2NMR
 
 | |
2NPJ
 
 | |
2NPC
 
 | |
2NPK
 
 | |
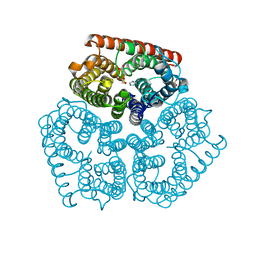
2NOP
 
 | |
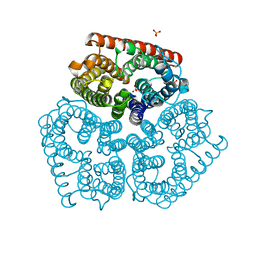
2NPG
 
 | |
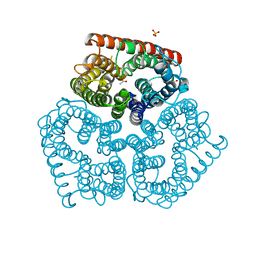
2NPE
 
 | |
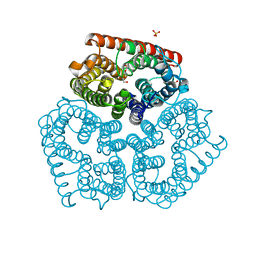
2NOW
 
 | |
2NPD
 
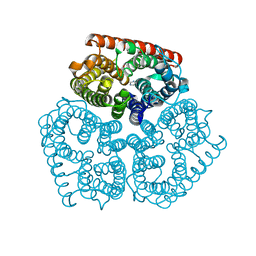 | |
1F97
 
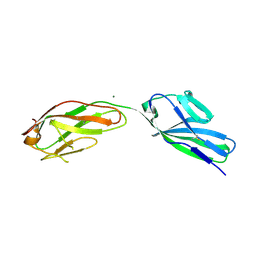 | | SOLUBLE PART OF THE JUNCTION ADHESION MOLECULE FROM MOUSE | | Descriptor: | JUNCTION ADHESION MOLECULE, MAGNESIUM ION | | Authors: | Kostrewa, D, Brockhaus, M, D'Arcy, A, Dale, G, Bazzoni, G, Dejana, E, Winkler, F, Hennig, M. | | Deposit date: | 2000-07-07 | | Release date: | 2001-01-10 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | X-ray structure of junctional adhesion molecule: structural basis for homophilic adhesion via a novel dimerization motif.
EMBO J., 20, 2001
|
|
8PNV
 
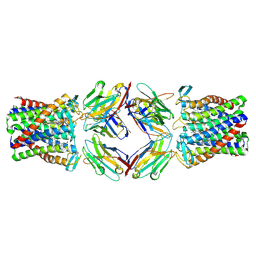 | | Cryo-EM structure of styrene oxide isomerase | | Descriptor: | Nanobody, PROTOPORPHYRIN IX CONTAINING FE, Styrene oxide isomerase | | Authors: | Khanppnavar, B, Korkhov, B, Li, X. | | Deposit date: | 2023-07-02 | | Release date: | 2024-04-03 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (2.048 Å) | | Cite: | Structural basis of the Meinwald rearrangement catalysed by styrene oxide isomerase.
Nat.Chem., 2024
|
|
8PNU
 
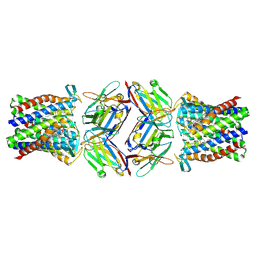 | |
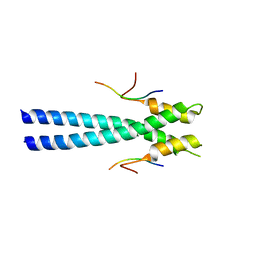
3GJO
 
 | |
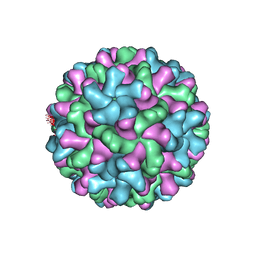
2TBV
 
 | |
1RV5
 
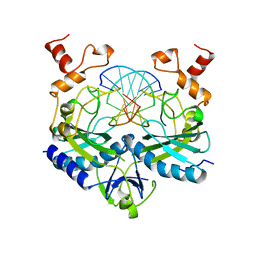 | | COMPLEX OF ECORV ENDONUCLEASE WITH D(AAAGAT)/D(ATCTT) | | Descriptor: | 5'-D(*AP*AP*AP*GP*AP*T*AP*TP*CP*TP*T)-3', ECORV ENDONUCLEASE | | Authors: | Horton, N.C, Perona, J.J. | | Deposit date: | 1998-06-01 | | Release date: | 1998-11-11 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Role of protein-induced bending in the specificity of DNA recognition: crystal structure of EcoRV endonuclease complexed with d(AAAGAT) + d(ATCTT).
J.Mol.Biol., 277, 1998
|
|
1THG
 
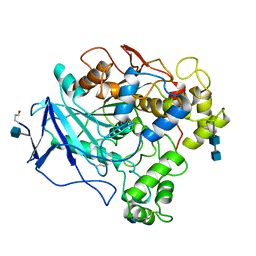 | | 1.8 ANGSTROMS REFINED STRUCTURE OF THE LIPASE FROM GEOTRICHUM CANDIDUM | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Schrag, J.D, Cygler, M. | | Deposit date: | 1992-07-28 | | Release date: | 1993-10-31 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | 1.8 A refined structure of the lipase from Geotrichum candidum.
J.Mol.Biol., 230, 1993
|
|
1AZ0
 
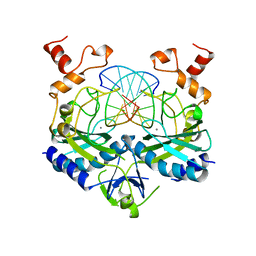 | | ECORV ENDONUCLEASE/DNA COMPLEX | | Descriptor: | CALCIUM ION, DNA (5'-D(*AP*AP*AP*GP*AP*TP*AP*TP*CP*TP*T)-3'), PROTEIN (TYPE II RESTRICTION ENZYME ECORV) | | Authors: | Perona, J.J, Martin, A.M. | | Deposit date: | 1997-11-24 | | Release date: | 1998-06-19 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Conformational transitions and structural deformability of EcoRV endonuclease revealed by crystallographic analysis.
J.Mol.Biol., 273, 1997
|
|
1AZ4
 
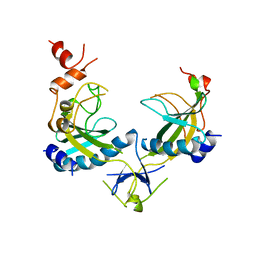 | | ECORV ENDONUCLEASE, UNLIGANDED, FORM B, T93A MUTANT | | Descriptor: | ECORV ENDONUCLEASE | | Authors: | Perona, J, Martin, A. | | Deposit date: | 1997-11-24 | | Release date: | 1998-05-27 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Conformational transitions and structural deformability of EcoRV endonuclease revealed by crystallographic analysis.
J.Mol.Biol., 273, 1997
|
|
1AZ3
 
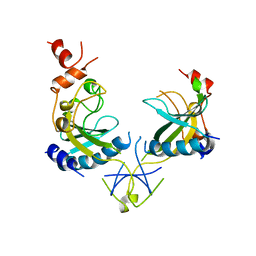 | | ECORV ENDONUCLEASE, UNLIGANDED, FORM B | | Descriptor: | ECORV ENDONUCLEASE | | Authors: | Perona, J, Martin, A. | | Deposit date: | 1997-11-24 | | Release date: | 1998-05-27 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Conformational transitions and structural deformability of EcoRV endonuclease revealed by crystallographic analysis.
J.Mol.Biol., 273, 1997
|
|
1DAN
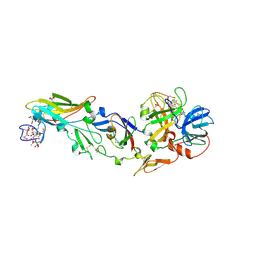 
 | |
1JR4
 
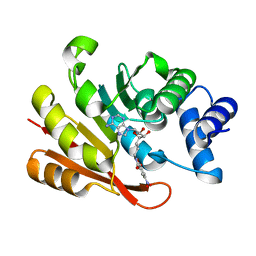 | | CATECHOL O-METHYLTRANSFERASE BISUBSTRATE-INHIBITOR COMPLEX | | Descriptor: | CATECHOL O-METHYLTRANSFERASE, SOLUBLE FORM, MAGNESIUM ION, ... | | Authors: | Ruf, A, Stihle, M. | | Deposit date: | 2001-08-10 | | Release date: | 2002-08-10 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | X-ray Crystal Structure of a Bisubstrate Inhibitor Bound to the Enzyme Catechol-O-methyltransferase: A Dramatic Effect of Inhibitor Preorganization on Binding Affinity.
Angew.Chem.Int.Ed.Engl., 40, 2001
|
|
4AYV
 
 | | Human thrombin - inhibitor complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, HIRUDIN-3A', SODIUM ION, ... | | Authors: | Banner, D.W, D'Arcy, A, Winkler, F.K, Hilpert, K. | | Deposit date: | 2012-06-22 | | Release date: | 2012-08-15 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Design and Synthesis of Potent and Highly Selective Thrombin Inhibitors.
J.Med.Chem., 37, 1994
|
|
4AX9
 
 | | Human thrombin complexed with Napsagatran, RO0466240 | | Descriptor: | 2-[[(2S)-4-[[(3S)-1-carbamimidoylpiperidin-3-yl]methylamino]-2-(naphthalen-2-ylsulfonylamino)-4-oxidanylidene-butanoyl] -cyclopropyl-amino]ethanoic acid, 2-acetamido-2-deoxy-beta-D-glucopyranose, HIRUDIN VARIANT-1, ... | | Authors: | Banner, D.W, D'Arcy, A, Winkler, F.K, Hilpert, K, Spinelli, S, Cambillau, C. | | Deposit date: | 2012-06-11 | | Release date: | 2012-06-20 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Design and Synthesis of Potent and Highly Selective Thrombin Inhibitors.
J.Med.Chem., 37, 1994
|
|
4AZ2
 
 | | Human thrombin - inhibitor complex | | Descriptor: | (R)-N-((S)-1-CARBAMIMIDOYL-PIPERIDIN-3-YLMETHYL)-2-(NAPHTHALENE-2-SULFONYLAMINO)-3-PHENYL-PROPIONAMIDE, 2-acetamido-2-deoxy-beta-D-glucopyranose, HIRUDIN-3A', ... | | Authors: | Banner, D.W, D'Arcy, A, Winkler, F.K, Hilpert, K. | | Deposit date: | 2012-06-22 | | Release date: | 2012-08-15 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Design and Synthesis of Potent and Highly Selective Thrombin Inhibitors.
J.Med.Chem., 37, 1994
|
|
1TNR
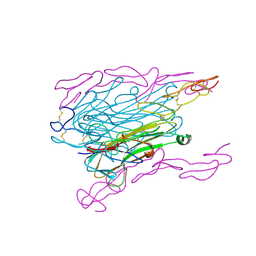 
 | |
