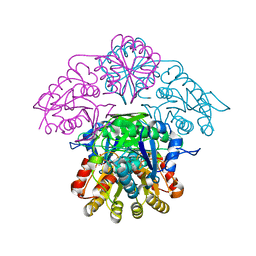
5K4G
 
 | |
5KK3
 
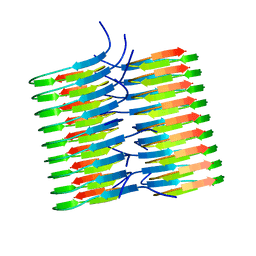 | | Atomic Resolution Structure of Monomorphic AB42 Amyloid Fibrils | | Descriptor: | Beta-amyloid protein 42 | | Authors: | Colvin, M.T, Silvers, R, Zhe Ni, Q, Can, T.V, Sergeyev, I, Rosay, M, Donovan, K.J, Michael, B, Wall, J, Linse, S, Griffin, R.G. | | Deposit date: | 2016-06-20 | | Release date: | 2016-07-13 | | Last modified: | 2024-05-01 | | Method: | SOLID-STATE NMR | | Cite: | Atomic Resolution Structure of Monomorphic A beta 42 Amyloid Fibrils.
J.Am.Chem.Soc., 138, 2016
|
|
7RYD
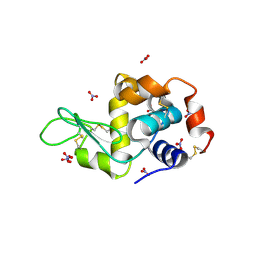 
 | | Hen egg-white lysozyme with ionic liquid butylammonium nitrate 1 mol% | | Descriptor: | Lysozyme C, NITRATE ION | | Authors: | Han, Q, Darmanin, C, Smith, K, Drummond, C, Greaves, T. | | Deposit date: | 2021-08-25 | | Release date: | 2023-03-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.18 Å) | | Cite: | Probing ion-binding at a protein interface: Modulation of protein properties by ionic liquids.
J Colloid Interface Sci, 650, 2023
|
|
7RZ0
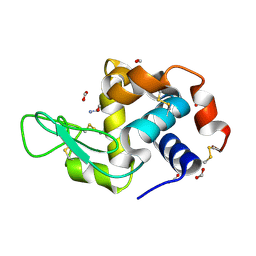 
 | | Hen egg-white lysozyme with ionic liquid ethanolammonium formate 6.7 mol% | | Descriptor: | ETHANOLAMINE, FORMIC ACID, Lysozyme C | | Authors: | Han, Q, Darmanin, C, Drummond, C, Greaves, T. | | Deposit date: | 2021-08-27 | | Release date: | 2023-03-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Probing ion-binding at a protein interface: Modulation of protein properties by ionic liquids.
J Colloid Interface Sci, 650, 2023
|
|
7RZ1
 
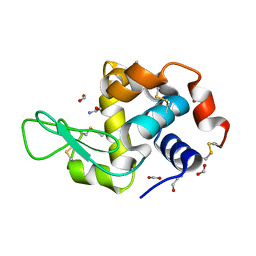 | | Hen egg-white lysozyme with ionic liquid ethanolammonium formate 14.4 mol% | | Descriptor: | ETHANOLAMINE, FORMIC ACID, Lysozyme C | | Authors: | Han, Q, Darmanin, C, Drummond, C, Greaves, T. | | Deposit date: | 2021-08-27 | | Release date: | 2023-03-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.046 Å) | | Cite: | Probing ion-binding at a protein interface: Modulation of protein properties by ionic liquids.
J Colloid Interface Sci, 650, 2023
|
|
7RZ2
 
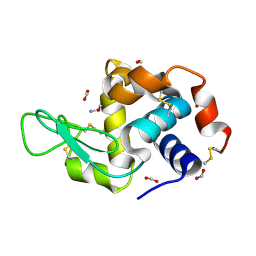 | | Hen egg-white lysozyme with ionic liquid ethanolammonium formate 4 mol% | | Descriptor: | ETHANOLAMINE, FORMIC ACID, Lysozyme C | | Authors: | Han, Q, Darmanin, C, Drummond, C, Greaves, T. | | Deposit date: | 2021-08-27 | | Release date: | 2023-03-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.07 Å) | | Cite: | Probing ion-binding at a protein interface: Modulation of protein properties by ionic liquids.
J Colloid Interface Sci, 650, 2023
|
|
7F0H
 
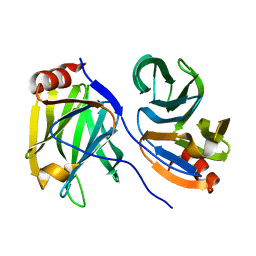 | |
6IRR
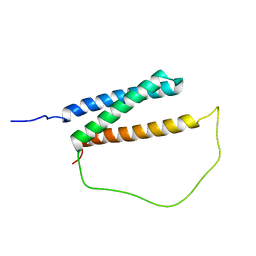 
 | | Solution structure of DISC1/ATF4 complex | | Descriptor: | Disrupted in schizophrenia 1 homolog,Cyclic AMP-dependent transcription factor ATF-4 | | Authors: | Ye, F, Yu, C, Zhang, M. | | Deposit date: | 2018-11-14 | | Release date: | 2019-09-25 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural interaction between DISC1 and ATF4 underlying transcriptional and synaptic dysregulation in an iPSC model of mental disorders.
Mol. Psychiatry, 2019
|
|
6MOB
 
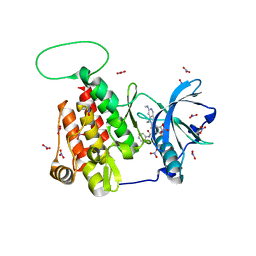 | | Crystal structure of KIT1 in complex with DP2976 via co-crystallization | | Descriptor: | Mast/stem cell growth factor receptor Kit, N-{4-chloro-5-[1-ethyl-7-(methylamino)-2-oxo-1,2-dihydro-1,6-naphthyridin-3-yl]-2-fluorophenyl}-N'-phenylurea, NITRATE ION | | Authors: | Edwards, T.E, Abendroth, J, Safford, K, Chun, L. | | Deposit date: | 2018-10-04 | | Release date: | 2019-07-31 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Ripretinib (DCC-2618) Is a Switch Control Kinase Inhibitor of a Broad Spectrum of Oncogenic and Drug-Resistant KIT and PDGFRA Variants.
Cancer Cell, 35, 2019
|
|
1C3E
 
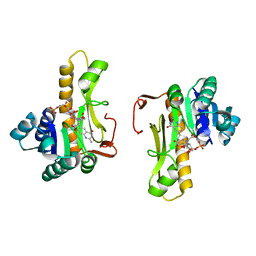 | | NEW INSIGHTS INTO INHIBITOR DESIGN FROM THE CRYSTAL STRUCTURE AND NMR STUDIES OF E. COLI GAR TRANSFORMYLATE IN COMPLEX WITH BETA-GAR AND 10-FORMYL-5,8,10-TRIDEAZAFOLIC ACID. | | Descriptor: | 2-{4-[2-(2-AMINO-4-HYDROXY-QUINAZOLIN-6-YL)-1-CARBOXY-ETHYL]-BENZOYLAMINO}-PENTANEDIOIC ACID, GLYCINAMIDE RIBONUCLEOTIDE, GLYCINAMIDE RIBONUCLEOTIDE TRANSFORMYLASE | | Authors: | Greasley, S.E, Yamashita, M.M, Cai, H, Benkovic, S.J, Boger, D.L, Wilson, I.A. | | Deposit date: | 1999-07-27 | | Release date: | 1999-12-29 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | New insights into inhibitor design from the crystal structure and NMR studies of Escherichia coli GAR transformylase in complex with beta-GAR and 10-formyl-5,8,10-trideazafolic acid.
Biochemistry, 38, 1999
|
|
1C2T
 
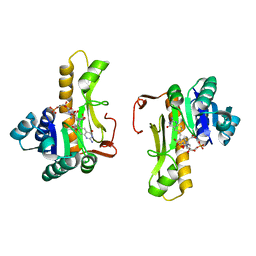 | | NEW INSIGHTS INTO INHIBITOR DESIGN FROM THE CRYSTAL STRUCTURE AND NMR STUDIES OF E. COLI GAR TRANSFORMYLASE IN COMPLEX WITH BETA-GAR AND 10-FORMYL-5,8,10-TRIDEAZAFOLIC ACID. | | Descriptor: | 10-FORMYL-5,8,10-TRIDEAZAFOLIC ACID, GLYCINAMIDE RIBONUCLEOTIDE, GLYCINAMIDE RIBONUCLEOTIDE TRANSFORMYLASE | | Authors: | Greasley, S.E, Yamashita, M.M, Cai, H, Benkovic, S.J, Boger, D.L, Wilson, I.A. | | Deposit date: | 1999-07-26 | | Release date: | 2000-01-05 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | New insights into inhibitor design from the crystal structure and NMR studies of Escherichia coli GAR transformylase in complex with beta-GAR and 10-formyl-5,8,10-trideazafolic acid.
Biochemistry, 38, 1999
|
|
