4D04
 
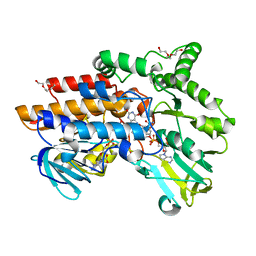 | | Structure of the Cys65Asp mutant of phenylacetone monooxygenase: reduced state | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, HEXAETHYLENE GLYCOL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Brondani, P.B, Dudek, H.M, Martinoli, C, Mattevi, A, Fraaije, M.W. | | Deposit date: | 2014-04-24 | | Release date: | 2014-12-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Finding the Switch: Turning a Baeyer-Villiger Monooxygenase Into a Nadph Oxidase.
J.Am.Chem.Soc., 136, 2014
|
|
6GZF
 
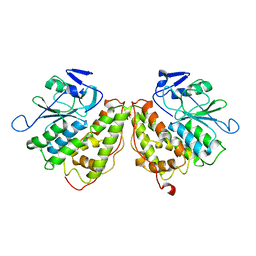 | | Xi Class GST from Natrialba magadii | | Descriptor: | Glutathione S-transferase, SULFATE ION | | Authors: | Di Matteo, A, Federici, L, Masulli, M, Carletti, E, Cassidy, J, Paradisi, F, Di Ilio, C, Allocati, N. | | Deposit date: | 2018-07-04 | | Release date: | 2019-02-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.614 Å) | | Cite: | Structural Characterization of the Xi Class Glutathione Transferase From the Haloalkaliphilic ArchaeonNatrialba magadii.
Front Microbiol, 10, 2019
|
|
6H4D
 
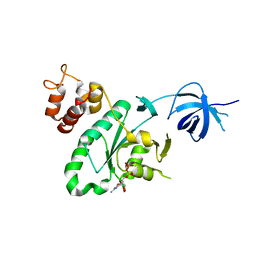 | | Crystal structure of RsgA from Pseudomonas aeruginosa | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, Small ribosomal subunit biogenesis GTPase RsgA, ZINC ION | | Authors: | Rocchio, S, Santorelli, D, Travaglini-Allocatelli, C, Federici, L, Di Matteo, A. | | Deposit date: | 2018-07-20 | | Release date: | 2019-06-26 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural and functional investigation of the Small Ribosomal Subunit Biogenesis GTPase A (RsgA) from Pseudomonas aeruginosa.
Febs J., 286, 2019
|
|
3GUS
 
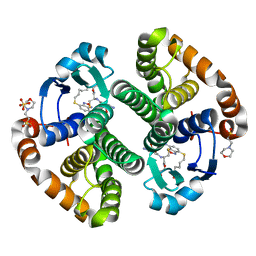 | | Crystal strcture of human Pi class glutathione S-transferase GSTP1-1 in complex with 6-(7-Nitro-2,1,3-benzoxadiazol-4-ylthio)hexanol (NBDHEX) | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 6-[(7-nitro-2,1,3-benzoxadiazol-4-yl)sulfanyl]hexan-1-ol, GLUTATHIONE, ... | | Authors: | Federici, L, Lo Sterzo, C, Di Matteo, A, Scaloni, F, Federici, G, Caccuri, A.M. | | Deposit date: | 2009-03-30 | | Release date: | 2009-10-27 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Structural basis for the binding of the anticancer compound 6-(7-nitro-2,1,3-benzoxadiazol-4-ylthio)hexanol to human glutathione s-transferases
Cancer Res., 69, 2009
|
|
1OJ9
 
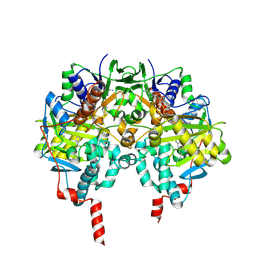 | | HUMAN MONOAMINE OXIDASE B IN COMPLEX WITH 1,4-DIPHENYL-2-BUTENE | | Descriptor: | 1,4-DIPHENYL-2-BUTENE, AMINE OXIDASE [FLAVIN-CONTAINING] B, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Binda, C, Edmondson, D.E, Mattevi, A. | | Deposit date: | 2003-07-08 | | Release date: | 2003-08-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Insights Into the Mode of Inhibition of Human Mitochondrial Monoamine Oxidase B from High-Resolution Crystal Structures
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1RZ0
 
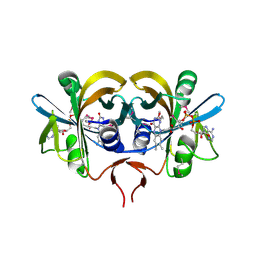 | | Flavin reductase PheA2 in native state | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, phenol 2-hydroxylase component B | | Authors: | van den Heuvel, R.H, Westphal, A.H, Heck, A.J, Walsh, M.A, Rovida, S, van Berkel, W.J, Mattevi, A. | | Deposit date: | 2003-12-23 | | Release date: | 2004-04-06 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural Studies on Flavin Reductase PheA2 Reveal Binding of NAD in an Unusual Folded Conformation and Support Novel Mechanism of Action.
J.Biol.Chem., 279, 2004
|
|
1OJC
 
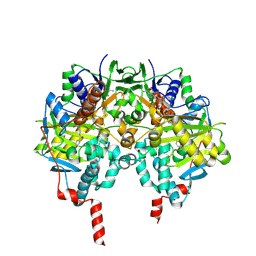 | |
2Z69
 
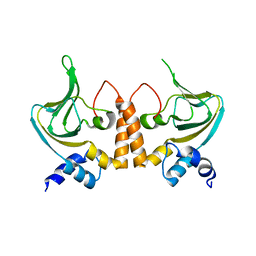 | |
1PBD
 
 | | CRYSTAL STRUCTURES OF WILD-TYPE P-HYDROXYBENZOATE HYDROXYLASE COMPLEXED WITH 4-AMINOBENZOATE, 2,4-DIHYDROXYBENZOATE AND 2-HYDROXY-4-AMINOBENZOATE AND OF THE TRY222ALA MUTANT, COMPLEXED WITH 2-HYDROXY-4-AMINOBENZOATE. EVIDENCE FOR A PROTON CHANNEL AND A NEW BINDING MODE OF THE FLAVIN RING | | Descriptor: | 4-AMINOBENZOIC ACID, FLAVIN-ADENINE DINUCLEOTIDE, P-HYDROXYBENZOATE HYDROXYLASE | | Authors: | Schreuder, H.A, Mattevi, A, Hol, W.G.J. | | Deposit date: | 1994-07-06 | | Release date: | 1994-09-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structures of wild-type p-hydroxybenzoate hydroxylase complexed with 4-aminobenzoate,2,4-dihydroxybenzoate, and 2-hydroxy-4-aminobenzoate and of the Tyr222Ala mutant complexed with 2-hydroxy-4-aminobenzoate. Evidence for a proton channel and a new binding mode of the flavin ring
Biochemistry, 33, 1994
|
|
1PBB
 
 | | CRYSTAL STRUCTURES OF WILD-TYPE P-HYDROXYBENZOATE HYDROXYLASE COMPLEXED WITH 4-AMINOBENZOATE, 2,4-DIHYDROXYBENZOATE AND 2-HYDROXY-4-AMINOBENZOATE AND OF THE TRY222ALA MUTANT, COMPLEXED WITH 2-HYDROXY-4-AMINOBENZOATE. EVIDENCE FOR A PROTON CHANNEL AND A NEW BINDING MODE OF THE FLAVIN RING | | Descriptor: | 2,4-DIHYDROXYBENZOIC ACID, FLAVIN-ADENINE DINUCLEOTIDE, P-HYDROXYBENZOATE HYDROXYLASE | | Authors: | Schreuder, H.A, Mattevi, A, Hol, W.G.J. | | Deposit date: | 1994-07-06 | | Release date: | 1994-09-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structures of wild-type p-hydroxybenzoate hydroxylase complexed with 4-aminobenzoate,2,4-dihydroxybenzoate, and 2-hydroxy-4-aminobenzoate and of the Tyr222Ala mutant complexed with 2-hydroxy-4-aminobenzoate. Evidence for a proton channel and a new binding mode of the flavin ring
Biochemistry, 33, 1994
|
|
2YG3
 
 | | Structure-based redesign of cofactor binding in Putrescine Oxidase: wild type enzyme | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, PUTRESCINE OXIDASE, ... | | Authors: | Kopacz, M.M, Rovida, S, van Duijn, E, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2011-04-11 | | Release date: | 2011-05-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure-Based Redesign of Cofactor Binding in Putrescine Oxidase.
Biochemistry, 50, 2011
|
|
2YG4
 
 | | Structure-based redesign of cofactor binding in Putrescine Oxidase: wild type bound to Putrescine | | Descriptor: | 4-HYDROXYBUTAN-1-AMINIUM, FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, ... | | Authors: | Kopacz, M.M, Rovida, S, van Duijn, E, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2011-04-11 | | Release date: | 2011-05-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure-Based Redesign of Cofactor Binding in Putrescine Oxidase.
Biochemistry, 50, 2011
|
|
2YG5
 
 | | Structure-based redesign of cofactor binding in Putrescine Oxidase: A394C mutant | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, PUTRESCINE OXIDASE, ... | | Authors: | Kopacz, M.M, Rovida, S, van Duijn, E, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2011-04-11 | | Release date: | 2011-05-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure-Based Redesign of Cofactor Binding in Putrescine Oxidase.
Biochemistry, 50, 2011
|
|
2YG6
 
 | | Structure-based redesign of cofactor binding in Putrescine Oxidase: P15I-A394C double mutant | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, PUTRESCINE OXIDASE, ... | | Authors: | Kopacz, M.M, Rovida, S, van Duijn, E, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2011-04-11 | | Release date: | 2011-05-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure-based redesign of cofactor binding in putrescine oxidase.
Biochemistry, 50, 2011
|
|
2YG7
 
 | | Structure-based redesign of cofactor binding in Putrescine Oxidase: A394C-A396T-Q431G Triple mutant | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, PUTRESCINE OXIDASE | | Authors: | Kopacz, M.M, Rovida, S, van Duijn, E, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2011-04-11 | | Release date: | 2011-05-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structure-Based Redesign of Cofactor Binding in Putrescine Oxidase.
Biochemistry, 50, 2011
|
|
1RZ1
 
 | | Reduced flavin reductase PheA2 in complex with NAD | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, phenol 2-hydroxylase component B | | Authors: | Van Den Heuvel, R.H, Westphal, A.H, Heck, A.J, Walsh, M.A, Rovida, S, Van Berkel, W.J, Mattevi, A. | | Deposit date: | 2003-12-23 | | Release date: | 2004-04-06 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Studies on Flavin Reductase PheA2 Reveal Binding of NAD in an Unusual Folded Conformation and Support Novel Mechanism of Action.
J.Biol.Chem., 279, 2004
|
|
1OJD
 
 | |
2NTO
 
 | | Structure of the Glutathione Transferase from Ochrobactrum anthropi in complex with glutathione | | Descriptor: | GLUTATHIONE, SULFATE ION, glutathione S-transferase | | Authors: | Federici, L, Bonivento, D, Di Matteo, A, Allocati, N. | | Deposit date: | 2006-11-08 | | Release date: | 2007-09-25 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.095 Å) | | Cite: | Role of Ser11 in the stabilization of the structure of Ochrobactrum anthropi glutathione transferase
Biochem.J., 403, 2007
|
|
1OJA
 
 | | HUMAN MONOAMINE OXIDASE B IN COMPLEX WITH ISATIN | | Descriptor: | AMINE OXIDASE [FLAVIN-CONTAINING] B, FLAVIN-ADENINE DINUCLEOTIDE, ISATIN | | Authors: | Binda, C, Edmondson, D.E, Mattevi, A. | | Deposit date: | 2003-07-08 | | Release date: | 2003-08-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Insights Into the Mode of Inhibition of Human Mitochondrial Monoamine Oxidase B from High-Resolution Crystal Structures
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
4A7A
 
 | | Crystal structure of human monoamine oxidase B (MAO B) in complex with rosiglitazone | | Descriptor: | (R)-ROSIGLITAZONE, AMINE OXIDASE [FLAVIN-CONTAINING] B, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Binda, C, Aldeco, M, Geldenhuys, W.J, Tortorici, M, Mattevi, A, Edmondson, D.E. | | Deposit date: | 2011-11-11 | | Release date: | 2012-04-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Molecular Insights Into Human Monoamine Oxidase B Inhibition by the Glitazone Anti-Diabetes Drugs
Acs Med. Chem. Lett., 3, 2012
|
|
4A79
 
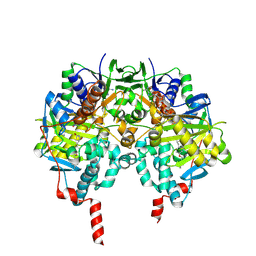 | | Crystal structure of human monoamine oxidase B (MAO B) in complex with pioglitazone | | Descriptor: | (5R)-5-{4-[2-(5-ethylpyridin-2-yl)ethoxy]benzyl}-1,3-thiazolidine-2,4-dione, AMINE OXIDASE [FLAVIN-CONTAINING] B, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Binda, C, Aldeco, M, Geldenhuys, W.J, Tortorici, M, Mattevi, A, Edmondson, D.E. | | Deposit date: | 2011-11-11 | | Release date: | 2012-04-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Molecular Insights Into Human Monoamine Oxidase B Inhibition by the Glitazone Anti-Diabetes Drugs
Acs Med. Chem. Lett., 3, 2012
|
|
4BAY
 
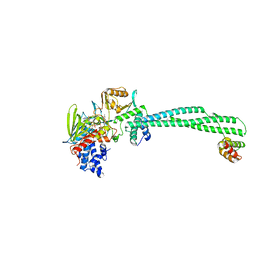 | | Phosphomimetic mutant of LSD1-8a splicing variant in complex with CoREST | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, LYSINE-SPECIFIC HISTONE DEMETHYLASE 1A, REST COREPRESSOR 1 | | Authors: | Toffolo, E, Paganini, L, Rusconi, F, Tortorici, M, Pilotto, S, Verpelli, C, Tedeschi, G, Maffioli, E, Sala, C, Mattevi, A, Battaglioli, E. | | Deposit date: | 2012-09-17 | | Release date: | 2013-11-27 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Phosphorylation of Neuronal Lysine-Specific Demethylase 1Lsd1/Kdm1A Impairs Transcriptional Repression by Regulating Interaction with Corest and Histone Deacetylases Hdac1/2.
J.Neurochem., 128, 2014
|
|
3ZYX
 
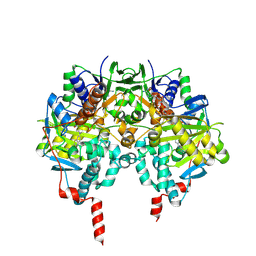 | | Crystal structure of human monoamine oxidase B in complex with methylene blue and bearing the double mutation I199A-Y326A | | Descriptor: | 3,7-BIS(DIMETHYLAMINO)PHENOTHIAZIN-5-IUM, AMINE OXIDASE [FLAVIN-CONTAINING] B, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Milczek, E.M, Binda, C, Rovida, S, Mattevi, A, Edmondson, D.E. | | Deposit date: | 2011-08-29 | | Release date: | 2011-12-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The 'Gating' Residues Ile199 and Tyr326 in Human Monoamine Oxidase B Function in Substrate and Inhibitor Recognition.
FEBS J., 278, 2011
|
|
5L3C
 
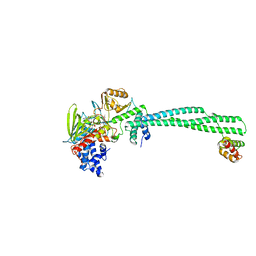 | | Human LSD1/CoREST: LSD1 E379K mutation | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, Lysine-specific histone demethylase 1A, REST corepressor 1 | | Authors: | Pilotto, S, Speranzini, V, Marabelli, C, Mattevi, A. | | Deposit date: | 2016-04-06 | | Release date: | 2016-05-04 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.31 Å) | | Cite: | LSD1/KDM1A mutations associated to a newly described form of intellectual disability impair demethylase activity and binding to transcription factors.
Hum.Mol.Genet., 25, 2016
|
|
5MLN
 
 | |
