3L2D
 
 | |
3L2E
 
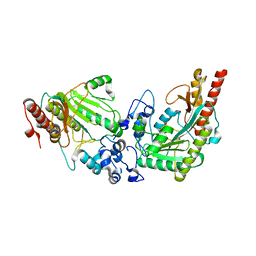
 | | Glycocyamine kinase, alpha-beta heterodimer from marine worm Namalycastis sp. | | Descriptor: | Glycocyamine kinase alpha chain, Glycocyamine kinase beta chain | | Authors: | Lim, K, Pullalarevu, S, Herzberg, O. | | Deposit date: | 2009-12-15 | | Release date: | 2010-03-02 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural basis for the mechanism and substrate specificity of glycocyamine kinase, a phosphagen kinase family member.
Biochemistry, 49, 2010
|
|
2M7T
 
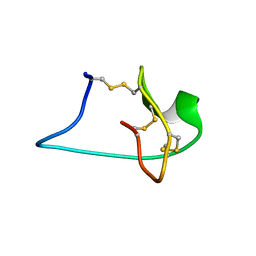 | | Solution NMR Structure of Engineered Cystine Knot Protein 2.5D | | Descriptor: | Cystine Knot Protein 2.5D | | Authors: | Cochran, F.V, Das, R. | | Deposit date: | 2013-04-30 | | Release date: | 2014-05-07 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Challenging the state of the art in protein structure prediction: Highlights of experimental target structures for the 10th Critical Assessment of Techniques for Protein Structure Prediction Experiment CASP10.
Proteins, 82 Suppl 2, 2014
|
|
7M6B
 
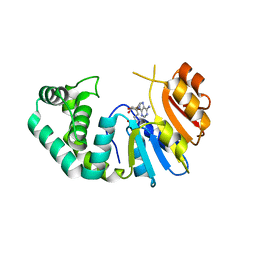 | | The Crystal Structure of Mcbe1 | | Descriptor: | 1,2-ETHANEDIOL, S-ADENOSYL-L-HOMOCYSTEINE, S-ADENOSYLMETHIONINE, ... | | Authors: | Alahuhta, P.M, Lunin, V.V. | | Deposit date: | 2021-03-25 | | Release date: | 2021-04-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Target highlights in CASP14: Analysis of models by structure providers.
Proteins, 89, 2021
|
|
5T87
 
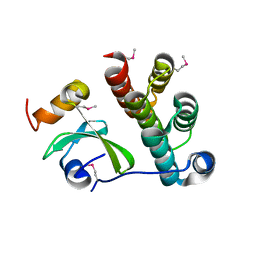 | | Crystal structure of CDI complex from Cupriavidus taiwanensis LMG 19424 | | Descriptor: | CdiA toxin, CdiI immunity protein | | Authors: | Michalska, K, Joachimiak, G, Jedrzejczak, R, Hayes, C.S, Goulding, C.W, Joachimiak, A, Structure-Function Analysis of Polymorphic CDI Toxin-Immunity Protein Complexes (UC4CDI), Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2016-09-06 | | Release date: | 2017-09-13 | | Last modified: | 2019-12-25 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Target highlights from the first post-PSI CASP experiment (CASP12, May-August 2016).
Proteins, 86 Suppl 1, 2018
|
|
1APN
 
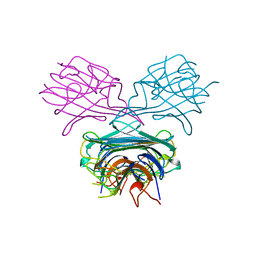 | |
1CLS
 
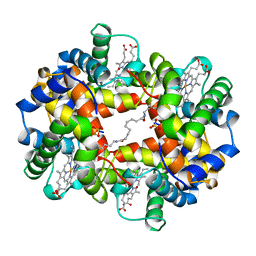 | | CROSS-LINKED HUMAN HEMOGLOBIN DEOXY | | Descriptor: | HEMOGLOBIN, OXYGEN ATOM, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Ji, X, Fronticelli, C, Bucci, E, Gilliland, G.L. | | Deposit date: | 1995-08-29 | | Release date: | 1996-10-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Positive and negative cooperativities at subsequent steps of oxygenation regulate the allosteric behavior of multistate sebacylhemoglobin.
Biochemistry, 35, 1996
|
|
1HDB
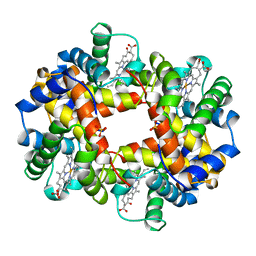 
 | | ANALYSIS OF THE CRYSTAL STRUCTURE, MOLECULAR MODELING AND INFRARED SPECTROSCOPY OF THE DISTAL BETA-HEME POCKET VALINE67(E11)-THREONINE MUTATION OF HEMOGLOBIN | | Descriptor: | HEMOGLOBIN (DEOXY) BETA-V67T, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION | | Authors: | Pechik, I, Ji, X, Fronticelli, C, Gilliland, G.L. | | Deposit date: | 1995-04-14 | | Release date: | 1996-04-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystallographic, molecular modeling, and biophysical characterization of the valine beta 67 (E11)-->threonine variant of hemoglobin.
Biochemistry, 35, 1996
|
|
5SGA
 
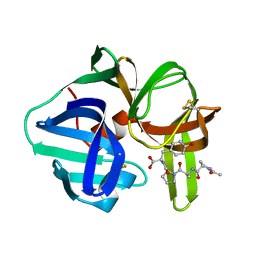 | |
4SGA
 
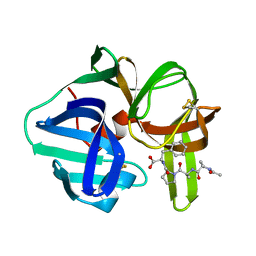 | |
6CP9
 
 | | Contact-dependent growth inhibition toxin - immunity protein complex from Klebsiella pneumoniae 342 | | Descriptor: | CdiA, CdiI | | Authors: | Michalska, K, Stols, L, Eschenfeldt, W, Hayes, C.S, Goulding, C.W, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Structure-Function Analysis of Polymorphic CDI Toxin-Immunity Protein Complexes (UC4CDI) | | Deposit date: | 2018-03-13 | | Release date: | 2019-03-13 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Convergent Evolution of the Barnase/EndoU/Colicin/RelE (BECR) Fold in Antibacterial tRNase Toxins.
Structure, 27, 2019
|
|
1APV
 
 | |
1KGG
 
 | | STRUCTURE OF BETA-LACTAMASE GLU166GLN:ASN170ASP MUTANT | | Descriptor: | PROTEIN (BETA-LACTAMASE), SULFATE ION | | Authors: | Chen, C.C.H, Herzberg, O. | | Deposit date: | 1999-05-20 | | Release date: | 1999-05-28 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Relocation of the catalytic carboxylate group in class A beta-lactamase: the structure and function of the mutant enzyme Glu166-->Gln:Asn170-->Asp.
Protein Eng., 12, 1999
|
|
1ALQ
 
 | |
1APW
 
 | |
6CP8
 
 | | Contact-dependent growth inhibition toxin-immunity protein complex from from E. coli 3006 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CdiA, CdiI, ... | | Authors: | Michalska, K, Stols, L, Eschenfeldt, W, Hayes, C.S, Goulding, C.W, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Structure-Function Analysis of Polymorphic CDI Toxin-Immunity Protein Complexes (UC4CDI) | | Deposit date: | 2018-03-13 | | Release date: | 2019-03-13 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (2.201 Å) | | Cite: | Convergent Evolution of the Barnase/EndoU/Colicin/RelE (BECR) Fold in Antibacterial tRNase Toxins.
Structure, 27, 2019
|
|
1PPK
 
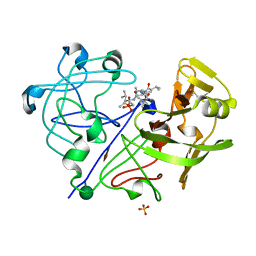 | |
1OME
 
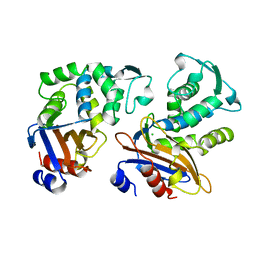 | | CRYSTAL STRUCTURE OF THE OMEGA LOOP DELETION MUTANT (RESIDUES 163-178 DELETED) OF BETA-LACTAMASE FROM STAPHYLOCOCCUS AUREUS PC1 | | Descriptor: | BETA-LACTAMASE, CHLORIDE ION | | Authors: | Banerjee, S, Pieper, U, Herzberg, O. | | Deposit date: | 1998-02-09 | | Release date: | 1998-05-27 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Role of the omega-loop in the activity, substrate specificity, and structure of class A beta-lactamase.
Biochemistry, 37, 1998
|
|
3SGA
 
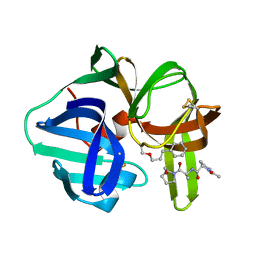 | |
1D18
 
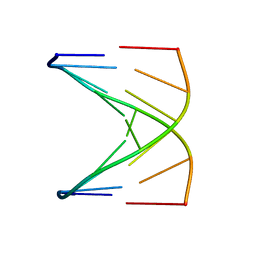 | |
1D19
 
 | |
1PIO
 
 | |
1D20
 
 | | SOLUTION STRUCTURE OF PHAGE LAMBDA HALF-OPERATOR DNA | | Descriptor: | DNA (5'-D(*CP*GP*GP*TP*GP*AP*TP*AP*GP*A)-3'), DNA (5'-D(*TP*CP*TP*AP*TP*CP*AP*CP*CP*G)-3') | | Authors: | Baleja, J.D, Sykes, B.D. | | Deposit date: | 1990-08-01 | | Release date: | 1991-07-15 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of phage lambda half-operator DNA by use of NMR, restrained molecular dynamics, and NOE-based refinement.
Biochemistry, 29, 1990
|
|
1DJC
 
 | |
1DJB
 
 | |
