1ZHH
 
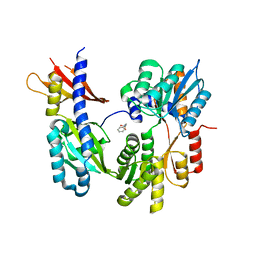 | | Crystal Structure of the Apo Form of Vibrio Harveyi LUXP Complexed with the Periplasmic Domain of LUXQ | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, Autoinducer 2 sensor kinase/phosphatase luxQ, Autoinducer 2-binding periplasmic protein luxP | | Authors: | Neiditch, M.B, Federle, M.J, Miller, S.T, Bassler, B.L, Hughson, F.M. | | Deposit date: | 2005-04-25 | | Release date: | 2005-05-24 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Regulation of LuxPQ Receptor Activity by the Quorum-Sensing Signal Autoinducer-2.
Mol.Cell, 18, 2005
|
|
2HJE
 
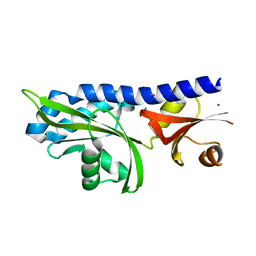 | |
2HJ9
 
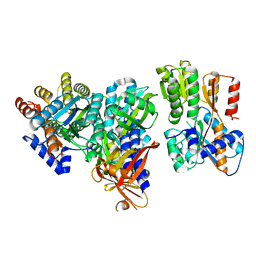 | | Crystal structure of the Autoinducer-2-bound form of Vibrio harveyi LuxP complexed with the periplasmic domain of LuxQ | | Descriptor: | 3A-METHYL-5,6-DIHYDRO-FURO[2,3-D][1,3,2]DIOXABOROLE-2,2,6,6A-TETRAOL, Autoinducer 2 sensor kinase/phosphatase luxQ, Autoinducer 2-binding periplasmic protein luxP | | Authors: | Neiditch, M.B, Hughson, F.M. | | Deposit date: | 2006-06-30 | | Release date: | 2006-09-26 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Ligand-induced asymmetry in histidine sensor kinase complex regulates quorum sensing.
Cell(Cambridge,Mass.), 126, 2006
|
|
4YV9
 
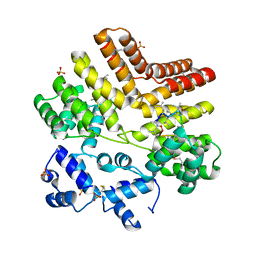 | |
4YV6
 
 | |
6W1E
 
 | |
6W1F
 | |
5W4N
 
 | |
7JI0
 
 | | CryoEM structure of Streptococcus thermophilus SHP pheromone receptor Rgg3 in complex with SHP3 | | Descriptor: | Positive transcriptional regulator MutR family, SHP3 | | Authors: | Petrou, V.I, Capodagli, G.C, Kaelber, J.T, Neiditch, M.B. | | Deposit date: | 2020-07-21 | | Release date: | 2020-10-07 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (2.95 Å) | | Cite: | Structure-function studies of Rgg binding to pheromones and target promoters reveal a model of transcription factor interplay.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
6DGJ
 
 | |
6DGG
 
 | |
3ULQ
 
 | |
8DFK
 
 | |
8DSS
 
 | |
6DGA
 
 | |
6DGN
 
 | |
4I1A
 
 | | Crystal Structure of the Apo Form of RapI | | Descriptor: | CHLORIDE ION, Response regulator aspartate phosphatase I | | Authors: | Parashar, V, Jeffrey, P.D, Neiditch, M.B. | | Deposit date: | 2012-11-20 | | Release date: | 2013-04-03 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.443 Å) | | Cite: | Conformational change-induced repeat domain expansion regulates rap phosphatase quorum-sensing signal receptors.
Plos Biol., 11, 2013
|
|
4GYO
 
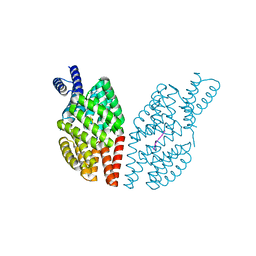 | |
3Q15
 
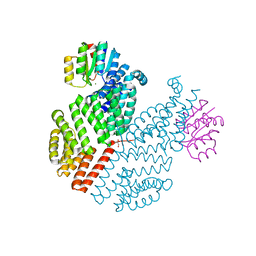 | | Crystal Structure of RapH complexed with Spo0F | | Descriptor: | GLYCEROL, MAGNESIUM ION, Response regulator aspartate phosphatase H, ... | | Authors: | Parashar, V, Neiditch, M.B. | | Deposit date: | 2010-12-16 | | Release date: | 2011-02-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.192 Å) | | Cite: | Structural basis of response regulator dephosphorylation by Rap phosphatases.
Plos Biol., 9, 2011
|
|
6W1A
 
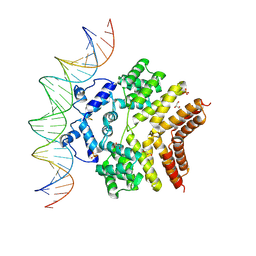 | |
6P9L
 
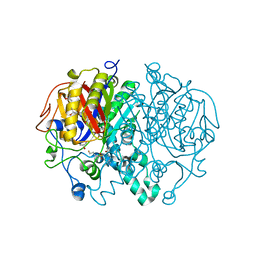 | |
6P9K
 
 | |
6P9M
 
 | |
6PRK
 
 | | X-ray Crystal Structure of Bacillus subtilis RicA in complex with RicF | | Descriptor: | RicA, RicF | | Authors: | Khaja, F.T, Jeffrey, P.D, Neiditch, M.B, Dubnau, D. | | Deposit date: | 2019-07-10 | | Release date: | 2019-10-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structure-Function Studies of the Bacillus subtilis Ric Proteins Identify the Fe-S Cluster-Ligating Residues and Their Roles in Development and RNA Processing.
Mbio, 10, 2019
|
|
6PRH
 
 | |
