2V66
 | |
4DS1
 
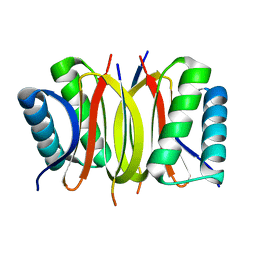 | |
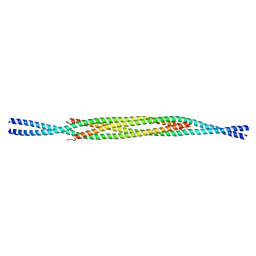
2V71
 
 | | Coiled-coil region of NudEL | | Descriptor: | NUCLEAR DISTRIBUTION PROTEIN NUDE-LIKE 1 | | Authors: | Derewenda, U, Cooper, D.R, Kim, M.H, Derewenda, Z.S. | | Deposit date: | 2007-07-25 | | Release date: | 2007-11-27 | | Last modified: | 2017-06-28 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | The Structure of the Coiled-Coil Domain of Ndel1 and the Basis of its Interaction with Lis1, the Causal Protein of Miller-Dieker Lissencephaly.
Structure, 15, 2007
|
|
6WY1
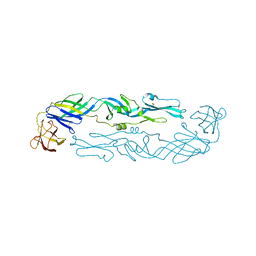 
 | | Crystal structure of an engineered thermostable dengue virus 2 envelope protein dimer | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Dengue 2 soluble recombinant envelope | | Authors: | Kudlacek, S.T, Lakshmanane, P, Kuhlman, B. | | Deposit date: | 2020-05-12 | | Release date: | 2021-11-10 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.42 Å) | | Cite: | Designed, highly expressing, thermostable dengue virus 2 envelope protein dimers elicit quaternary epitope antibodies.
Sci Adv, 7, 2021
|
|
3P8H
 
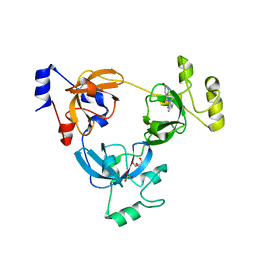 | | Crystal structure of L3MBTL1 (MBT repeat) in complex with a nicotinamide antagonist | | Descriptor: | 3-bromo-5-[(4-pyrrolidin-1-ylpiperidin-1-yl)carbonyl]pyridine, GLYCEROL, Lethal(3)malignant brain tumor-like protein, ... | | Authors: | Lam, R, Herold, J.M, Ouyang, H, Tempel, W, Gao, C, Ravichandran, M, Senisterra, G, Bountra, C, Weigelt, J, Arrowsmith, C.H, Edwards, A.M, Vedadi, M, Kireev, D, Frye, S.V, Brown, P.J, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-10-13 | | Release date: | 2010-11-03 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Small-molecule ligands of methyl-lysine binding proteins.
J.Med.Chem., 54, 2011
|
|
7TJL
 
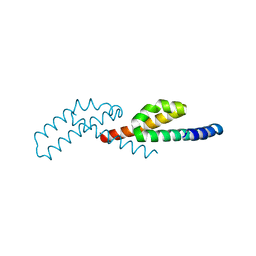 | |
2M0O
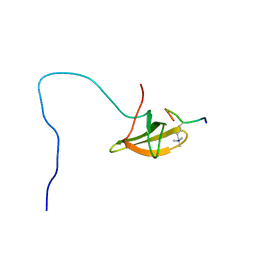 
 | |
6MVH
 
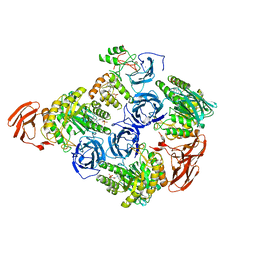 | |
6MVF
 
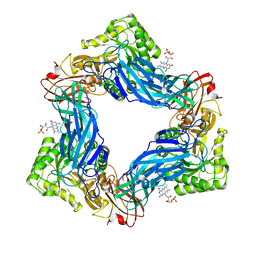 | |
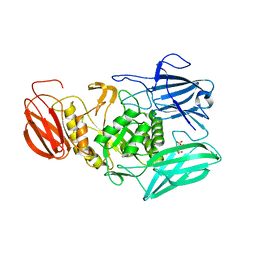
6MVG
 
 | |
4KGH
 
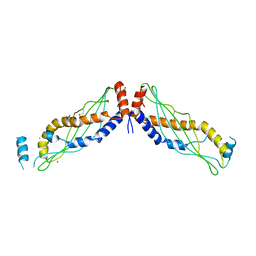 | |
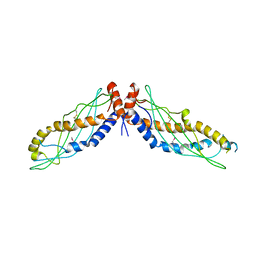
4KGO
 
 | |
2HTF
 
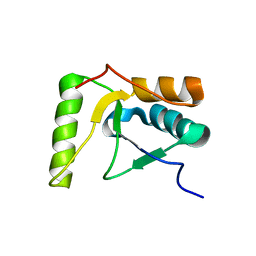 | | The solution structure of the BRCT domain from human polymerase reveals homology with the TdT BRCT domain | | Descriptor: | DNA polymerase mu | | Authors: | DeRose, E.F, Clarkson, M.W, Gilmore, S.A, Ramsden, D.A, Mueller, G.A, London, R.E, Lee, A.L. | | Deposit date: | 2006-07-25 | | Release date: | 2007-02-27 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of polymerase mu's BRCT Domain reveals an element essential for its role in nonhomologous end joining.
Biochemistry, 46, 2007
|
|
6DXU
 
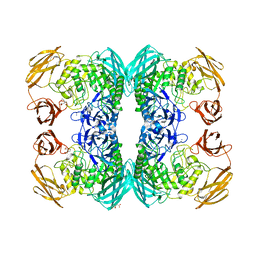 | |
6ED2
 
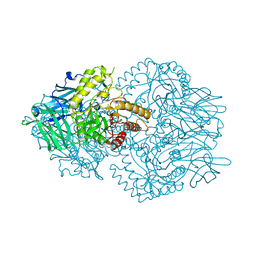 | | Faecalibacterium prausnitzii beta-glucuronidase | | Descriptor: | FORMIC ACID, GLYCEROL, Glycosyl hydrolase family 2, ... | | Authors: | Pellock, S.J, Biernat, K.A, Redinbo, M.R. | | Deposit date: | 2018-08-08 | | Release date: | 2019-02-13 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure, function, and inhibition of drug reactivating human gut microbial beta-glucuronidases.
Sci Rep, 9, 2019
|
|
6ED1
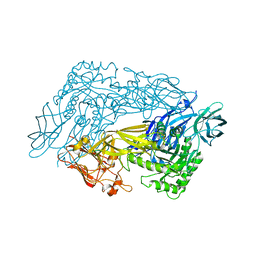 
 | | Bacteroides dorei Beta-glucuronidase | | Descriptor: | Glycosyl hydrolase family 2, sugar binding domain protein, SODIUM ION | | Authors: | Biernat, K.A, Redinbo, M.R. | | Deposit date: | 2018-08-08 | | Release date: | 2019-02-13 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structure, function, and inhibition of drug reactivating human gut microbial beta-glucuronidases.
Sci Rep, 9, 2019
|
|
6EC6
 
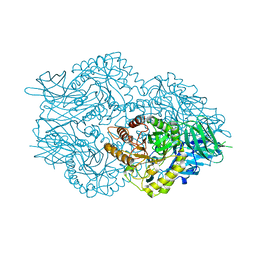 | | Ruminococcus gnavus Beta-glucuronidase | | Descriptor: | Beta-glucuronidase, CHLORIDE ION, GLYCEROL | | Authors: | Biernat, K.A, Redinbo, M.R. | | Deposit date: | 2018-08-07 | | Release date: | 2019-02-13 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structure, function, and inhibition of drug reactivating human gut microbial beta-glucuronidases.
Sci Rep, 9, 2019
|
|
6D6W
 
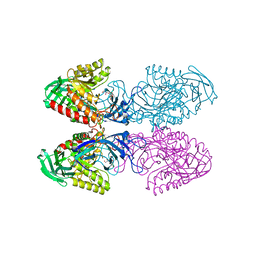 | | Bacteroides uniformis beta-glucuronidase 1 bound to glucuronate | | Descriptor: | Beta-galactosidase/beta-glucuronidase, CHLORIDE ION, GLYCEROL, ... | | Authors: | Walton, W.G, Pellock, S.J, Redinbo, M.R. | | Deposit date: | 2018-04-23 | | Release date: | 2018-10-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Three structurally and functionally distinct beta-glucuronidases from the human gut microbeBacteroides uniformis.
J. Biol. Chem., 293, 2018
|
|
6D8G
 
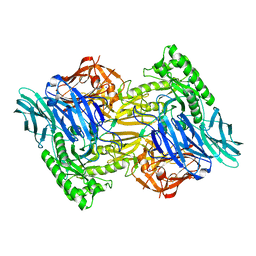 | |
6AP0
 
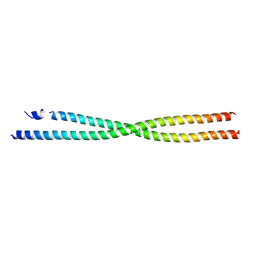 | |
6AZ6
 
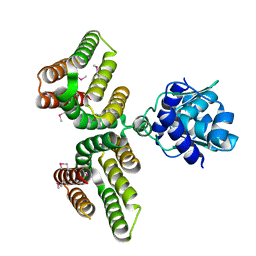 | | Streptococcus agalactiae GntR | | Descriptor: | GntR family transcriptional regulator | | Authors: | Pellock, S.J, Redinbo, M.R. | | Deposit date: | 2017-09-10 | | Release date: | 2017-12-20 | | Last modified: | 2019-12-04 | | Method: | X-RAY DIFFRACTION (1.909 Å) | | Cite: | Structural basis for the regulation of beta-glucuronidase expression by human gut Enterobacteriaceae.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6AZH
 
 | |
6ANO
 
 | |
6AYI
 
 | | Escherichia coli GusR | | Descriptor: | 4-nitrophenyl beta-D-glucopyranosiduronic acid, HTH-type transcriptional regulator UidR | | Authors: | Little, M.S, Pellock, S.J. | | Deposit date: | 2017-09-08 | | Release date: | 2017-12-20 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Structural basis for the regulation of beta-glucuronidase expression by human gut Enterobacteriaceae.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6AOZ
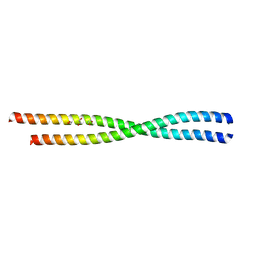 
 | |
