4KMS
 
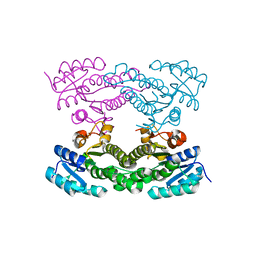 | | Crystal structure of Acetoacetyl-CoA reductase from Rickettsia felis | | Descriptor: | Acetoacetyl-CoA reductase | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID), Abendroth, J, Lukacs, C, Edwards, T.E, Lorimer, D. | | Deposit date: | 2013-05-08 | | Release date: | 2013-05-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of acetoacetyl-CoA reductase from Rickettsia felis.
Acta Crystallogr.,Sect.F, 77, 2021
|
|
3T7C
 
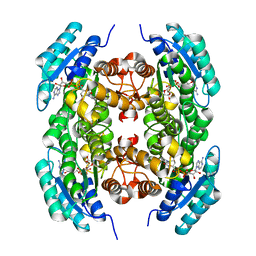 | |
3TK1
 
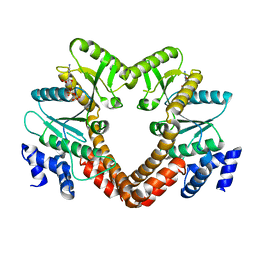 | |
3SX2
 
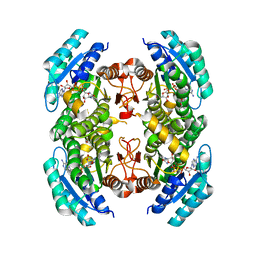 | |
3S55
 
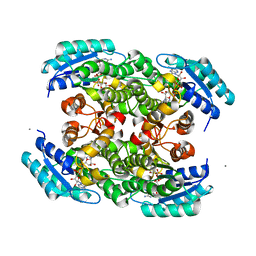 | |
3K2H
 
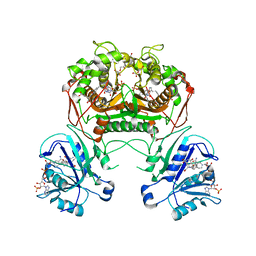 | | Co-crystal structure of dihydrofolate reductase/thymidylate synthase from Babesia bovis with dUMP, Pemetrexed and NADP | | Descriptor: | 1,2-ETHANEDIOL, 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, 2-{4-[2-(2-AMINO-4-OXO-4,7-DIHYDRO-3H-PYRROLO[2,3-D]PYRIMIDIN-5-YL)-ETHYL]-BENZOYLAMINO}-PENTANEDIOIC ACID, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2009-09-30 | | Release date: | 2009-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Inhibitor-bound complexes of dihydrofolate reductase-thymidylate synthase from Babesia bovis.
Acta Crystallogr.,Sect.F, 67, 2011
|
|
3U0B
 
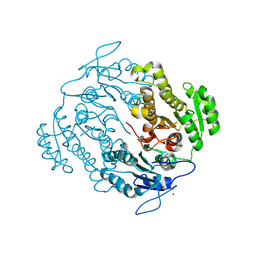 | | Crystal structure of an oxidoreductase from Mycobacterium smegmatis | | Descriptor: | Oxidoreductase, short chain dehydrogenase/reductase family protein, SODIUM ION | | Authors: | Arakaki, T.L, Staker, B.L, Clifton, M.C, Abendroth, J, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2011-09-28 | | Release date: | 2011-10-05 | | Last modified: | 2024-07-17 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural and functional characterization of FabG4 from Mycolicibacterium smegmatis.
Acta Crystallogr.,Sect.F, 80, 2024
|
|
3MEB
 
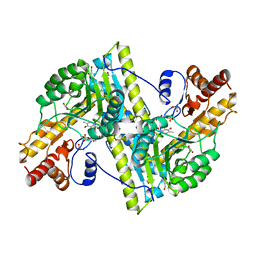 | |
3TSC
 
 | |
6NOT
 
 | |
6NBM
 
 | |
6NHI
 
 | |
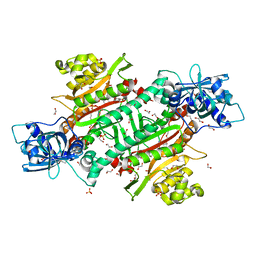
6O3F
 
 | |
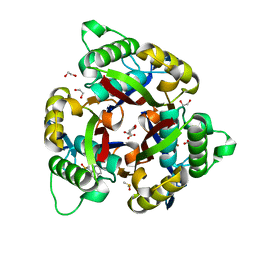
6NMO
 
 | |
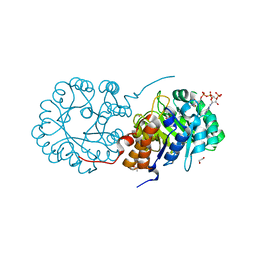
6NDS
 
 | |
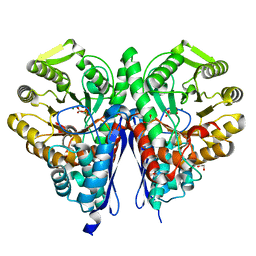
6NB2
 
 | |
6NS0
 
 | |
6NE6
 
 | |
6NQ4
 
 | |
6NUP
 
 | |
6NRZ
 
 | |
6NLX
 
 | |
6D46
 
 | |
6DEG
 
 | |
6D47
 
 | |
