5MGL
 
 | |
5MWG
 
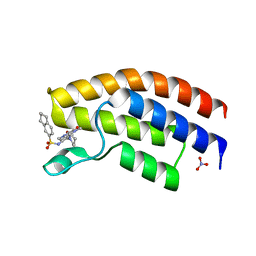 | | Crystal structure of the human BRPF1 bromodomain in complex with BZ091 | | Descriptor: | NITRATE ION, Peregrin, ~{N}-[1,4-dimethyl-2,3-bis(oxidanylidene)-7-piperidin-1-yl-quinoxalin-6-yl]-5,6,7,8-tetrahydronaphthalene-2-sulfonamide | | Authors: | Zhu, J, Caflisch, A. | | Deposit date: | 2017-01-18 | | Release date: | 2018-02-14 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structure-based discovery of selective BRPF1 bromodomain inhibitors.
Eur J Med Chem, 155, 2018
|
|
5MGE
 
 | |
7QWU
 
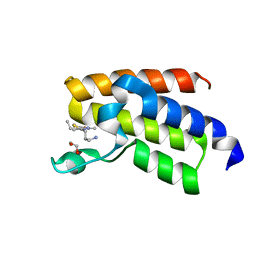 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 44 | | Descriptor: | 1,2-ETHANEDIOL, 1-[5-[2-(3-azanylpropyl)-1,3-thiazol-4-yl]-4-ethyl-2-methyl-1~{H}-pyrrol-3-yl]ethanone, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-25 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QVU
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative fragment 25 | | Descriptor: | 2-[4-(4-ethanoyl-3-ethyl-5-methyl-1~{H}-pyrrol-2-yl)-1,3-thiazol-2-yl]guanidine, Bromodomain adjacent to zinc finger domain protein 2A, MAGNESIUM ION | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-23 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QWF
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 45 | | Descriptor: | 1,2-ETHANEDIOL, 1-[4-ethyl-2-methyl-5-[2-[2-(methylamino)ethyl]-1,3-thiazol-4-yl]-1~{H}-pyrrol-3-yl]ethanone, Bromodomain adjacent to zinc finger domain protein 2A, ... | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-25 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QWY
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 61 | | Descriptor: | 1-[4-ethyl-2-methyl-5-[2-(piperazin-1-ylmethyl)-1,3-thiazol-4-yl]-1~{H}-pyrrol-3-yl]ethanone, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-26 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.443 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QX9
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 65 | | Descriptor: | 3-[[4-(4-ethanoyl-3,5-dimethyl-1~{H}-pyrrol-2-yl)-1,3-thiazol-2-yl]methyl]imidazolidine-2,4-dione, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-26 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QZ0
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 83 | | Descriptor: | 2-[4-(3-methyl-4-oxidanylidene-2,5,6,7-tetrahydroisoindol-1-yl)-1,3-thiazol-2-yl]guanidine, Bromodomain adjacent to zinc finger domain protein 2A, MAGNESIUM ION | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-30 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QZI
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 113 | | Descriptor: | (2~{R})-~{N}-(2,2-dimethylpropyl)-1-[4-(4-ethanoyl-3-ethyl-5-methyl-1~{H}-pyrrol-2-yl)-1,3-thiazol-2-yl]piperazine-2-carboxamide, 1,2-ETHANEDIOL, Bromodomain adjacent to zinc finger domain protein 2A, ... | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-31 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QZT
 
 | | BAZ2A bromodomain in complex with isoquinoline derivative fragment 9 | | Descriptor: | Bromodomain adjacent to zinc finger domain protein 2A, ~{N}-[(3~{S})-pyrrolidin-3-yl]isoquinoline-5-sulfonamide | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-31 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.183 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QYV
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 109 | | Descriptor: | (2~{R})-1-[4-(4-ethanoyl-3-ethyl-5-methyl-1~{H}-pyrrol-2-yl)-1,3-thiazol-2-yl]-~{N}-(4-oxidanylbutyl)piperazine-2-carboxamide, Bromodomain adjacent to zinc finger domain protein 2A, MAGNESIUM ION | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-29 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QYW
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 111 | | Descriptor: | (2~{R})-1-[4-(4-ethanoyl-3-ethyl-5-methyl-1~{H}-pyrrol-2-yl)-1,3-thiazol-2-yl]-~{N}-[3,3,3-tris(fluoranyl)propyl]piperazine-2-carboxamide, Bromodomain adjacent to zinc finger domain protein 2A, MAGNESIUM ION | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-29 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QYT
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 80 | | Descriptor: | 3-methyl-1-(2-piperazin-1-yl-1,3-thiazol-4-yl)-2,5,6,7-tetrahydroisoindol-4-one, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-29 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QYU
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 88 | | Descriptor: | 2-[4-[[4-[4-(4-ethanoyl-3-ethyl-5-methyl-1~{H}-pyrrol-2-yl)-1,3-thiazol-2-yl]piperazin-1-yl]methyl]-1,2,3-triazol-1-yl]ethanamide, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-29 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7R0B
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 47 | | Descriptor: | 1,2-ETHANEDIOL, 1-[4-ethyl-2-methyl-5-(2-piperazin-1-yl-1,3-thiazol-4-yl)-1~{H}-pyrrol-3-yl]ethanone, Bromodomain adjacent to zinc finger domain protein 2A, ... | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-02-01 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.345 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QZB
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 98 | | Descriptor: | 1,2-ETHANEDIOL, 3-methyl-1-[2-[4-[(4-methyl-1~{H}-pyrazol-3-yl)methyl]piperazin-1-yl]-1,3-thiazol-4-yl]-2,5,6,7-tetrahydroisoindol-4-one, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-31 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.151 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QVV
 | | BAZ2A bromodomain in complex with pyridone derivative fragment 36 | | Descriptor: | 5-(4-azanylpiperidin-1-yl)carbonyl-1~{H}-pyridin-2-one, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-23 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.601 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QX2
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 63 | | Descriptor: | 1-[5-[2-[(4-azanylpiperidin-1-yl)methyl]-1,3-thiazol-4-yl]-4-ethyl-2-methyl-1~{H}-pyrrol-3-yl]ethanone, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-26 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.428 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QYO
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 79 | | Descriptor: | 1-[2-methyl-4-(3-methylbutyl)-5-(2-piperazin-1-yl-1,3-thiazol-4-yl)-1~{H}-pyrrol-3-yl]ethanone, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-28 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (0.983 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QZ4
 
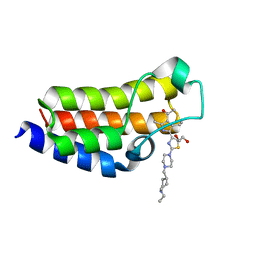 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 87 | | Descriptor: | 1,2-ETHANEDIOL, 1-[4-ethyl-5-[2-[4-[(1-ethylpiperidin-4-yl)methyl]piperazin-1-yl]-1,3-thiazol-4-yl]-2-methyl-1~{H}-pyrrol-3-yl]ethanone, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-30 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QZC
 
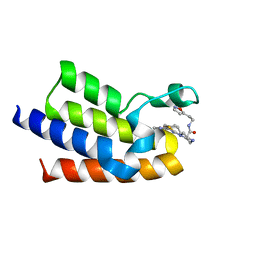 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 104 | | Descriptor: | (2R)-1-[4-(4-ethanoyl-3-ethyl-5-methyl-1H-pyrrol-2-yl)-1,3-thiazol-2-yl]-N-[2-(1,2-oxazol-5-yl)ethyl]piperazine-2-carboxamide, Bromodomain adjacent to zinc finger domain protein 2A, MAGNESIUM ION | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-31 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7R01
 
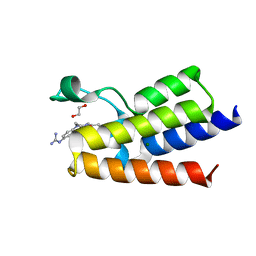 | | BAZ2A bromodomain in complex with N-acetylpiperazine derivative fragment 18 | | Descriptor: | 1,2-ETHANEDIOL, 1-[4-(4-ethanoylpiperazin-1-yl)phenyl]guanidine, Bromodomain adjacent to zinc finger domain protein 2A, ... | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-02-01 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.256 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QVT
 
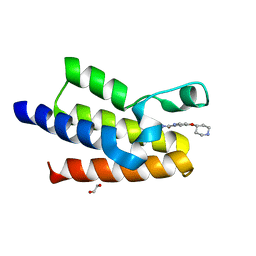 | | BAZ2A bromodomain in complex with picolinamide derivative fragment 13 | | Descriptor: | 1,2-ETHANEDIOL, 4-piperidin-4-yloxypyridine-2-carboxamide, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-23 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
7QXL
 
 | | BAZ2A bromodomain in complex with acetylpyrrole derivative compound 77 | | Descriptor: | 1-[2-methyl-5-(2-piperazin-1-yl-1,3-thiazol-4-yl)-4-[3,3,3-tris(fluoranyl)propyl]-1~{H}-pyrrol-3-yl]ethanone, Bromodomain adjacent to zinc finger domain protein 2A | | Authors: | Dalle Vedove, A, Cazzanelli, G, Caflisch, A, Lolli, G. | | Deposit date: | 2022-01-26 | | Release date: | 2022-09-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Identification of a BAZ2A-Bromodomain Hit Compound by Fragment Growing.
Acs Med.Chem.Lett., 13, 2022
|
|
