8RLK
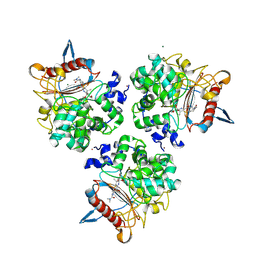 
 | | Structure of the apo form of PIB-1 in an Orthorombic space group | | Descriptor: | (4R,5S)-3-{[(3S,5S)-5-(dimethylcarbamoyl)pyrrolidin-3-yl]sulfanyl}-5-[(2S,3R)-3-hydroxy-1-oxobutan-2-yl]-4-methyl-4,5-d ihydro-1H-pyrrole-2-carboxylic acid, Class C beta-lactamase-related serine hydrolase, MAGNESIUM ION, ... | | Authors: | Medrano, F.J, Romero, A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-08-14 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A new type of Class C beta-lactamases defined by PIB-1. A metal-dependent carbapenem-hydrolyzing beta-lactamase, from Pseudomonas aeruginosa: Structural and functional analysis.
Int.J.Biol.Macromol., 277, 2024
|
|
8RLL
 
 | | Structure of the apo form of PIB-1 in an Orthorombic space group | | Descriptor: | Class C beta-lactamase-related serine hydrolase | | Authors: | Medrano, F.J, Romero, A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-08-14 | | Last modified: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (2.72 Å) | | Cite: | A new type of Class C beta-lactamases defined by PIB-1. A metal-dependent carbapenem-hydrolyzing beta-lactamase, from Pseudomonas aeruginosa: Structural and functional analysis.
Int.J.Biol.Macromol., 277, 2024
|
|
4W7O
 
 | |
4W7L
 
 | |
4Y8F
 
 | |
4Y96
 
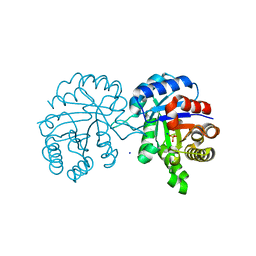 | | Crystal structure of Triosephosphate Isomerase from Gemmata obscuriglobus | | Descriptor: | CALCIUM ION, PHOSPHATE ION, SODIUM ION, ... | | Authors: | Romero-Romero, S, Rodriguez-Romero, A, Fernandez-Velasco, D.A. | | Deposit date: | 2015-02-17 | | Release date: | 2015-07-15 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.581 Å) | | Cite: | Reversibility and two state behaviour in the thermal unfolding of oligomeric TIM barrel proteins.
Phys Chem Chem Phys, 17, 2015
|
|
5C9K
 
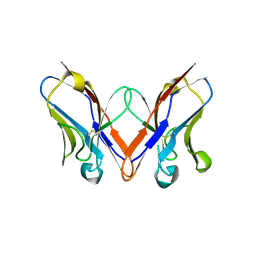 | |
4Y90
 
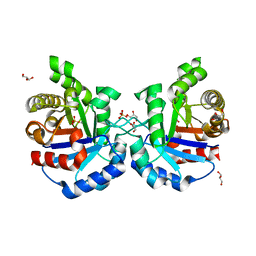 | | Crystal structure of Triosephosphate Isomerase from Deinococcus radiodurans | | Descriptor: | CALCIUM ION, GLYCEROL, SODIUM ION, ... | | Authors: | Romero-Romero, S, Rodriguez-Romero, A, Fernadez-Velasco, D.A. | | Deposit date: | 2015-02-16 | | Release date: | 2015-07-15 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Reversibility and two state behaviour in the thermal unfolding of oligomeric TIM barrel proteins.
Phys Chem Chem Phys, 17, 2015
|
|
4Y9A
 
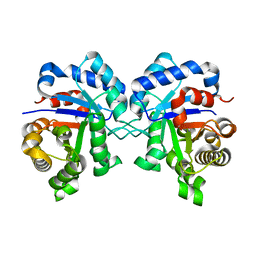 | |
2BML
 
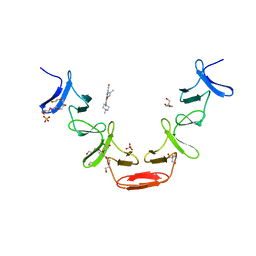 | | Ofloxacin-like antibiotics inhibit pneumococcal cell wall degrading virulence factors | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, AUTOLYSIN, DEXTROFLOXACINE, ... | | Authors: | Fernandez-Tornero, C, Gimenez-Gallego, G, Romero, A, Garcia, E, Pascual-Teresa, B.D, Lopez, R. | | Deposit date: | 2005-03-15 | | Release date: | 2005-03-30 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Ofloxacin-Like Antibiotics Inhibit Pneumococcal Cell Wall-Degrading Virulence Factors
J.Biol.Chem., 280, 2005
|
|
4HPG
 
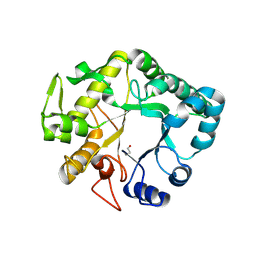 | | Crystal structure of a glycosylated beta-1,3-glucanase (HEV B 2), an allergen from Hevea brasiliensis | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Beta-1,3-glucanase, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Rodriguez-Romero, A, Hernandez-Santoyo, A. | | Deposit date: | 2012-10-23 | | Release date: | 2013-11-27 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.5364 Å) | | Cite: | Structural analysis of the endogenous glycoallergen Hev b 2 (endo-beta-1,3-glucanase) from Hevea brasiliensis and its recognition by human basophils.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4IIS
 
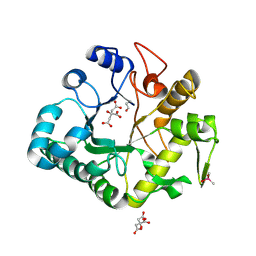 | | Crystal structure of a glycosylated beta-1,3-glucanase (HEV B 2), An allergen from Hevea Brasiliensis (Space group P41) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Beta-1,3-glucanase form 'RRII Gln 2', CACODYLATE ION, ... | | Authors: | Rodriguez-Romero, A, Hernandez-Santoyo, A. | | Deposit date: | 2012-12-20 | | Release date: | 2013-11-27 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.6676 Å) | | Cite: | Structural analysis of the endogenous glycoallergen Hev b 2 (endo-beta-1,3-glucanase) from Hevea brasiliensis and its recognition by human basophils.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
7KOT
 
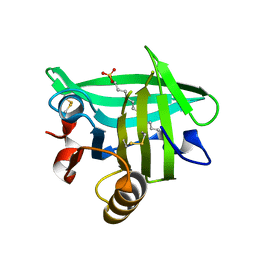 | |
4V3F
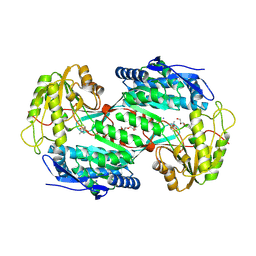 
 | |
4V37
 
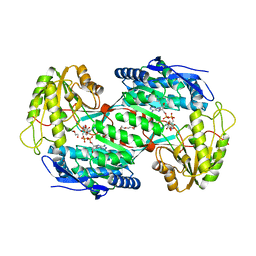 | |
7ZBP
 
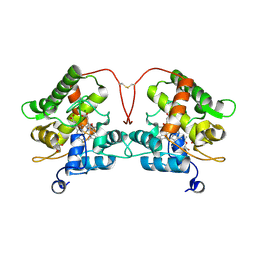 | | Unspecific peroxygenase from Marasmius rotula | | Descriptor: | ACETATE ION, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Santillana, E, Romero, A. | | Deposit date: | 2022-03-24 | | Release date: | 2022-05-18 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structural Characterization of Two Short Unspecific Peroxygenases: Two Different Dimeric Arrangements.
Antioxidants, 11, 2022
|
|
7ZCL
 
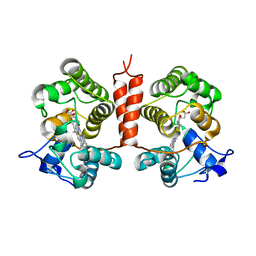 | | Unspecific peroxygenase from Collariella virescens | | Descriptor: | Collariella virescens UPO, HEME C, MAGNESIUM ION | | Authors: | Santillana, E, Romero, A. | | Deposit date: | 2022-03-28 | | Release date: | 2022-05-18 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural Characterization of Two Short Unspecific Peroxygenases: Two Different Dimeric Arrangements.
Antioxidants, 11, 2022
|
|
4W7N
 
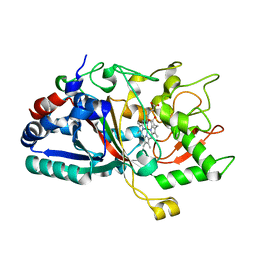 | |
4W7J
 
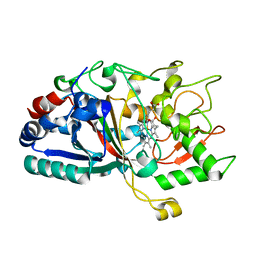 | |
4W7K
 
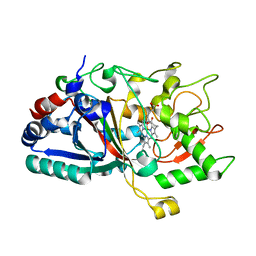 | |
4MST
 
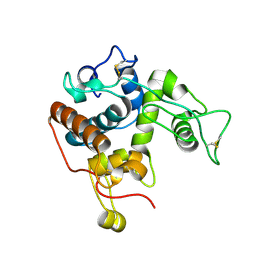 | |
4MPI
 
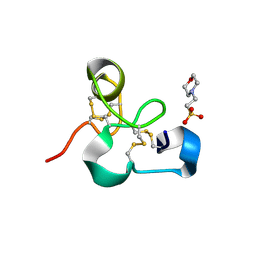 | | Crystal structure of the chitin-binding module (CBM18) of a chitinase-like protein from Hevea brasiliensis | | Descriptor: | 1,4-DIETHYLENE DIOXIDE, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Class I chitinase | | Authors: | Martinez-Caballero, C.S, Hermoso, J.A, Rodriguez-Romero, A. | | Deposit date: | 2013-09-12 | | Release date: | 2014-08-27 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.602 Å) | | Cite: | Comparative study of two GH19 chitinase-like proteins from Hevea brasiliensis, one exhibiting a novel carbohydrate-binding domain.
Febs J., 281, 2014
|
|
8EYM
 
 | | CRYSTAL STRUCTURE OF NAGB-II PHOSPHOSUGAR ISOMERASE FROM SHEWANELLA DENITRIFICANS OS217 IN COMPLEX WITH GLUCITOLAMINE-6-PHOSPHATE AND N-ACETYLGLUCOSAMINE-6-PHOSPHATE AT 2.31 A RESOLUTION | | Descriptor: | 2-DEOXY-2-AMINO GLUCITOL-6-PHOSPHATE, 2-acetamido-2-deoxy-6-O-phosphono-alpha-D-glucopyranose, GLUCOSAMINE-6-PHOSPHATE DEAMINASE | | Authors: | Rodriguez-Hernandez, A, Marcos-Viquez, J, Rodriguez-Romero, A, Bustos-Jaimes, I. | | Deposit date: | 2022-10-27 | | Release date: | 2023-05-17 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.311 Å) | | Cite: | Substrate binding in the allosteric site mimics homotropic cooperativity in the SIS-fold glucosamine-6-phosphate deaminases.
Protein Sci., 32, 2023
|
|
8FDB
 
 | | CRYSTAL STRUCTURE OF NAGB-II PHOSPHOSUGAR ISOMERASE FROM Shewanella denitrificans OS217 IN COMPLEX WITH GLUCITOLAMINE-6-PHOSPHATE AT 3.06 A RESOLUTION. | | Descriptor: | 2-DEOXY-2-AMINO GLUCITOL-6-PHOSPHATE, GLYCEROL, Glutamine-fructose-6-phosphate transaminase (Isomerizing), ... | | Authors: | Rodriguez-Romero, A, Rodriguez-Hernandez, A, Marcos-Viquez, J, Bustos-Jaimes, I. | | Deposit date: | 2022-12-02 | | Release date: | 2023-05-17 | | Last modified: | 2023-06-07 | | Method: | X-RAY DIFFRACTION (3.06 Å) | | Cite: | Substrate binding in the allosteric site mimics homotropic cooperativity in the SIS-fold glucosamine-6-phosphate deaminases.
Protein Sci., 32, 2023
|
|
8EOL
 
 | |
