1M93
 
 | |
5YOY
 
 | | Crystal structure of the human tumor necrosis factor in complex with golimumab Fv | | Descriptor: | Golimumab heavy chain variable region, Golimumab light chain variable region, Tumor necrosis factor | | Authors: | Ono, M, Horita, S, Sato, Y, Nomura, Y, Iwata, S, Nomura, N. | | Deposit date: | 2017-10-31 | | Release date: | 2018-05-09 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.727 Å) | | Cite: | Structural basis for tumor necrosis factor blockade with the therapeutic antibody golimumab
Protein Sci., 27, 2018
|
|
1C8O
 
 | |
1JBE
 
 | |
3CME
 
 | |
2FOT
 
 | | Crystal structure of the complex between calmodulin and alphaII-spectrin | | Descriptor: | CALCIUM ION, Calmodulin, alpha-II spectrin Spectrin | | Authors: | Simonovic, M, Zhang, Z, Cianci, C.D, Steitz, T.A, Morrow, J.S. | | Deposit date: | 2006-01-13 | | Release date: | 2006-09-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Structure of the calmodulin alphaII-spectrin complex provides insight into the regulation of cell plasticity.
J.Biol.Chem., 281, 2006
|
|
3CMA
 
 | |
1IMV
 
 | | 2.85 A crystal structure of PEDF | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, PIGMENT EPITHELIUM-DERIVED FACTOR | | Authors: | Simonovic, M, Gettins, P.G.W, Volz, K. | | Deposit date: | 2001-05-11 | | Release date: | 2001-09-26 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Crystal structure of human PEDF, a potent anti-angiogenic and neurite growth-promoting factor.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
1J8E
 
 | | Crystal structure of ligand-binding repeat CR7 from LRP | | Descriptor: | CALCIUM ION, LOW-DENSITY LIPOPROTEIN RECEPTOR-RELATED PROTEIN 1 | | Authors: | Simonovic, M, Dolmer, K, Huang, W, Strickland, D.K, Volz, K, Gettins, P.G.W. | | Deposit date: | 2001-05-21 | | Release date: | 2001-12-19 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Calcium coordination and pH dependence of the calcium affinity of ligand-binding repeat CR7 from the LRP. Comparison with related domains from the LRP and the LDL receptor.
Biochemistry, 40, 2001
|
|
7P55
 
 | |
7P5Q
 
 | |
7P5G
 
 | |
7P5S
 
 | |
3EDU
 | | Crystal structure of the ankyrin-binding domain of human erythroid spectrin | | Descriptor: | Spectrin beta chain, erythrocyte | | Authors: | Simonovic, M, Stabach, P, Simonovic, I, Steitz, T.A, Morrow, J.S. | | Deposit date: | 2008-09-03 | | Release date: | 2009-02-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The structure of the ankyrin-binding site of {beta}-spectrin reveals how tandem spectrin-repeats generate unique ligand-binding properties
Blood, 113, 2009
|
|
3UH0
 
 | | Crystal structure of the yeast mitochondrial threonyl-tRNA synthetase (MST1) in complex with threonyl sulfamoyl adenylate | | Descriptor: | 5'-O-(N-(L-THREONYL)-SULFAMOYL)ADENOSINE, SULFATE ION, Threonyl-tRNA synthetase, ... | | Authors: | Peterson, K.M, Ling, J, Simonovic, I, Cho, C, Soll, D, Simonovic, M. | | Deposit date: | 2011-11-03 | | Release date: | 2012-02-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Yeast mitochondrial threonyl-tRNA synthetase recognizes tRNA isoacceptors by distinct mechanisms and promotes CUN codon reassignment.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
3UGQ
 
 | | Crystal structure of the apo form of the yeast mitochondrial threonyl-tRNA synthetase determined at 2.1 Angstrom resolution | | Descriptor: | POTASSIUM ION, SULFATE ION, Threonyl-tRNA synthetase, ... | | Authors: | Peterson, K.M, Ling, J, Simonovic, I, Cho, C, Soll, D, Simonovic, M. | | Deposit date: | 2011-11-02 | | Release date: | 2012-02-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Yeast mitochondrial threonyl-tRNA synthetase recognizes tRNA isoacceptors by distinct mechanisms and promotes CUN codon reassignment.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
3UGT
 
 | | Crystal structure of the yeast mitochondrial threonyl-tRNA synthetase - orthorhombic crystal form | | Descriptor: | Threonyl-tRNA synthetase, mitochondrial, ZINC ION | | Authors: | Peterson, K.M, Ling, J, Simonovic, I, Cho, C, Soll, D, Simonovic, M. | | Deposit date: | 2011-11-02 | | Release date: | 2012-02-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Yeast mitochondrial threonyl-tRNA synthetase recognizes tRNA isoacceptors by distinct mechanisms and promotes CUN codon reassignment.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
3HJL
 
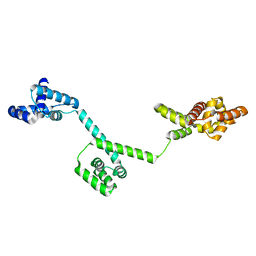 | | The structure of full-length FliG from Aquifex aeolicus | | Descriptor: | Flagellar motor switch protein fliG | | Authors: | Lee, L.K, Ginsburg, M.A, Crovace, C, Donohoe, M, Stock, D. | | Deposit date: | 2009-05-22 | | Release date: | 2010-08-04 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of the torque ring of the flagellar motor and the molecular basis for rotational switching
Nature, 466, 2010
|
|
5FZO
 
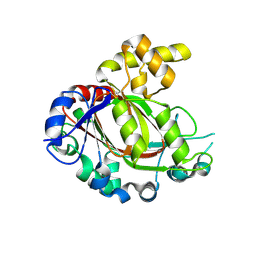 | | Crystal structure of the catalytic domain of human JmjD1C | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, MANGANESE (II) ION, ... | | Authors: | Nowak, R, Talon, R, Krojer, T, Goubin, S, McDonough, M, Fairhead, M, Oppermann, U, Johansson, C. | | Deposit date: | 2016-03-15 | | Release date: | 2016-03-30 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Crystal Structure of the Catalytic Domain of Human Jmjd1C
To be Published
|
|
6O2R
 
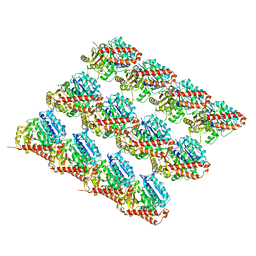 | | Deacetylated Microtubules | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Eshun-Wilson, L, Zhang, R, Portran, D, Nachury, M.V, Toso, D, Lohr, T, Vendruscolo, M, Bonomi, M, Fraser, J.S, Nogales, E. | | Deposit date: | 2019-02-24 | | Release date: | 2019-05-22 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Effects of alpha-tubulin acetylation on microtubule structure and stability.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6O2Q
 
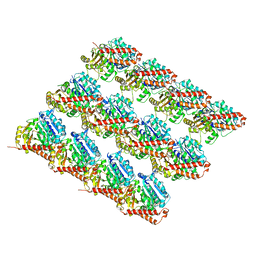 | | Acetylated Microtubules | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Eshun-Wilson, L, Zhang, R, Portran, D, Nachury, M.V, Toso, D, Lohr, T, Vendruscolo, M, Bonomi, M, Fraser, J.S, Nogales, E. | | Deposit date: | 2019-02-24 | | Release date: | 2019-05-22 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Effects of alpha-tubulin acetylation on microtubule structure and stability.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
4WYZ
 | | The crystal structure of the A109G mutant of RNase A in complex with 3'UMP | | Descriptor: | 3'-URIDINEMONOPHOSPHATE, Ribonuclease pancreatic | | Authors: | French, R.L, Gagne, D, Doucet, N, Simonovic, M. | | Deposit date: | 2014-11-18 | | Release date: | 2015-11-18 | | Last modified: | 2019-12-25 | | Method: | X-RAY DIFFRACTION (1.449 Å) | | Cite: | Perturbation of the Conformational Dynamics of an Active-Site Loop Alters Enzyme Activity.
Structure, 23, 2015
|
|
4WYP
 
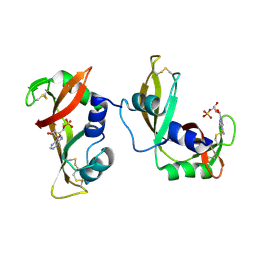 | | The crystal structure of the A109G mutant of RNase A in complex with 5'AMP | | Descriptor: | ADENOSINE MONOPHOSPHATE, Ribonuclease pancreatic | | Authors: | French, R.L, Gagne, D, Doucet, N, Simonovic, M. | | Deposit date: | 2014-11-17 | | Release date: | 2015-11-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.502 Å) | | Cite: | Perturbation of the Conformational Dynamics of an Active-Site Loop Alters Enzyme Activity.
Structure, 23, 2015
|
|
4WYN
 | |
5IZL
 
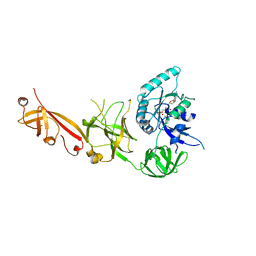 | | The crystal structure of human eEFSec in complex with GDPCP | | Descriptor: | MAGNESIUM ION, PHOSPHOMETHYLPHOSPHONIC ACID GUANYLATE ESTER, Selenocysteine-specific elongation factor | | Authors: | Dobosz-Bartoszek, M, Simonovic, M. | | Deposit date: | 2016-03-25 | | Release date: | 2016-10-12 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.72 Å) | | Cite: | Crystal structures of the human elongation factor eEFSec suggest a non-canonical mechanism for selenocysteine incorporation.
Nat Commun, 7, 2016
|
|
