4J1A
 
 | |
4N9R
 
 | | X-ray structure of the complex between hen egg white lysozyme and pentacholrocarbonyliridate(III) (1 day) | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Petruk, A.A, Bikiel, D.E, Vergara, A, Merlino, A. | | Deposit date: | 2013-10-21 | | Release date: | 2015-12-30 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Interaction between proteins and Ir based CO releasing molecules: mechanism of adduct formation and CO release.
Inorg.Chem., 53, 2014
|
|
4J1B
 
 | |
4KXH
 
 | | The X-ray crystal structure of a dimeric variant of human pancreatic ribonuclease | | Descriptor: | CHLORIDE ION, Ribonuclease pancreatic, SODIUM ION, ... | | Authors: | Pica, A, Merlino, A, Mazzarella, L, Sica, F. | | Deposit date: | 2013-05-26 | | Release date: | 2013-10-02 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Three-dimensional domain swapping and supramolecular protein assembly: insights from the X-ray structure of a dimeric swapped variant of human pancreatic RNase.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4L55
 
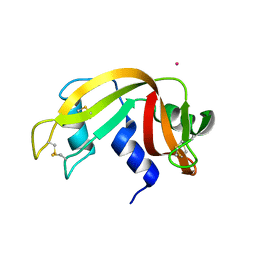 | |
8PH7
 
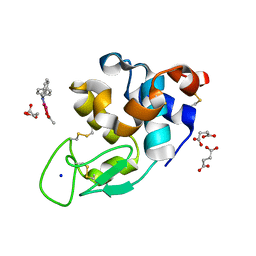 | | X-ray structure of the adduct formed upon reaction of Lysozyme with [Ru2Cl(DPhF)(O2CCH3)3] in condition A | | Descriptor: | Lysozyme C, SODIUM ION, SUCCINIC ACID, ... | | Authors: | Teran, A, Ferraro, G, Merlino, A. | | Deposit date: | 2023-06-19 | | Release date: | 2023-09-13 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.406 Å) | | Cite: | Steric hindrance and charge influence on the cytotoxic activity and protein binding properties of diruthenium complexes.
Int.J.Biol.Macromol., 253, 2023
|
|
8PH8
 
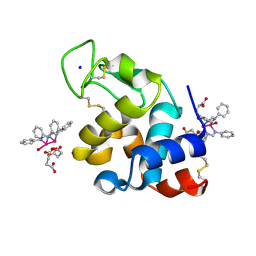 | | X-ray structure of the adduct formed upon reaction of Lysozyme with [Ru2Cl(DPhF)2(O2CCH3)2] in condition A | | Descriptor: | 1-oxidanyl-2,4,6,8-tetraphenyl-2,4,6,8-tetraza-1$l^{4},5$l^{3}-diruthenabicyclo[3.3.0]octane, Lysozyme C, SODIUM ION, ... | | Authors: | Teran, A, Ferraro, G, Merlino, A. | | Deposit date: | 2023-06-19 | | Release date: | 2023-09-13 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.29 Å) | | Cite: | Steric hindrance and charge influence on the cytotoxic activity and protein binding properties of diruthenium complexes.
Int.J.Biol.Macromol., 253, 2023
|
|
8PH5
 
 | | X-ray structure of the adduct formed upon reaction of Lysozyme with [Ru2Cl(DPhF)2(O2CCH3)2] in condition B | | Descriptor: | 1-oxidanyl-6,8,9,11-tetraphenyl-2,4-dioxa-6,8,9,11-tetraza-1$l^{5},5$l^{4}-diruthenatricyclo[3.3.3.0^{1,5}]undecane, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Lysozyme C | | Authors: | Teran, A, Ferraro, G, Merlino, A. | | Deposit date: | 2023-06-19 | | Release date: | 2023-09-13 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.29 Å) | | Cite: | Steric hindrance and charge influence on the cytotoxic activity and protein binding properties of diruthenium complexes.
Int.J.Biol.Macromol., 253, 2023
|
|
8PH6
 
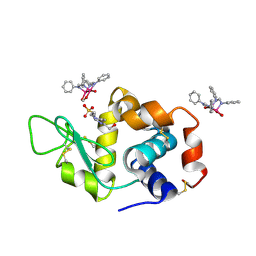
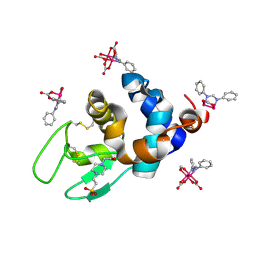 | | X-ray structure of the adduct formed upon reaction of Lysozyme with K2[Ru2(DPhF)(CO3)3] in condition B | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Lysozyme C, Ru2(DPhF)(CO3)2(Formate), ... | | Authors: | Teran, A, Ferraro, G, Merlino, A. | | Deposit date: | 2023-06-19 | | Release date: | 2023-09-13 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.07 Å) | | Cite: | Steric hindrance and charge influence on the cytotoxic activity and protein binding properties of diruthenium complexes.
Int.J.Biol.Macromol., 253, 2023
|
|
6ETN
 
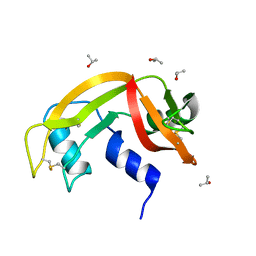 | |
8Q0S
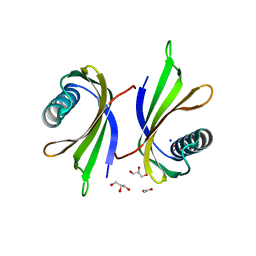 
 | | X-ray structure of the single chain monellin derivative MNEI | | Descriptor: | ACETATE ION, GLYCEROL, Monellin chain B,Monellin chain A, ... | | Authors: | Ferraro, G, Merlino, A, Lucignano, R, Picone, D. | | Deposit date: | 2023-07-29 | | Release date: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.19 Å) | | Cite: | Structural insights and aggregation propensity of a super-stable monellin mutant: A new potential building block for protein-based nanostructured materials.
Int.J.Biol.Macromol., 254, 2024
|
|
8Q0R
 
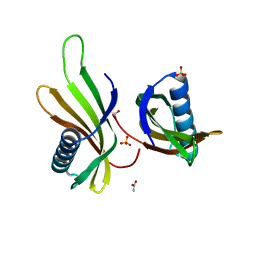 | | X-ray structure of MNEI mutant Mut9 (E23A, C41A, Y65R, S76Y) | | Descriptor: | ACETATE ION, Monellin chain B,Monellin chain A, SULFATE ION | | Authors: | Ferraro, G, Merlino, A, Lucignano, R, Picone, D. | | Deposit date: | 2023-07-29 | | Release date: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural insights and aggregation propensity of a super-stable monellin mutant: A new potential building block for protein-based nanostructured materials.
Int.J.Biol.Macromol., 254, 2024
|
|
7ZU6
 
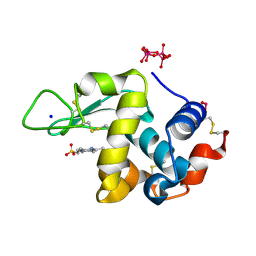 | | Polyoxidovanadate interaction with proteins: crystal structure of lysozyme bound to tetra-vanadate ion (structure 1) | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CYCLO-TETRAMETAVANADATE, Lysozyme, ... | | Authors: | Tito, G, Merlino, A, Ferraro, G. | | Deposit date: | 2022-05-11 | | Release date: | 2023-05-24 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.183 Å) | | Cite: | Stabilization and Binding of [V 4 O 12 ] 4- and Unprecedented [V 20 O 54 (NO 3 )] n- to Lysozyme upon Loss of Ligands and Oxidation of the Potential Drug V IV O(acetylacetonato) 2.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
7QHR
 
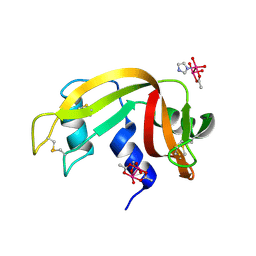 | |
8AJ4
 
 | | X-ray structure of lysozyme obtained upon reaction with [VIVO(malt)2] (Structure A') | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, 8,8-bis($l^{1}-oxidanyl)-2,2'-dimethyl-8,8'-spirobi[3$l^{4},7,9-trioxa-8$l^{6}-vanadabicyclo[4.3.0]nona-1(6),2,4-triene], Lysozyme, ... | | Authors: | Paolillo, M, Merlino, A, Ferraro, G. | | Deposit date: | 2022-07-27 | | Release date: | 2022-11-23 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.22 Å) | | Cite: | Multiple and Variable Binding of Pharmacologically Active Bis(maltolato)oxidovanadium(IV) to Lysozyme.
Inorg.Chem., 61, 2022
|
|
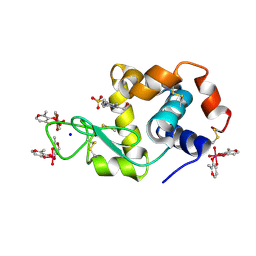
8AJ3
 
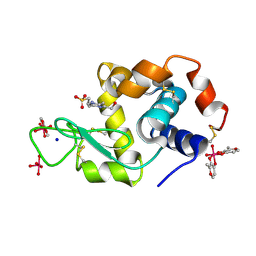 | | X-ray structure of lysozyme obtained upon reaction with [VIVO(malt)2] (Structure A) | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, 8,8,8,8-tetrakis($l^{1}-oxidanyl)-2-methyl-3,7,9-trioxa-8$l^{6}-vanadabicyclo[4.3.0]nona-1,5-diene, 8,8-bis($l^{1}-oxidanyl)-2,2'-dimethyl-8,8'-spirobi[3$l^{4},7,9-trioxa-8$l^{6}-vanadabicyclo[4.3.0]nona-1(6),2,4-triene], ... | | Authors: | Paolillo, M, Merlino, A, Ferraro, G. | | Deposit date: | 2022-07-27 | | Release date: | 2022-11-23 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | Multiple and Variable Binding of Pharmacologically Active Bis(maltolato)oxidovanadium(IV) to Lysozyme.
Inorg.Chem., 61, 2022
|
|
8AJ5
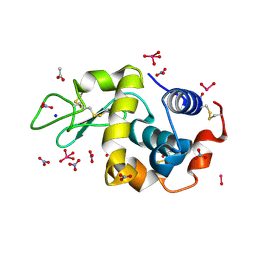 
 | | X-ray structure of lysozyme obtained upon reaction with [VIVO(malt)2] (Structure B) | | Descriptor: | ACETATE ION, Lysozyme, NITRATE ION, ... | | Authors: | Paolillo, M, Merlino, A, Ferraro, G. | | Deposit date: | 2022-07-27 | | Release date: | 2022-11-23 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.31 Å) | | Cite: | Multiple and Variable Binding of Pharmacologically Active Bis(maltolato)oxidovanadium(IV) to Lysozyme.
Inorg.Chem., 61, 2022
|
|
5L4R
 
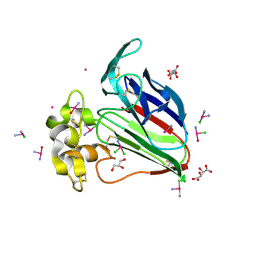 | | X-ray structure of the adduct between thaumatin and cisplatin | | Descriptor: | Cisplatin, GLYCEROL, L(+)-TARTARIC ACID, ... | | Authors: | Russo Krauss, I, Ferraro, G, Merlino, A. | | Deposit date: | 2016-05-26 | | Release date: | 2016-12-07 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Cisplatin-Protein Interactions: Unexpected Drug Binding to N-Terminal Amine and Lysine Side Chains.
Inorg.Chem., 55, 2016
|
|
4L0A
 
 | | X-ray structure of an all LNA quadruplex | | Descriptor: | DNA/RNA (5'-R(*(TLN)P*(LCG)P*(LCG)P*(LCG)P*(TLN))-3'), GLYCEROL, POTASSIUM ION | | Authors: | Russo Krauss, I, Parkinson, G, Merlino, A, Mazzarella, L, Sica, F. | | Deposit date: | 2013-05-31 | | Release date: | 2014-03-05 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A regular thymine tetrad and a peculiar supramolecular assembly in the first crystal structure of an all-LNA G-quadruplex.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4QGZ
 
 | |
9GYS
 
 | | X-ray structure of the adduct formed upon reaction of RNase A with [Ru2(D-p-FPhF)(O2CCH3)2(O2CO)] complex | | Descriptor: | Ribonuclease pancreatic, SULFATE ION, [Ru2-N,N'-diphenylformamidinate(O2CCH3)2(O2CO)] complex | | Authors: | Teran, A, Ferraro, G, Merlino, A. | | Deposit date: | 2024-10-02 | | Release date: | 2024-12-11 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Deciphering the role of neutral diruthenium complexes in protein binding.
Int.J.Biol.Macromol., 283, 2024
|
|
4QH3
 
 | |
4OT4
 
 | |
9FYW
 
 | | X-ray structure of the adduct formed upon reaction of RNase A with [Ru2Cl(D-p-CNPhF)(O2CCH3)3] | | Descriptor: | 4-[11-(4-cyanophenyl)-1,5-bis($l^{1}-oxidanyl)-3,7-dimethyl-2,4,6,8-tetraoxa-9,11-diaza-1$l^{5},5$l^{5}-diruthenatricyclo[3.3.3.0^{1,5}]undecan-9-yl]benzenecarbonitrile, Ribonuclease pancreatic, SULFATE ION | | Authors: | Teran, A, Ferraro, G, Merlino, A. | | Deposit date: | 2024-07-04 | | Release date: | 2024-11-27 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Exchange of equatorial ligands in protein-bound paddlewheel Ru25+ complexes: new insights from X-ray crystallography and quantum chemistry
Inorg Chem Front, 11, 2024
|
|
9EX0
 
 | | X-ray structure of a polyoxidovanadate/lysozyme adduct obtained when the protein is treated with [VIVO(acac)2] in 1.1 M NaCl, 0.1 M sodium acetate at pH 4.0 (Structure A) | | Descriptor: | CHLORIDE ION, Lysozyme C, Polyoxidovanadate complex, ... | | Authors: | Tito, G, Merlino, A, Ferraro, G. | | Deposit date: | 2024-04-05 | | Release date: | 2024-06-26 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Non-Covalent and Covalent Binding of New Mixed-Valence Cage-like Polyoxidovanadate Clusters to Lysozyme.
Angew.Chem.Int.Ed.Engl., 63, 2024
|
|
