3RDC
 
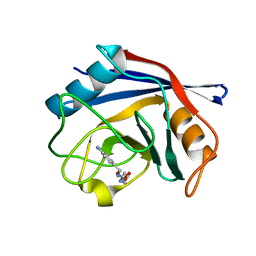 | | Human Cyclophilin D Complexed with an Inhibitor | | Descriptor: | Peptidyl-prolyl cis-trans isomerase F, mitochondrial, ethyl N-[(4-aminobenzyl)carbamoyl]glycinate | | Authors: | Colliandre, L, Ahmed-Belkacem, H, Bessin, Y, Pawlotsky, J.M, Guichou, J.F. | | Deposit date: | 2011-04-01 | | Release date: | 2012-03-21 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Combining 'dry' co-crystallization and in situ diffraction to facilitate ligand screening by X-ray crystallography.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
7YXY
 
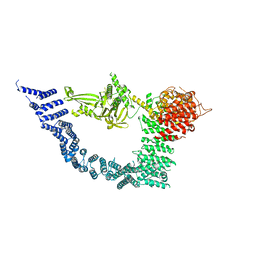 | |
7YXX
 
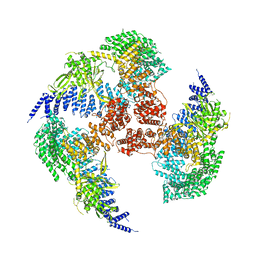 | |
2LB6
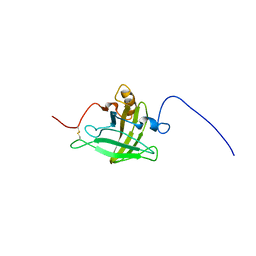 
 | | Structure of 18694Da MUP, typical to the major urinary protein family: MUP9, MUP11, MUP15, MUP18 & MUP19 | | Descriptor: | Major urinary protein 6 | | Authors: | Phelan, M.M, Mclean, L, Beynon, R.J, Hurst, J.L, Lian, L. | | Deposit date: | 2011-03-23 | | Release date: | 2012-03-28 | | Last modified: | 2023-12-06 | | Method: | SOLUTION NMR | | Cite: | Structural insights into the specificity of darcin, an atypical major urinary protein.
To be Published
|
|
2L9C
 
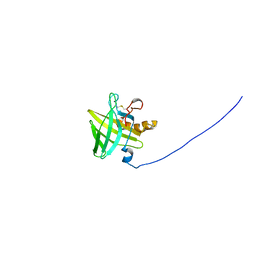 | | Structural insights into the specificity of darcin, an atypical major urinary protein. | | Descriptor: | Darcin | | Authors: | Phelan, M.M, Mclean, L, Beynon, R.J, Hurst, J.L, Lian, L. | | Deposit date: | 2011-02-07 | | Release date: | 2012-02-08 | | Last modified: | 2023-12-06 | | Method: | SOLUTION NMR | | Cite: | Structural insights into the specificity of darcin, an atypical major urinary protein.
To be Published
|
|
2MHO
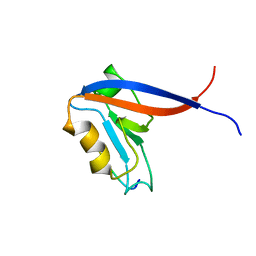 
 | |
6RP5
 
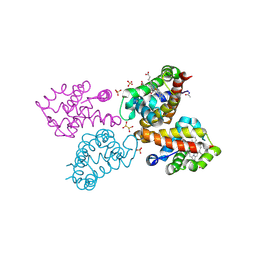 | | Crystal structure of monocarboxylated hemoglobin from the sub-Antarctic fish Eleginops maclovinus | | Descriptor: | CARBON MONOXIDE, DITHIONITE, Hemoglobin subunit alpha 1, ... | | Authors: | Balasco, N, Vitagliano, L, Merlino, A, Verde, C, Mazzarella, L, Vergara, A. | | Deposit date: | 2019-05-13 | | Release date: | 2019-12-25 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | The unique structural features of carbonmonoxy hemoglobin from the sub-Antarctic fish Eleginops maclovinus.
Sci Rep, 9, 2019
|
|
1EF7
 
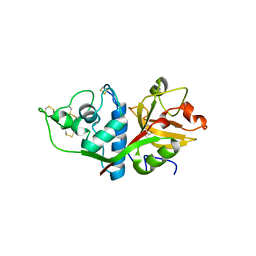 | | CRYSTAL STRUCTURE OF HUMAN CATHEPSIN X | | Descriptor: | CATHEPSIN X | | Authors: | Guncar, G, Klemencic, I, Turk, B, Turk, V, Karaoglanovic-Carmona, A, Juliano, L, Turk, D. | | Deposit date: | 2000-02-07 | | Release date: | 2000-03-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.67 Å) | | Cite: | Crystal structure of cathepsin X: a flip-flop of the ring of His23 allows carboxy-monopeptidase and carboxy-dipeptidase activity of the protease.
Structure Fold.Des., 8, 2000
|
|
1VYH
 
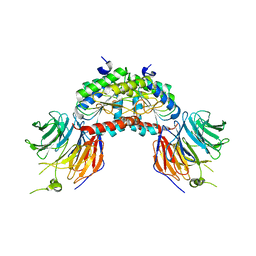 | | PAF-AH Holoenzyme: Lis1/Alfa2 | | Descriptor: | PLATELET-ACTIVATING FACTOR ACETYLHYDROLASE IB ALPHA SUBUNIT, PLATELET-ACTIVATING FACTOR ACETYLHYDROLASE IB BETA SUBUNIT | | Authors: | Tarricone, C, Perrina, F, Monzani, S, Massimiliano, L, Knapp, S, Tsai, L.-H, Derewenda, Z.S, Musacchio, A. | | Deposit date: | 2004-04-30 | | Release date: | 2005-05-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Coupling Paf Signaling to Dynein Regulation: Structure of Lis1 in Complex with Paf-Acetylhydrolase.
Neuron, 44, 2004
|
|
4PSH
 
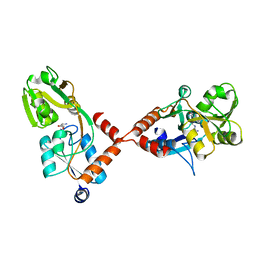 | | Structure of holo ArgBP from T. maritima | | Descriptor: | ABC-type transporter, periplasmic subunit family 3, ARGININE | | Authors: | Ruggiero, A, Dattelbaum, J.D, Staiano, M, Berisio, R, D'Auria, S, Vitagliano, L. | | Deposit date: | 2014-03-07 | | Release date: | 2014-07-23 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | A loose domain swapping organization confers a remarkable stability to the dimeric structure of the arginine binding protein from Thermotoga maritima
Plos One, 9, 2014
|
|
4PRS
 
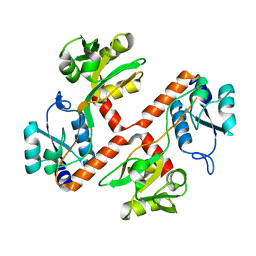 | | Structure of apo ArgBP from T. maritima | | Descriptor: | ABC-type transporter, periplasmic subunit family 3 | | Authors: | Ruggiero, A, Dattelbaum, J.D, Staiano, M, Berisio, R, D'Auria, S, Vitagliano, L. | | Deposit date: | 2014-03-06 | | Release date: | 2014-07-23 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | A loose domain swapping organization confers a remarkable stability to the dimeric structure of the arginine binding protein from Thermotoga maritima
Plos One, 9, 2014
|
|
7RUO
 
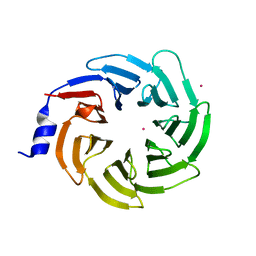 | | Crystal structure of human UTP15 | | Descriptor: | U3 small nucleolar RNA-associated protein 15 homolog, UNKNOWN ATOM OR ION | | Authors: | Dehghani-Tafti, S, Dong, A, Zeng, H, Hutchinson, A, Seitova, A, Loppnau, P, Arrowsmith, C.H, Edwards, A.M, Halabelian, L, Structural Genomics Consortium (SGC) | | Deposit date: | 2021-08-17 | | Release date: | 2021-11-10 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of human UTP15
To Be Published
|
|
1K6F
 | |
3NFE
 
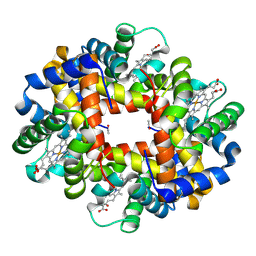 | | The crystal structure of hemoglobin I from trematomus newnesi in deoxygenated state | | Descriptor: | Hemoglobin subunit alpha-1, Hemoglobin subunit beta-1/2, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Vergara, A, Vitagliano, L, Merlino, A, Sica, F, Marino, K, Mazzarella, L. | | Deposit date: | 2010-06-10 | | Release date: | 2010-07-07 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | An order-disorder transition plays a role in switching off the root effect in fish hemoglobins.
J.Biol.Chem., 285, 2010
|
|
7RBQ
 
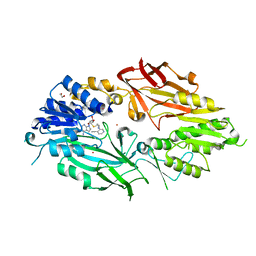 | | Co-crystal structure of human PRMT9 in complex with MT556 inhibitor | | Descriptor: | 1,2-ETHANEDIOL, 7-[5-S-(4-{[(4-ethylpyridin-3-yl)methyl]amino}butyl)-5-thio-beta-D-ribofuranosyl]-7H-pyrrolo[2,3-d]pyrimidin-4-amine, Protein arginine N-methyltransferase 9, ... | | Authors: | Zeng, H, Dong, A, Hutchinson, A, Seitova, A, Li, Y, Gao, Y.D, Schneider, S, Siliphaivanh, P, Sloman, D, Nicholson, B, Fischer, C, Hicks, J, Brown, P.J, Arrowsmith, C.H, Edwards, A.M, Halabelian, L, Structural Genomics Consortium (SGC) | | Deposit date: | 2021-07-06 | | Release date: | 2021-08-11 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Co-crystal structure of human PRMT9 in complex with MT556 inhibitor
To Be Published
|
|
3NG6
 
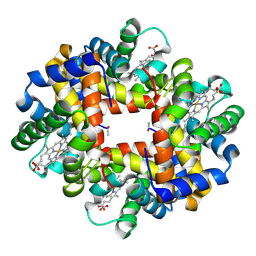 | | The crystal structure of hemoglobin I from Trematomus newnesi in deoxygenated state obtained through an oxidation/reduction cycle in which potassium hexacyanoferrate and sodium dithionite were alternatively added | | Descriptor: | Hemoglobin subunit alpha-1, Hemoglobin subunit beta-1/2, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Vergara, A, Vitagliano, L, Merlino, A, Sica, F, Marino, K, Mazzarella, L. | | Deposit date: | 2010-06-11 | | Release date: | 2010-07-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | An order-disorder transition plays a role in switching off the root effect in fish hemoglobins.
J.Biol.Chem., 285, 2010
|
|
1LA6
 
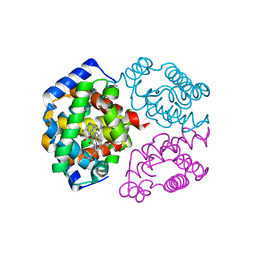 | | The crystal structure of Trematomus newnesi hemoglobin in a partial hemichrome state | | Descriptor: | CARBON MONOXIDE, Hemoglobin alpha-1 chain, Hemoglobin beta-1/2 chain, ... | | Authors: | Riccio, A, Vitagliano, L, di Prisco, G, Zagari, A, Mazzarella, L. | | Deposit date: | 2002-03-28 | | Release date: | 2002-07-31 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The crystal structure of a tetrameric hemoglobin in a partial hemichrome state
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
8G5E
 
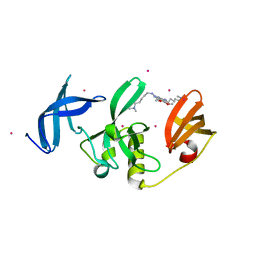 | | Crystal Structure of SETDB1 Tudor domain in complex with UNC6535 | | Descriptor: | Histone-lysine N-methyltransferase SETDB1, N~4~-[6-(dimethylamino)hexyl]-N~2~-[5-(dimethylamino)pentyl]-6,7-dimethoxyquinazoline-2,4-diamine, UNKNOWN ATOM OR ION | | Authors: | Beldar, S, Dong, A, Brown, P.J, Arrowsmith, C.H, Edwards, A.M, Halabelian, L, Structural Genomics Consortium (SGC) | | Deposit date: | 2023-02-13 | | Release date: | 2023-02-22 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Crystal Structure of SETDB1 Tudor domain in complex with UNC6535
To be published
|
|
8GCY
 
 | | Co-crystal structure of CBL-B in complex with N-Aryl isoindolin-1-one inhibitor | | Descriptor: | 1,2-ETHANEDIOL, 2-{3-[(1s,3R)-3-methyl-1-(4-methyl-4H-1,2,4-triazol-3-yl)cyclobutyl]phenyl}-6-{[(3S)-3-methylpiperidin-1-yl]methyl}-4-(trifluoromethyl)-2,3-dihydro-1H-isoindol-1-one, E3 ubiquitin-protein ligase CBL-B, ... | | Authors: | Kimani, S, Zeng, H, Dong, A, Li, Y, Santhakumar, V, Arrowsmith, C.H, Edwards, A.M, Halabelian, L, Structural Genomics Consortium (SGC) | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-22 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | The co-crystal structure of Cbl-b and a small-molecule inhibitor reveals the mechanism of Cbl-b inhibition.
Commun Biol, 6, 2023
|
|
1DY5
 
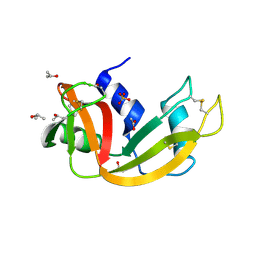 | | Deamidated derivative of bovine pancreatic ribonuclease | | Descriptor: | ACETATE ION, ISOPROPYL ALCOHOL, RIBONUCLEASE A, ... | | Authors: | Esposito, L, Vitagliano, L, Sica, F, Zagari, A, Mazzarella, L. | | Deposit date: | 2000-01-27 | | Release date: | 2000-03-28 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (0.87 Å) | | Cite: | The Ultrahigh Resolution Crystal Structure of Ribonuclease A Containing an Isoaspartyl Residue: Hydration and Sterochemical Analysis.
J.Mol.Biol., 297, 2000
|
|
2H8F
 | | Crystal structure of deoxy hemoglobin from Trematomus bernacchii at pH 6.2 | | Descriptor: | Hemoglobin alpha subunit, Hemoglobin beta subunit, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Mazzarella, L, Vergara, A, Vitagliano, L, Merlino, A, Bonomi, G, Scala, S, Verde, C, di Prisco, G. | | Deposit date: | 2006-06-07 | | Release date: | 2006-08-29 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | High resolution crystal structure of deoxy hemoglobin from Trematomus bernacchii at different pH values: The role of histidine residues in modulating the strength of the root effect.
Proteins, 65, 2006
|
|
5E27
 
 | | The structure of Resuscitation Promoting Factor B from M. tuberculosis reveals unexpected ubiquitin-like domains | | Descriptor: | Resuscitation-promoting factor RpfB | | Authors: | Ruggiero, A, Squeglia, F, Romano, M, Vitagliano, L, De Simone, A, Berisio, R. | | Deposit date: | 2015-09-30 | | Release date: | 2015-11-18 | | Last modified: | 2015-12-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The structure of Resuscitation promoting factor B from M. tuberculosis reveals unexpected ubiquitin-like domains.
Biochim.Biophys.Acta, 1860, 2015
|
|
2H8D
 | | Crystal structure of deoxy hemoglobin from Trematomus bernacchii at pH 8.4 | | Descriptor: | Hemoglobin alpha subunit, Hemoglobin beta subunit, POTASSIUM ION, ... | | Authors: | Mazzarella, L, Vergara, A, Vitagliano, L, Merlino, A, Bonomi, G, Scala, S, Verde, C, di Prisco, G. | | Deposit date: | 2006-06-07 | | Release date: | 2006-08-29 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | High resolution crystal structure of deoxy hemoglobin from Trematomus bernacchii at different pH values: The role of histidine residues in modulating the strength of the root effect.
Proteins, 65, 2006
|
|
1LSQ
 | | RIBONUCLEASE A WITH ASN 67 REPLACED BY A BETA-ASPARTYL RESIDUE | | Descriptor: | RIBONUCLEASE A, SULFATE ION | | Authors: | Esposito, L, Sica, F, Vitagliano, L, Zagari, A, Mazzarella, L. | | Deposit date: | 1996-06-25 | | Release date: | 1997-01-11 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Deamidation in proteins: the crystal structure of bovine pancreatic ribonuclease with an isoaspartyl residue at position 67.
J.Mol.Biol., 257, 1996
|
|
6NM4
 
 | | Crystal structure of SAM-bound PRDM9 in complex with MRK-740 inhibitor | | Descriptor: | 4-[3-(3,5-dimethoxyphenyl)-1,2,4-oxadiazol-5-yl]-1-methyl-9-(2-methylpyridin-4-yl)-1,4,9-triazaspiro[5.5]undecane, Histone-lysine N-methyltransferase PRDM9, S-ADENOSYLMETHIONINE, ... | | Authors: | Ivanochko, D, Halabelian, L, Fischer, C, Sanders, J.M, Kattar, S.D, Brown, P.J, Edwards, A.M, Bountra, C, Arrowsmith, C.H, Structural Genomics Consortium (SGC) | | Deposit date: | 2019-01-10 | | Release date: | 2019-02-13 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.58 Å) | | Cite: | Discovery of a chemical probe for PRDM9.
Nat Commun, 10, 2019
|
|
