8PTG
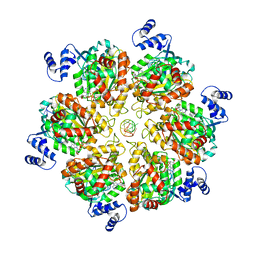 
 | | Structure of the transcription termination factor Rho bound to RNA at the PBS and SBS | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, BERYLLIUM TRIFLUORIDE ION, MAGNESIUM ION, ... | | Authors: | Said, N, Hilal, T, Wahl, M.C. | | Deposit date: | 2023-07-14 | | Release date: | 2024-04-17 | | Last modified: | 2024-04-24 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Sm-like protein Rof inhibits transcription termination factor rho by binding site obstruction and conformational insulation.
Nat Commun, 15, 2024
|
|
8PTO
 
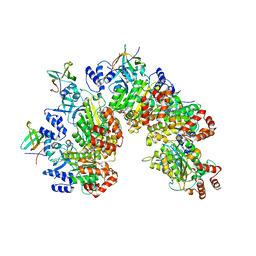 | | Structure of Rho pentamer in complex with Rof and ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Protein rof, Transcription termination factor Rho | | Authors: | Said, N, Hilal, T, Wahl, M.C. | | Deposit date: | 2023-07-14 | | Release date: | 2024-04-17 | | Last modified: | 2024-04-24 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Sm-like protein Rof inhibits transcription termination factor rho by binding site obstruction and conformational insulation.
Nat Commun, 15, 2024
|
|
8BCE
 
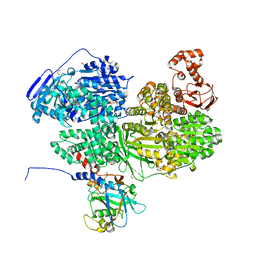 | | Human Brr2 Helicase Region in complex with C-tail deleted Jab1 and compound 76 | | Descriptor: | 1,2-ETHANEDIOL, N-methoxybenzenesulfonamide, Pre-mRNA-processing-splicing factor 8, ... | | Authors: | Vester, K, Loll, B, Wahl, M.C. | | Deposit date: | 2022-10-15 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Conformation-dependent ligand hot spots in the spliceosomal RNA helicase BRR2.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8BCF
 
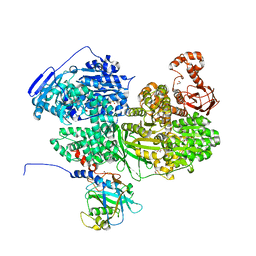 | | Human Brr2 Helicase Region in complex with C-tail deleted Jab1 and compound 78 | | Descriptor: | 1,2-ETHANEDIOL, Pre-mRNA-processing-splicing factor 8, U5 small nuclear ribonucleoprotein 200 kDa helicase, ... | | Authors: | Vester, K, Loll, B, Wahl, M.C. | | Deposit date: | 2022-10-15 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Conformation-dependent ligand hot spots in the spliceosomal RNA helicase BRR2.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8BCH
 
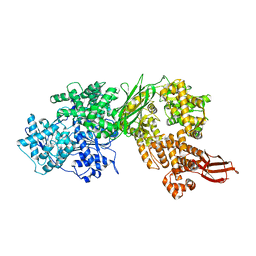 | | Human Brr2 Helicase Region in complex with Sulfaguanidine | | Descriptor: | 1-(4-aminophenyl)sulfonylguanidine, U5 small nuclear ribonucleoprotein 200 kDa helicase | | Authors: | Vester, K, Loll, B, Wahl, M.C. | | Deposit date: | 2022-10-15 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.87 Å) | | Cite: | Conformation-dependent ligand hot spots in the spliceosomal RNA helicase BRR2.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8ALZ
 
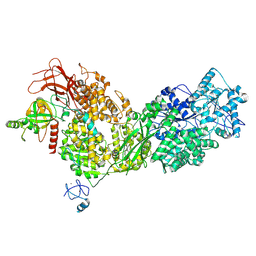 | | Cryo-EM structure of ASCC3 in complex with ASC1 | | Descriptor: | Activating signal cointegrator 1, Activating signal cointegrator 1 complex subunit 3, ZINC ION | | Authors: | Jia, J, Hilal, T, Loll, B, Wahl, M.C. | | Deposit date: | 2022-08-01 | | Release date: | 2023-03-08 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Cryo-EM structure of ASCC3 in complex with ASC1
Nat Commun, 2023
|
|
8BCA
 
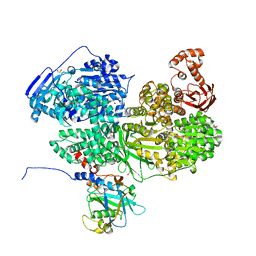 | | Human Brr2 Helicase Region in complex with C-tail deleted Jab1 and compound 26 | | Descriptor: | 1,2-ETHANEDIOL, 3-azanyl-~{N}-methyl-4-(methylamino)benzenesulfonamide, GLYCEROL, ... | | Authors: | Vester, K, Loll, B, Wahl, M.C. | | Deposit date: | 2022-10-15 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Conformation-dependent ligand hot spots in the spliceosomal RNA helicase BRR2.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8BCC
 
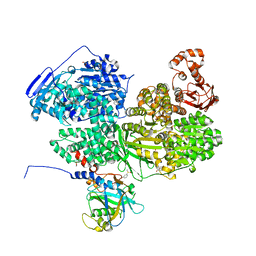 | | Human Brr2 Helicase Region in complex with C-tail deleted Jab1 and compound 39 | | Descriptor: | 1,2-ETHANEDIOL, 3-oxidanylbenzenesulfonamide, Pre-mRNA-processing-splicing factor 8, ... | | Authors: | Vester, K, Loll, B, Wahl, M.C. | | Deposit date: | 2022-10-15 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Conformation-dependent ligand hot spots in the spliceosomal RNA helicase BRR2.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8BCD
 
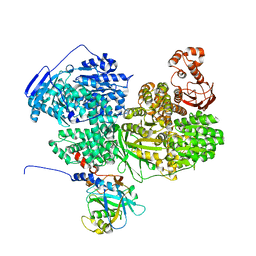 | |
8BC9
 
 | | Human Brr2 Helicase Region in complex with C-tail deleted Jab1 and compound 24 | | Descriptor: | 1,2-ETHANEDIOL, N-hydroxybenzenesulfonamide, Pre-mRNA-processing-splicing factor 8, ... | | Authors: | Vester, K, Loll, B, Wahl, M.C. | | Deposit date: | 2022-10-15 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Conformation-dependent ligand hot spots in the spliceosomal RNA helicase BRR2.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8BC8
 
 | | Human Brr2 Helicase Region in complex with C-tail deleted Jab1 and compound 18 | | Descriptor: | 1,2-ETHANEDIOL, 3-azanyl-4-oxidanyl-benzenesulfonamide, Pre-mRNA-processing-splicing factor 8, ... | | Authors: | Vester, K, Loll, B, Wahl, M.C. | | Deposit date: | 2022-10-15 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Conformation-dependent ligand hot spots in the spliceosomal RNA helicase BRR2.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8BCB
 
 | | Human Brr2 Helicase Region in complex with C-tail deleted Jab1 and compound 34 | | Descriptor: | 1,2-ETHANEDIOL, Pre-mRNA-processing-splicing factor 8, SULFANILAMIDE, ... | | Authors: | Vester, K, Loll, B, Wahl, M.C. | | Deposit date: | 2022-10-15 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Conformation-dependent ligand hot spots in the spliceosomal RNA helicase BRR2.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8BCG
 
 | | Human Brr2 Helicase Region in complex with C-tail deleted Jab1 and compound 86 | | Descriptor: | 1,2-ETHANEDIOL, 4-chloranyl-~{N}-methoxy-~{N}-methyl-benzenesulfonamide, Pre-mRNA-processing-splicing factor 8, ... | | Authors: | Vester, K, Loll, B, Wahl, M.C. | | Deposit date: | 2022-10-15 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Conformation-dependent ligand hot spots in the spliceosomal RNA helicase BRR2.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
7PJH
 
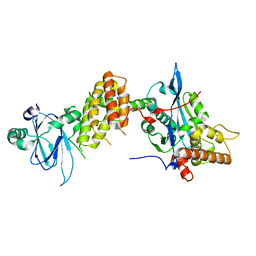 | | Crystal structure of the human spliceosomal maturation factor AAR2 bound to the RNAse H domain of PRPF8 | | Descriptor: | Pre-mRNA-processing-splicing factor 8, Protein AAR2 homolog | | Authors: | Preussner, M, Santos, K, Heroven, A.C, Alles, J, Heyd, F, Wahl, M.C, Weber, G. | | Deposit date: | 2021-08-24 | | Release date: | 2022-11-02 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural and functional investigation of the human snRNP assembly factor AAR2 in complex with the RNase H-like domain of PRPF8.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
1HH2
 
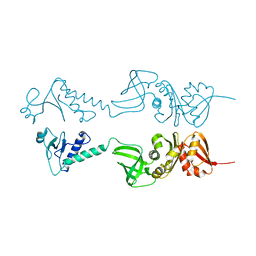 | | Crystal structure of NusA from Thermotoga maritima | | Descriptor: | N UTILIZATION SUBSTANCE PROTEIN A | | Authors: | Worbs, M, Bourenkov, G.P, Bartunik, H.D, Huber, R, Wahl, M.C. | | Deposit date: | 2000-12-18 | | Release date: | 2001-10-19 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | An Extended RNA Binding Surface Through Arrayed S1 and Kh Domains in Transcription Factor Nusa
Mol.Cell, 7, 2001
|
|
1M1G
 
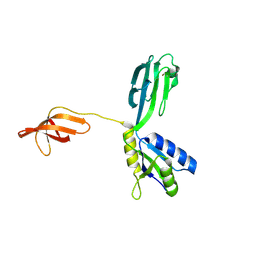 | | Crystal Structure of Aquifex aeolicus N-utilization substance G (NusG), Space Group P2(1) | | Descriptor: | Transcription antitermination protein nusG | | Authors: | Steiner, T, Kaiser, J.T, Marinkovic, S, Huber, R, Wahl, M.C. | | Deposit date: | 2002-06-19 | | Release date: | 2003-02-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of transcription factor NusG in light of its nucleic
acid- and protein-binding activities
Embo J., 21, 2002
|
|
1M1H
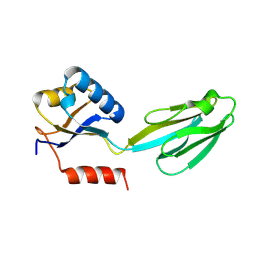 
 | | Crystal structure of Aquifex aeolicus N-utilization substance G (NusG), Space group I222 | | Descriptor: | Transcription antitermination protein nusG | | Authors: | Steiner, T, Kaiser, J.T, Marinkovic, S, Huber, R, Wahl, M.C. | | Deposit date: | 2002-06-19 | | Release date: | 2003-02-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structures of transcription factor NusG in light of its nucleic acid- and protein-binding activities
Embo J., 21, 2002
|
|
2V9V
 
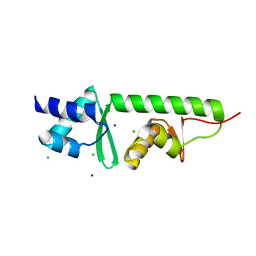 | | Crystal Structure of Moorella thermoacetica SelB(377-511) | | Descriptor: | CHLORIDE ION, SELENOCYSTEINE-SPECIFIC ELONGATION FACTOR, SODIUM ION | | Authors: | Ganichkin, O, Wahl, M.C. | | Deposit date: | 2007-08-27 | | Release date: | 2007-09-11 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Conformational Switches in Winged-Helix Domains 1 and 2 of Bacterial Translation Elongation Factor Selb.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2WFJ
 
 | | Atomic resolution crystal structure of the PPIase domain of human cyclophilin G in complex with cyclosporin A. | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, CYCLOSPORIN A, ... | | Authors: | Stegmann, C.M, Sheldrick, G.M, Wahl, M.C. | | Deposit date: | 2009-04-06 | | Release date: | 2009-06-16 | | Last modified: | 2019-05-22 | | Method: | X-RAY DIFFRACTION (0.75 Å) | | Cite: | The Thermodynamic Influence of Trapped Water Molecules on a Protein-Ligand Interaction.
Angew.Chem.Int.Ed.Engl., 48, 2009
|
|
2WFI
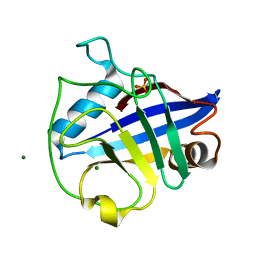 
 | |
2X7K
 
 | | The crystal structure of PPIL1 in complex with cyclosporine A suggests a binding mode for SKIP | | Descriptor: | CADMIUM ION, CYCLOSPORIN A, PEPTIDYL-PROLYL CIS-TRANS ISOMERASE-LIKE 1, ... | | Authors: | Stegmann, C.M, Luehrmann, R, Wahl, M.C. | | Deposit date: | 2010-03-01 | | Release date: | 2010-03-23 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | The Crystal Structure of Ppil1 Bound to Cyclosporine a Suggests a Binding Mode for a Linear Epitope of the Skip Protein.
Plos One, 5, 2010
|
|
2XHA
 
 | |
2XHE
 
 | | Crystal structure of the Unc18-syntaxin 1 complex from Monosiga brevicollis | | Descriptor: | SYNTAXIN1, UNC18 | | Authors: | Burkhardt, P, Stegmann, C.M, Wahl, M.C, Fasshauer, D. | | Deposit date: | 2010-06-14 | | Release date: | 2011-06-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Primordial Neurosecretory Apparatus Identified in the Choanoflagellate Monosiga Brevicollis.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
2XHC
 
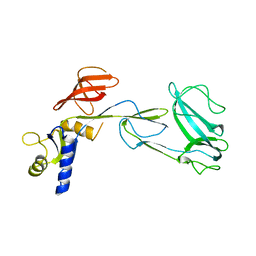 | |
378D
 
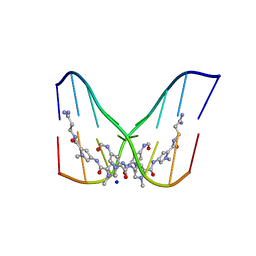 | | STRUCTURE OF THE SIDE-BY-SIDE BINDING OF DISTAMYCIN TO DNA | | Descriptor: | DISTAMYCIN A, DNA (5'-D(*GP*TP*AP*TP*AP*TP*AP*C)-3'), SODIUM ION | | Authors: | Mitra, S.N, Wahl, M.C, Sundaralingam, M. | | Deposit date: | 1998-01-28 | | Release date: | 1999-03-04 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of the side-by-side binding of distamycin to d(GTATATAC)2.
Acta Crystallogr.,Sect.D, 55, 1999
|
|
