1AI6
 
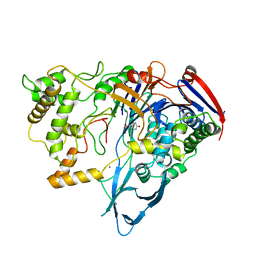 | | PENICILLIN ACYLASE WITH P-HYDROXYPHENYLACETIC ACID | | Descriptor: | 4-HYDROXYPHENYLACETATE, CALCIUM ION, PENICILLIN AMIDOHYDROLASE | | Authors: | Done, S.H. | | Deposit date: | 1997-05-01 | | Release date: | 1997-11-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Ligand-induced conformational change in penicillin acylase.
J.Mol.Biol., 284, 1998
|
|
1BNJ
 
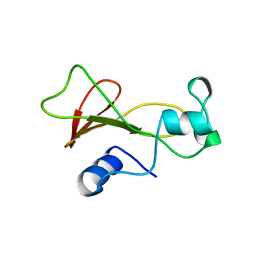 | | BARNASE WILDTYPE STRUCTURE AT PH 9.0 | | Descriptor: | BARNASE | | Authors: | Cameron, A, Henrick, K, Fersht, A.R, Dodson, G, Buckle, A.M. | | Deposit date: | 1995-05-17 | | Release date: | 1995-09-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structural analysis of mutations in the hydrophobic cores of barnase.
J.Mol.Biol., 234, 1993
|
|
1RFE
 
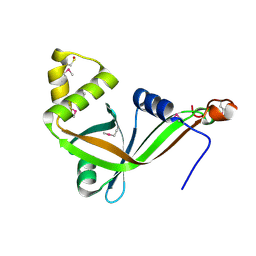 | | Crystal structure of conserved hypothetical protein Rv2991 from Mycobacterium tuberculosis | | Descriptor: | hypothetical protein Rv2991 | | Authors: | Benini, S, Haouz, A, Proux, F, Betton, J.M, Alzari, P, Dodson, G.G, Wilson, K.S, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2003-11-08 | | Release date: | 2004-12-28 | | Last modified: | 2019-04-17 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The crystal structure of Rv2991 from Mycobacterium tuberculosis: An F420binding protein with unknown function.
J. Struct. Biol., 2019
|
|
1GTF
 
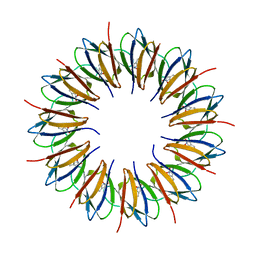 | | The structure of the trp RNA-binding attenuation protein (TRAP) bound to a 53-nucleotide RNA molecule containing GAGUU repeats | | Descriptor: | (GAGUU)10GAG 53-NUCLEOTIDE RNA, TRP RNA-BINDING ATTENUATION PROTEIN (TRAP), TRYPTOPHAN | | Authors: | Hopcroft, N.H, Wendt, A.L, Gollnick, P, Antson, A.A. | | Deposit date: | 2002-01-15 | | Release date: | 2002-04-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Specificity of Trap-RNA Interactions: Crystal Structures of Two Complexes with Different RNA Sequences
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1GDJ
 
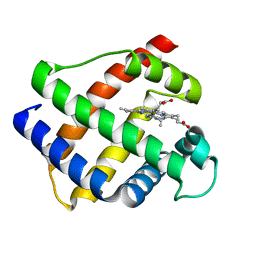 | |
1GTN
 
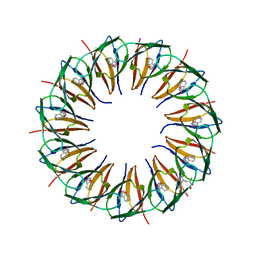 | | Structure of the trp RNA-binding attenuation protein (TRAP) bound to an RNA molecule containing 11 GAGCC repeats | | Descriptor: | (GAGCC)11G 56-NUCLEOTIDE RNA, TRP RNA-BINDING ATTENUATION PROTEIN, TRYPTOPHAN | | Authors: | Hopcroft, N.H, Wendt, A.L, Gollnick, P, Antson, A.A. | | Deposit date: | 2002-01-16 | | Release date: | 2002-04-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Specificity of Trap-RNA Interactions: Crystal Structures of Two Complexes with Different RNA Sequences
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1GDI
 
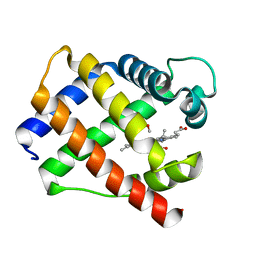 | |
1GDL
 
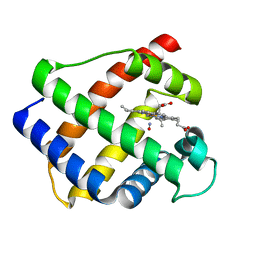 | |
1B21
 
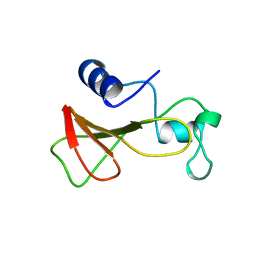 | | DELETION OF A BURIED SALT BRIDGE IN BARNASE | | Descriptor: | PROTEIN (BARNASE), ZINC ION | | Authors: | Vaughan, C.K, Harryson, P, Buckle, A.M, Oliveberg, M, Fersht, A.R. | | Deposit date: | 1998-12-03 | | Release date: | 1998-12-09 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A structural double-mutant cycle: estimating the strength of a buried salt bridge in barnase.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1BAN
 
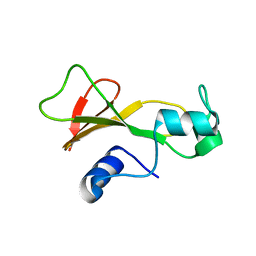 | |
1B27
 | |
1AQZ
 
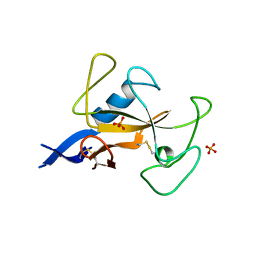 | |
1B2S
 | |
1BGS
 
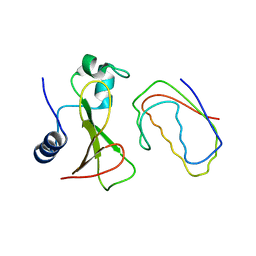 | | RECOGNITION BETWEEN A BACTERIAL RIBONUCLEASE, BARNASE, AND ITS NATURAL INHIBITOR, BARSTAR | | Descriptor: | BARNASE, BARSTAR | | Authors: | Guillet, V, Lapthorn, A, Mauguen, Y. | | Deposit date: | 1993-11-02 | | Release date: | 1994-04-30 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Recognition between a bacterial ribonuclease, barnase, and its natural inhibitor, barstar.
Structure, 1, 1993
|
|
1B20
 
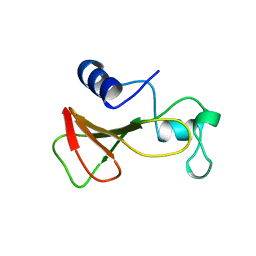 | | DELETION OF A BURIED SALT-BRIDGE IN BARNASE | | Descriptor: | PROTEIN (BARNASE), ZINC ION | | Authors: | Vaughan, C.K, Harryson, P, Buckle, A.M, Oliveberg, M, Fersht, A.R. | | Deposit date: | 1998-12-03 | | Release date: | 1998-12-09 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A structural double-mutant cycle: estimating the strength of a buried salt bridge in barnase.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1B3S
 | |
1BAO
 
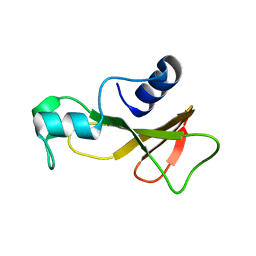 | |
1B2Z
 
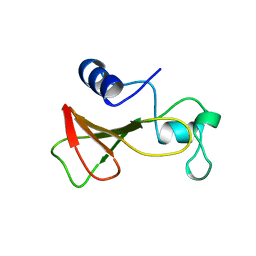 | | DELETION OF A BURIED SALT BRIDGE IN BARNASE | | Descriptor: | PROTEIN (BARNASE), ZINC ION | | Authors: | Vaughan, C.K, Harryson, P, Buckle, A.M, Oliveberg, M, Fersht, A.R. | | Deposit date: | 1998-12-03 | | Release date: | 1998-12-09 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | A structural double-mutant cycle: estimating the strength of a buried salt bridge in barnase.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1B2X
 | | BARNASE WILDTYPE STRUCTURE AT PH 7.5 FROM A CRYO_COOLED CRYSTAL AT 100K | | Descriptor: | PROTEIN (BARNASE), ZINC ION | | Authors: | Harrison, P, Vaughan, C.K, Buckle, A.M, Fersht, A.R. | | Deposit date: | 1998-12-03 | | Release date: | 1998-12-09 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A structural double-mutant cycle: estimating the strength of a buried salt bridge in barnase.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1B2U
 | |
1BNS
 
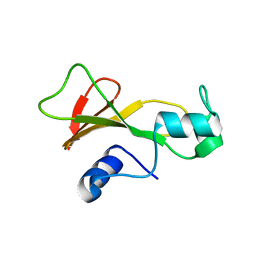 | | STRUCTURAL STUDIES OF BARNASE MUTANTS | | Descriptor: | BARNASE | | Authors: | Chen, Y.W. | | Deposit date: | 1994-04-11 | | Release date: | 1994-06-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Contribution of buried hydrogen bonds to protein stability. The crystal structures of two barnase mutants.
J.Mol.Biol., 234, 1993
|
|
1BRS
 
 | |
1BSE
 
 | |
1BSA
 
 | |
1BSD
 
 | |
