9JYW
 
 | |
8YR4
 
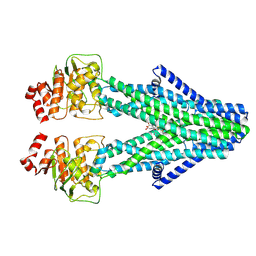 | | Cryo-EM structure of the human ABCB6 in complex with Cd(II):Phytochelatin 2 | | Descriptor: | ATP-binding cassette sub-family B member 6, CADMIUM ION, Phytochelatin 2 | | Authors: | Choi, S.H, Lee, S.S, Lee, H.Y, Kim, S, Kim, J.W, Jin, M.S. | | Deposit date: | 2024-03-20 | | Release date: | 2024-06-19 | | Last modified: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Cryo-EM structure of cadmium-bound human ABCB6.
Commun Biol, 7, 2024
|
|
8YR3
 
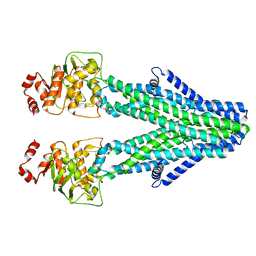 | | Cryo-EM structure of the human ABCB6 in complex with Cd(II):GSH | | Descriptor: | ATP-binding cassette sub-family B member 6, CADMIUM ION, GLUTATHIONE | | Authors: | Choi, S.H, Lee, S.S, Lee, H.Y, Kim, S, Kim, J.W, Jin, M.S. | | Deposit date: | 2024-03-20 | | Release date: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Cryo-EM structure of cadmium-bound human ABCB6.
Commun Biol, 7, 2024
|
|
5X3D
 
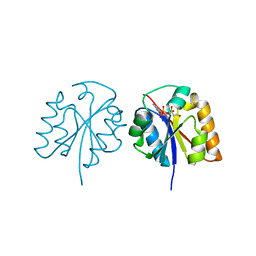 | | Crystal structure of HEP-CMP-bound form of cytidylyltransferase (CyTase) domain of Fom1 from Streptomyces wedmorensis | | Descriptor: | Phosphoenolpyruvate phosphomutase, [[(2R,3S,4R,5R)-5-(4-azanyl-2-oxidanylidene-pyrimidin-1-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl]oxy-(2-hydroxyethyl)phosphinic acid | | Authors: | Tomita, T, Cho, S.H, Kuzuyama, T, Nishiyama, M. | | Deposit date: | 2017-02-04 | | Release date: | 2017-09-13 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Fosfomycin Biosynthesis via Transient Cytidylylation of 2-Hydroxyethylphosphonate by the Bifunctional Fom1 Enzyme
ACS Chem. Biol., 12, 2017
|
|
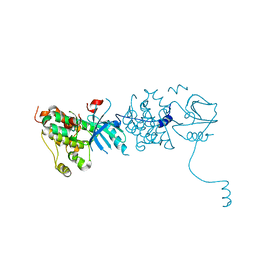
3GOP
 
 | |
2FUJ
 
 | |
2FUK
 
 | | Crystal structure of XC6422 from Xanthomonas campestris: a member of a/b serine hydrolase without lid at 1.6 resolution | | Descriptor: | XC6422 protein | | Authors: | Yang, C.Y, Chin, K.H, Chou, C.C, Wang, A.H.J, Chou, S.H. | | Deposit date: | 2006-01-27 | | Release date: | 2006-07-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure of XC6422 from Xanthomonas campestris at 1.6 A resolution: a small serine alpha/beta-hydrolase
Acta Crystallogr.,Sect.F, 62, 2006
|
|
9LSI
 
 | | Cryo-EM structure of the S82C-S82C diabody complex (CitS-diabody #2-TLR3) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Kim, S, Kim, J.W, Park, J.G, Lee, S.S, Choi, S.H, Lee, J.-O, Jin, M.S. | | Deposit date: | 2025-02-04 | | Release date: | 2025-03-12 | | Last modified: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Disulfide-stabilized diabodies enable near-atomic cryo-EM imaging of small proteins: A case study of the bacterial Na + /citrate symporter CitS.
Structure, 2025
|
|
9LSK
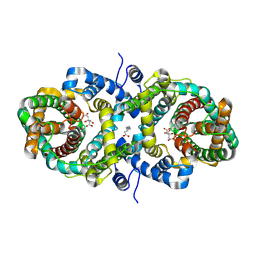 
 | | Cryo-EM structure of the Klebsiella pneumoniae CitS (citrate-bound occluded state) | | Descriptor: | CITRIC ACID, Citrate/sodium symporter, PALMITIC ACID | | Authors: | Kim, S, Kim, J.W, Park, J.G, Lee, S.S, Choi, S.H, Lee, J.-O, Jin, M.S. | | Deposit date: | 2025-02-04 | | Release date: | 2025-03-12 | | Last modified: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Disulfide-stabilized diabodies enable near-atomic cryo-EM imaging of small proteins: A case study of the bacterial Na + /citrate symporter CitS.
Structure, 2025
|
|
9LSH
 
 | | Cryo-EM structure of the wild-type diabody complex (CitS-diabody #1-TLR3) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Kim, S, Kim, J.W, Park, J.G, Lee, S.S, Choi, S.H, Lee, J.-O, Jin, M.S. | | Deposit date: | 2025-02-04 | | Release date: | 2025-03-12 | | Last modified: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Disulfide-stabilized diabodies enable near-atomic cryo-EM imaging of small proteins: A case study of the bacterial Na + /citrate symporter CitS.
Structure, 2025
|
|
9LSJ
 
 | | Cryo-EM structure of the G15C-R66C and T83C-T83C diabody complex (CitS-diabody #7-TLR3) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CITRIC ACID, ... | | Authors: | Kim, S, Kim, J.W, Park, J.G, Lee, S.S, Choi, S.H, Lee, J.-O, Jin, M.S. | | Deposit date: | 2025-02-04 | | Release date: | 2025-03-12 | | Last modified: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Disulfide-stabilized diabodies enable near-atomic cryo-EM imaging of small proteins: A case study of the bacterial Na + /citrate symporter CitS.
Structure, 2025
|
|
4YP1
 
 | | Misting with CDA | | Descriptor: | (2R,3R,3aS,5R,7aR,9R,10R,10aS,12R,14aR)-2,9-bis(6-amino-9H-purin-9-yl)octahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8 ]tetraoxadiphosphacyclododecine-3,5,10,12-tetrol 5,12-dioxide, Stimulator of interferon genes protein | | Authors: | Chin, K.H, Chen, C.K, Tu, Z.I, Chou, S.H. | | Deposit date: | 2015-03-12 | | Release date: | 2015-10-07 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structural Insights into the Distinct Binding Mode of Cyclic Di-AMP with SaCpaA_RCK.
Biochemistry, 54, 2015
|
|
4YS2
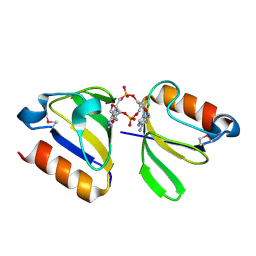 
 | | RCK domain with CDA | | Descriptor: | (2R,3R,3aS,5R,7aR,9R,10R,10aS,12R,14aR)-2,9-bis(6-amino-9H-purin-9-yl)octahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8 ]tetraoxadiphosphacyclododecine-3,5,10,12-tetrol 5,12-dioxide, Na+/H+ antiporter-like protein | | Authors: | Chin, K.H, Chou, S.H. | | Deposit date: | 2015-03-16 | | Release date: | 2016-04-27 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.968 Å) | | Cite: | RCK domain with CDA
To Be Published
|
|
4E6S
 
 | | Crystal structure of the SCAN domain from mouse Zfp206 | | Descriptor: | Zinc finger and SCAN domain-containing protein 10 | | Authors: | Liang, Y, Choo, S.H, Rossbach, M, Baburajendran, N, Palasingam, P, Kolatkar, P.R. | | Deposit date: | 2012-03-15 | | Release date: | 2012-05-02 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal optimization and preliminary diffraction data analysis of the SCAN domain of Zfp206.
Acta Crystallogr.,Sect.F, 68, 2012
|
|
2FA5
 
 | | The crystal structure of an unliganded multiple antibiotic-resistance repressor (MarR) from Xanthomonas campestris | | Descriptor: | CHLORIDE ION, transcriptional regulator marR/emrR family | | Authors: | Chin, K.H, Tu, Z.L, Li, J.N, Chou, C.C, Wang, A.H.J, Chou, S.H. | | Deposit date: | 2005-12-06 | | Release date: | 2006-11-14 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The crystal structure of XC1739: a putative multiple antibiotic-resistance repressor (MarR) from Xanthomonas campestris at 1.8 A resolution
Proteins, 65, 2006
|
|
2GBZ
 
 | | The Crystal Structure of XC847 from Xanthomonas campestris: a 3-5 Oligoribonuclease of DnaQ fold family with a Novel Opposingly-Shifted Helix | | Descriptor: | MAGNESIUM ION, Oligoribonuclease | | Authors: | Chin, K.H, Yang, C.Y, Chou, C.C, Wang, A.H.J, Chou, S.H. | | Deposit date: | 2006-03-12 | | Release date: | 2007-01-16 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The crystal structure of XC847 from Xanthomonas campestris: a 3'-5' oligoribonuclease of DnaQ fold family with a novel opposingly shifted helix
Proteins, 65, 2006
|
|
1C11
 
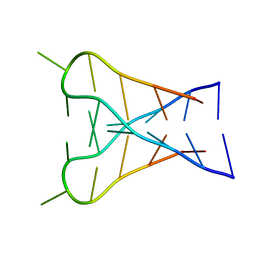 | | INTERCALATED D(TCCCGTTTCCA) DIMER, NMR, 7 STRUCTURES | | Descriptor: | DNA (5'-D(*TP*CP*CP*CP*GP*TP*TP*TP*CP*CP*A)-3') | | Authors: | Gallego, J, Chou, S.H, Reid, B.R. | | Deposit date: | 1998-07-15 | | Release date: | 1998-07-22 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Centromeric pyrimidine strands fold into an intercalated motif by forming a double hairpin with a novel T:G:G:T tetrad: solution structure of the d(TCCCGTTTCCA) dimer.
J.Mol.Biol., 273, 1997
|
|
2XSD
 
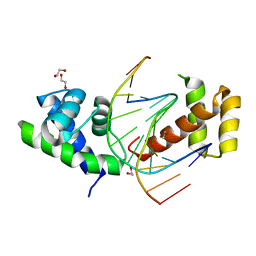 | | Crystal Structure of the dimeric Oct-6 (Pou3f1) POU domain bound to palindromic MORE DNA | | Descriptor: | 1,2-ETHANEDIOL, 5'-D(*AP*TP*GP*CP*AP*TP*GP*AP*GP*GP*AP)-3', 5'-D(*CP*CP*TP*CP*AP*TP*GP*CP*AP*TP*AP)-3', ... | | Authors: | Jauch, R, Choo, S.H, Ng, C.K.L, Kolatkar, P.R. | | Deposit date: | 2010-09-28 | | Release date: | 2010-10-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.049 Å) | | Cite: | Crystal Structure of the Dimeric Oct6 (POU3F1) POU Domain Bound to Palindromic More DNA.
Proteins, 79, 2011
|
|
2ZE3
 
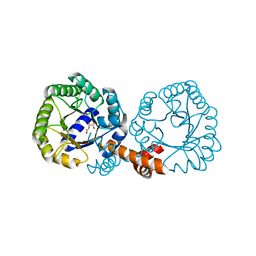 | |
1P0U
 
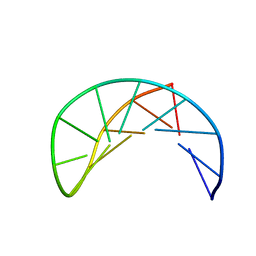 | | Sheared G/C Base Pair | | Descriptor: | 5'-D(*GP*CP*AP*TP*CP*GP*AP*CP*GP*AP*TP*GP*C)-3' | | Authors: | Chin, K.H, Chou, S.H. | | Deposit date: | 2003-04-11 | | Release date: | 2003-06-03 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Sheared-type G(anti).C(syn) Base-pair: A Unique d(GXC) Loop Closure Motif.
J.Mol.Biol., 329, 2003
|
|
3DSG
 
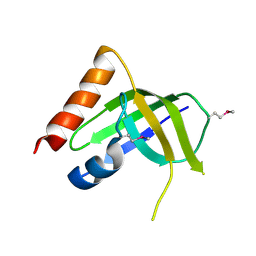 | | XC1028 from Xanthomonas campestris Adopts a PilZ Domain-like Structure Yet with Trivial c-di-GMP Binding Activity | | Descriptor: | Type IV fimbriae assembly protein | | Authors: | Li, T.N, Chin, K.H, Liu, J.H, Wang, A.H.J, Chou, S.H. | | Deposit date: | 2008-07-12 | | Release date: | 2009-05-19 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | XC1028 from Xanthomonas campestris adopts a PilZ domain-like structure without a c-di-GMP switch.
Proteins, 75, 2009
|
|
2GSC
 
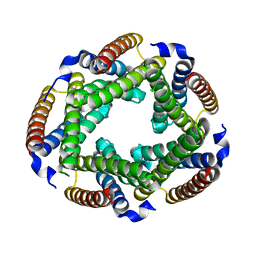 | | Crystal Structure of the Conserved Hypothetical Cytosolic Protein Xcc0516 from Xanthomonas campestris | | Descriptor: | Putative uncharacterized protein XCC0516 | | Authors: | Lin, L.Y, Ching, C.L, Chin, K.H, Chou, S.H, Chan, N.L. | | Deposit date: | 2006-04-26 | | Release date: | 2006-10-03 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Crystal structure of the conserved hypothetical cytosolic protein Xcc0516 from Xanthomonas campestris reveals a novel quaternary structure assembled by five four-helix bundles.
Proteins, 65, 2006
|
|
8ZL3
 
 | | Cryo-EM structure of the Drosophila INDY (DIDS-bound outward-open, pH 6) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4,4'-Diisothiocyano-2,2'-stilbenedisulfonic acid, PHOSPHATIDYLETHANOLAMINE, ... | | Authors: | Kim, S, Park, J.G, Choi, S.H, Kim, J.W, Jin, M.S. | | Deposit date: | 2024-05-17 | | Release date: | 2025-01-29 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Cryo-EM structures reveal the cation-independent citrate transport mechanism of the Drosophila longevity protein, INDY
To Be Published
|
|
8ZKW
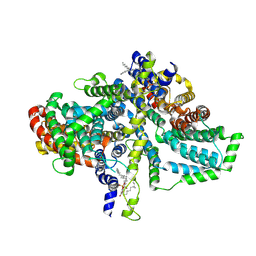 
 | | Cryo-EM structure of the Drosophila INDY (apo-asymmetric, pH 6) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, PHOSPHATIDYLETHANOLAMINE, Protein I'm not dead yet | | Authors: | Kim, S, Park, J.G, Choi, S.H, Kim, J.W, Jin, M.S. | | Deposit date: | 2024-05-17 | | Release date: | 2025-01-29 | | Method: | ELECTRON MICROSCOPY (2.77 Å) | | Cite: | Cryo-EM structures reveal the cation-independent citrate transport mechanism of the Drosophila longevity protein, INDY
To Be Published
|
|
8ZL6
 
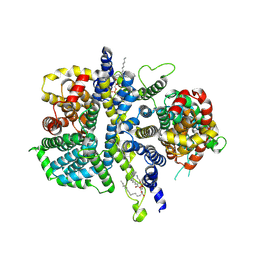 | | Cryo-EM structure of the Drosophila INDY (apo inward-open, pH 8) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, PHOSPHATIDYLETHANOLAMINE, Protein I'm not dead yet | | Authors: | Kim, S, Park, J.G, Choi, S.H, Kim, J.W, Jin, M.S. | | Deposit date: | 2024-05-17 | | Release date: | 2025-01-29 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Cryo-EM structures reveal the cation-independent citrate transport mechanism of the Drosophila longevity protein, INDY
To Be Published
|
|
