at two remote sites to ensure DNA repair by non-homologous end joining.
Nat. Struct. Mol. Biol., 25, 2018
|
|
6ERH
 
 | | Complex of XLF and heterodimer Ku bound to DNA | | Descriptor: | DNA (21-MER), DNA (34-MER), Non-homologous end-joining factor 1, ... | | Authors: | Nemoz, C, Legrand, P, Ropars, V, Charbonnier, J.B. | | Deposit date: | 2017-10-18 | | Release date: | 2018-10-17 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | XLF and APLF bind Ku80 at two remote sites to ensure DNA repair by non-homologous end joining.
Nat. Struct. Mol. Biol., 25, 2018
|
|
6ERF
 
 | | Complex of APLF factor and Ku heterodimer bound to DNA | | Descriptor: | Aprataxin and PNK-like factor, DNA (34-MER), DNA (5'-D(*GP*TP*TP*TP*TP*TP*AP*GP*TP*TP*TP*AP*TP*TP*GP*GP*GP*CP*GP*CP*G)-3'), ... | | Authors: | Nemoz, C, Legrand, P, Ropars, V, Charbonnier, J.B. | | Deposit date: | 2017-10-18 | | Release date: | 2018-10-17 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.01 Å) | | Cite: | XLF and APLF bind Ku80 at two remote sites to ensure DNA repair by non-homologous end joining.
Nat.Struct.Mol.Biol., 25, 2018
|
|
