1XTT
 
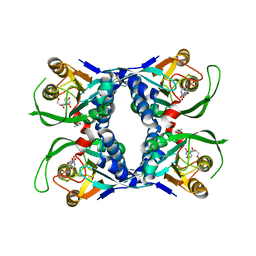 | | Sulfolobus solfataricus uracil phosphoribosyltransferase in complex with uridine 5'-monophosphate (UMP) | | Descriptor: | ACETIC ACID, Probable uracil phosphoribosyltransferase, URIDINE-5'-MONOPHOSPHATE | | Authors: | Arent, S, Harris, P, Jensen, K.F, Larsen, S. | | Deposit date: | 2004-10-24 | | Release date: | 2005-02-08 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Allosteric Regulation and Communication between Subunits in Uracil Phosphoribosyltransferase from Sulfolobus solfataricus(,)
Biochemistry, 44, 2005
|
|
1XTV
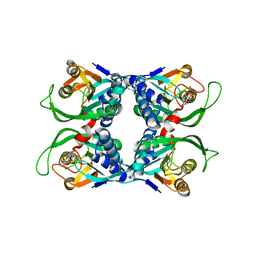 
 | | Sulfolobus solfataricus uracil phosphoribosyltransferase with uridine 5'-monophosphate (UMP) bound to half of the subunits | | Descriptor: | Probable uracil phosphoribosyltransferase, URIDINE-5'-MONOPHOSPHATE | | Authors: | Arent, S, Harris, P, Jensen, K.F, Larsen, S. | | Deposit date: | 2004-10-24 | | Release date: | 2005-02-08 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Allosteric Regulation and Communication between Subunits in Uracil Phosphoribosyltransferase from Sulfolobus solfataricus(,)
Biochemistry, 44, 2005
|
|
1XTU
 
 | | Sulfolobus solfataricus uracil phosphoribosyltransferase in complex with uridine 5'-monophosphate (UMP) and cytidine 5'-triphosphate (CTP) | | Descriptor: | CYTIDINE-5'-TRIPHOSPHATE, Probable uracil phosphoribosyltransferase, URIDINE-5'-MONOPHOSPHATE | | Authors: | Arent, S, Harris, P, Jensen, K.F, Larsen, S. | | Deposit date: | 2004-10-24 | | Release date: | 2005-02-08 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Allosteric Regulation and Communication between Subunits in Uracil Phosphoribosyltransferase from Sulfolobus solfataricus(,)
Biochemistry, 44, 2005
|
|
1LY8
 
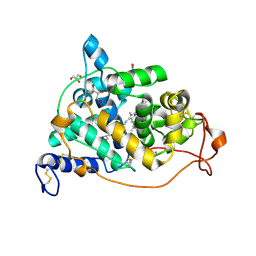 | | The crystal structure of a mutant enzyme of Coprinus cinereus peroxidase provides an understanding of its increased thermostability and insight into modelling of protein structures | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, GLYCEROL, ... | | Authors: | Houborg, K, Harris, P, Poulsen, J.-C.N, Svendsen, A, Schneider, P, Larsen, S. | | Deposit date: | 2002-06-07 | | Release date: | 2002-06-14 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | The structure of a mutant enzyme of Coprinus cinereus peroxidase provides an understanding of its increased thermostability.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1JJK
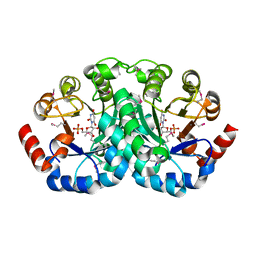 
 | | Selenomethionine Substitution of Orotidine-5'-monophosphate Decarboxylase from E. coli Causes a Change in Crystal Contacts and Space Group | | Descriptor: | 6-HYDROXYURIDINE-5'-PHOSPHATE, OROTIDINE 5'-PHOSPHATE DECARBOXYLASE | | Authors: | Poulsen, J.-C.N, Harris, P, Jensen, K.F, Larsen, S. | | Deposit date: | 2001-07-06 | | Release date: | 2001-08-01 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Selenomethionine substitution of orotidine-5'-monophosphate decarboxylase causes a change in crystal contacts and space group.
Acta Crystallogr.,Sect.D, 57, 2001
|
|
2QLP
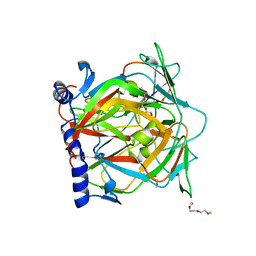 
 | |
7QRI
 
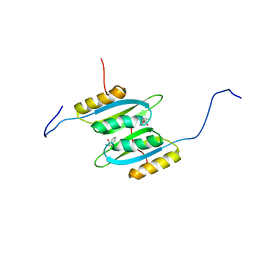 | | Regulatory domain dimer of tryptophan hydroxylase 2 in complex with L-Phe | | Descriptor: | PHENYLALANINE, Tryptophan 5-hydroxylase 2 | | Authors: | Vedel, I.M, Prestel, A, Harris, P, Peters, G.H.J, Kragelund, B.B. | | Deposit date: | 2022-01-11 | | Release date: | 2023-05-03 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of human tryptophan hydroxylase 2 reveals that L-Phe is superior to L-Trp as the regulatory domain ligand.
Structure, 31, 2023
|
|
4A6A
 
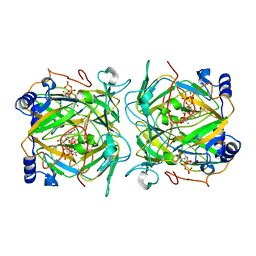 | |
1SIZ
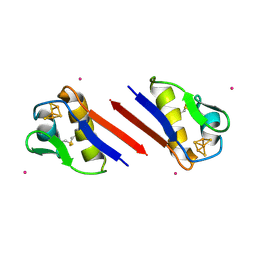 
 | |
1SJ1
 
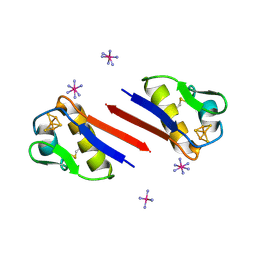 | | The 1.5 A Resolution Crystal Structure of [Fe3S4]-Ferredoxin from the hyperthermophilic Archaeon Pyrococcus furiosus | | Descriptor: | COBALT HEXAMMINE(III), FE3-S4 CLUSTER, Ferredoxin | | Authors: | Nielsen, M.S, Harris, P, Ooi, B.L, Christensen, H.E. | | Deposit date: | 2004-03-02 | | Release date: | 2004-05-25 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The 1.5 A Resolution Crystal Structure of [Fe3S4]-Ferredoxin from the Hyperthermophilic Archaeon Pyrococcus furiosus
Biochemistry, 43, 2004
|
|
3PNI
 | |
1M6Z
 
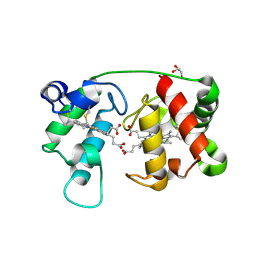 | | Crystal structure of reduced recombinant cytochrome c4 from Pseudomonas stutzeri | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Cytochrome c4, GLYCEROL, ... | | Authors: | Noergaard, A, Harris, P, Larsen, S, Christensen, H.E.M. | | Deposit date: | 2002-07-18 | | Release date: | 2003-09-16 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Structural comparison of recombinant Pseudomonas stutzeri cytochrome c4 in two oxidation states
To be Published
|
|
1M70
 
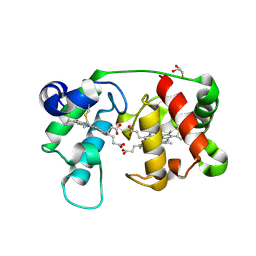 | |
2GIM
 
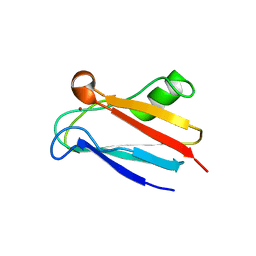 | |
2GJP
 
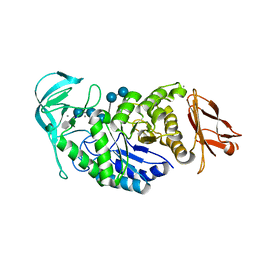 | | Structure of Bacillus halmapalus alpha-amylase, crystallized with the substrate analogue acarbose and maltose | | Descriptor: | 4,6-dideoxy-4-{[(1S,5R,6S)-3-formyl-5,6-dihydroxy-4-oxocyclohex-2-en-1-yl]amino}-alpha-D-xylo-hex-5-enopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 4,6-dideoxy-4-{[(1S,5R,6S)-3-formyl-5,6-dihydroxy-4-oxocyclohex-2-en-1-yl]amino}-alpha-D-xylo-hex-5-enopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Lyhne-Iversen, L, Hobley, T.J, Kaasgaard, S.G, Harris, P. | | Deposit date: | 2006-03-31 | | Release date: | 2006-09-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of Bacillus halmapalus alpha-amylase crystallized with and without the substrate analogue acarbose and maltose.
Acta Crystallogr.,Sect.F, 62, 2006
|
|
2GJR
 
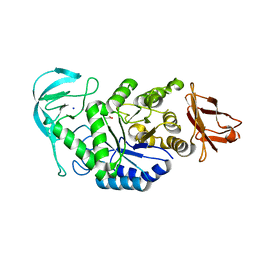 | | Structure of bacillus halmapalus alpha-amylase without any substrate analogues | | Descriptor: | ACETATE ION, CALCIUM ION, SODIUM ION, ... | | Authors: | Lyhne-Iversen, L, Hobley, T.J, Kaasgaard, S.G, Harris, P. | | Deposit date: | 2006-03-31 | | Release date: | 2006-09-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of Bacillus halmapalus alpha-amylase crystallized with and without the substrate analogue acarbose and maltose.
Acta Crystallogr.,Sect.F, 62, 2006
|
|
2Z8Q
 
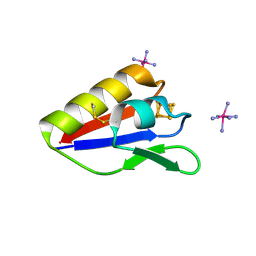 | |
4CG0
 
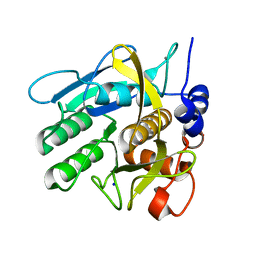 | | Savinase crystal structures for combined single crystal diffraction and powder diffraction analysis | | Descriptor: | CALCIUM ION, SODIUM ION, SUBTILISIN SAVINASE | | Authors: | Frankaer, C.G, Moroz, O.V, Turkenburg, J.P, Aspmo, S.I, Thymark, M, Friis, E.P, Stahla, K, Nielsen, J.E, Wilson, K.S, Harris, P. | | Deposit date: | 2013-11-19 | | Release date: | 2014-04-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.36 Å) | | Cite: | Analysis of an Industrial Production Suspension of Bacillus Lentus Subtilisin Crystals by Powder Diffraction: A Powerful Quality-Control Tool.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4CFY
 
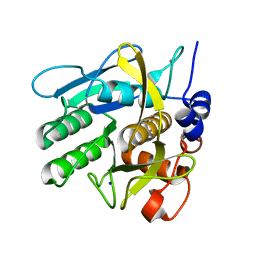 | | SAVINASE CRYSTAL STRUCTURES FOR COMBINED SINGLE CRYSTAL DIFFRACTION AND POWDER DIFFRACTION ANALYSIS | | Descriptor: | CALCIUM ION, SODIUM ION, SUBTILISIN SAVINASE | | Authors: | Frankaer, C.G, Moroz, O.V, Turkenburg, J.P, Aspmo, S.I, Thymark, M, Friis, E.P, Stahla, K, Nielsen, J.E, Wilson, K.S, Harris, P. | | Deposit date: | 2013-11-19 | | Release date: | 2014-04-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.17 Å) | | Cite: | Analysis of an Industrial Production Suspension of Bacillus Lentus Subtilisin Crystals by Powder Diffraction: A Powerful Quality-Control Tool.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4CFZ
 
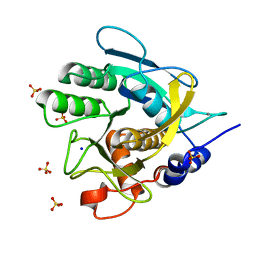 | | SAVINASE CRYSTAL STRUCTURES FOR COMBINED SINGLE CRYSTAL DIFFRACTION AND POWDER DIFFRACTION ANALYSIS | | Descriptor: | CALCIUM ION, SODIUM ION, SUBTILISIN SAVINASE, ... | | Authors: | Frankaer, C.G, Moroz, O.V, Turkenburg, J.P, Aspmo, S.I, Thymark, M, Friis, E.P, Stahla, K, Nielsen, J.E, Wilson, K.S, Harris, P. | | Deposit date: | 2013-11-19 | | Release date: | 2014-04-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | Analysis of an Industrial Production Suspension of Bacillus Lentus Subtilisin Crystals by Powder Diffraction: A Powerful Quality-Control Tool.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
6FXB
 
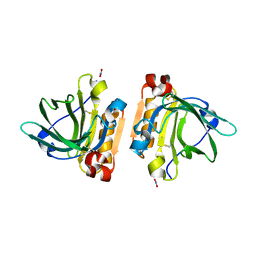 | | Bovine beta-lactoglobulin variant A at pH 4.0 | | Descriptor: | DI(HYDROXYETHYL)ETHER, Major allergen beta-lactoglobulin, NITRATE ION | | Authors: | Khan, S, Ipsen, R, Almdal, K, Svensson, B, Harris, P. | | Deposit date: | 2018-03-08 | | Release date: | 2018-05-23 | | Last modified: | 2019-02-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Revealing the Dimeric Crystal and Solution Structure of beta-Lactoglobulin at pH 4 and Its pH and Salt Dependent Monomer-Dimer Equilibrium.
Biomacromolecules, 19, 2018
|
|
3E2T
 
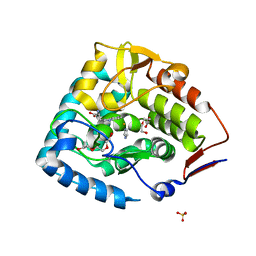 | | The catalytic domain of chicken tryptophan hydroxylase 1 with bound tryptophan | | Descriptor: | FE (III) ION, IMIDAZOLE, SULFATE ION, ... | | Authors: | Windahl, M.S, Petersen, C.R, Christensen, H.E.C, Harris, P. | | Deposit date: | 2008-08-06 | | Release date: | 2008-11-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of tryptophan hydroxylase with bound amino acid substrate
Biochemistry, 47, 2008
|
|
4M4J
 | | Radiation damage study of Cu T6-insulin - 0.30 MGy | | Descriptor: | COPPER (II) ION, Insulin | | Authors: | Frankaer, C.G, Harris, P, Stahl, K. | | Deposit date: | 2013-08-07 | | Release date: | 2014-01-15 | | Last modified: | 2018-03-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Towards accurate structural characterization of metal centres in protein crystals: the structures of Ni and Cu T6 bovine insulin derivatives.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4M4F
 | | Radiation damage study of Cu T6-insulin - 0.01 MGy | | Descriptor: | COPPER (II) ION, Insulin | | Authors: | Frankaer, C.G, Harris, P, Stahl, K. | | Deposit date: | 2013-08-07 | | Release date: | 2014-01-15 | | Last modified: | 2018-03-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Towards accurate structural characterization of metal centres in protein crystals: the structures of Ni and Cu T6 bovine insulin derivatives.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4M4M
 | | The structure of Ni T6 bovine insulin | | Descriptor: | Insulin, NICKEL (II) ION | | Authors: | Frankaer, C.G, Harris, P, Stahl, K. | | Deposit date: | 2013-08-07 | | Release date: | 2014-01-15 | | Last modified: | 2018-03-07 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Towards accurate structural characterization of metal centres in protein crystals: the structures of Ni and Cu T6 bovine insulin derivatives.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
