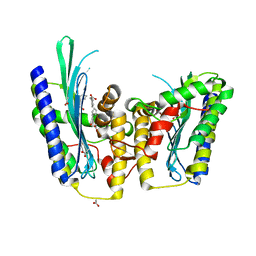
3DWR
 
 | |
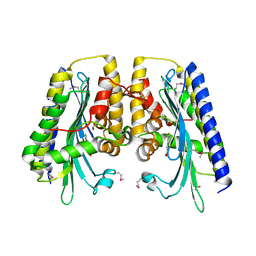
3DWS
 
 | |
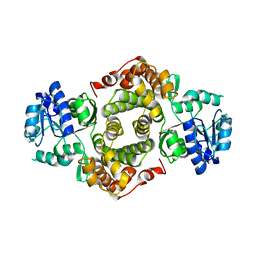
1YJ8
 
 | |
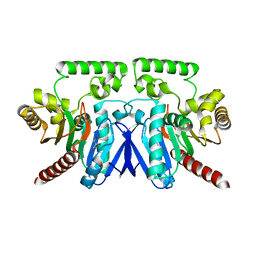
2Q0X
 
 | |
1SQ6
 
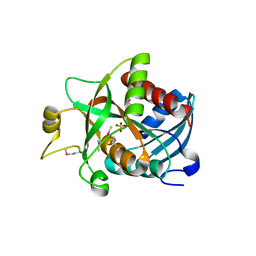 | |
3LJN
 
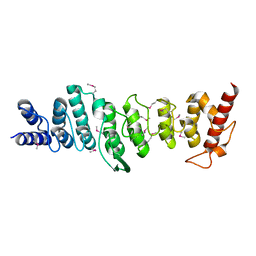 | |
3KPA
 
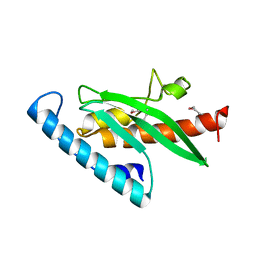 | |
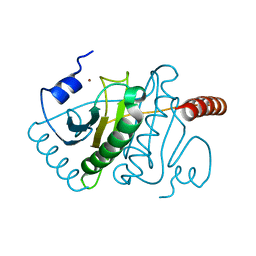
3KSV
 
 | |
2QT8
 
 | |
2QOR
 
 | |
2QAC
 
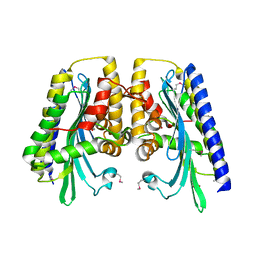
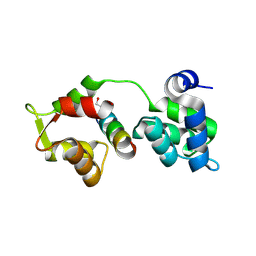 | | The closed MTIP-MyosinA-tail complex from the malaria parasite invasion machinery | | Descriptor: | Myosin A tail domain interacting protein MTIP, Myosin-A | | Authors: | Bosch, J, Turley, S, Roach, C.M, Daly, T.M, Bergman, L.W, Hol, W.G.J, Structural Genomics of Pathogenic Protozoa Consortium (SGPP) | | Deposit date: | 2007-06-14 | | Release date: | 2007-06-26 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Closed MTIP-Myosin A-Tail Complex from the Malaria Parasite Invasion Machinery.
J.Mol.Biol., 372, 2007
|
|
2PC4
 
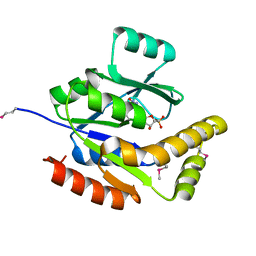
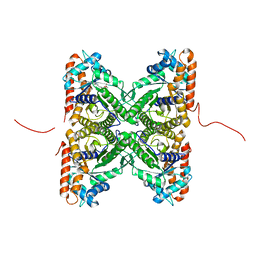 | | Crystal structure of fructose-bisphosphate aldolase from Plasmodium falciparum in complex with TRAP-tail determined at 2.4 angstrom resolution | | Descriptor: | Fructose-bisphosphate aldolase, PbTRAP | | Authors: | Bosch, J, Buscaglia, C.A, Krumm, B, Cardozo, T, Nussenzweig, V, Hol, W.G.J, Structural Genomics of Pathogenic Protozoa Consortium (SGPP) | | Deposit date: | 2007-03-29 | | Release date: | 2007-04-17 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Aldolase provides an unusual binding site for thrombospondin-related anonymous protein in the invasion machinery of the malaria parasite.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
2HQJ
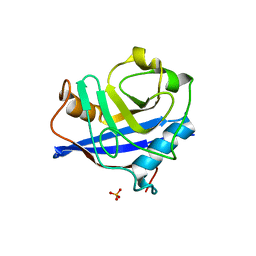 
 | |
1ZSO
 
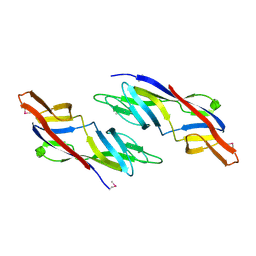 | |
2QVN
 
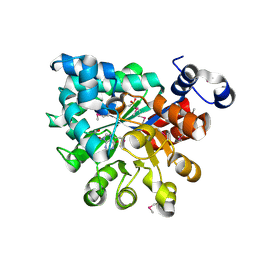 | |
1XN4
 
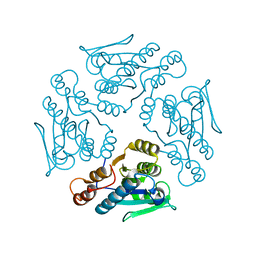 | |
1X9G
 
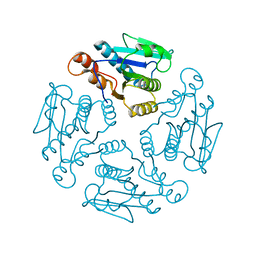 | |
1YZV
 
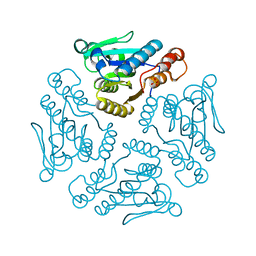 | |
2AR1
 
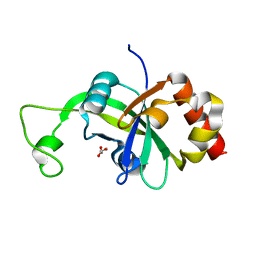 | |
2B4T
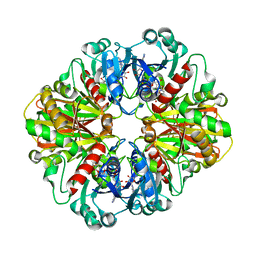 
 | | Crystal structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum at 2.25 Angstrom resolution reveals intriguing extra electron density in the active site | | Descriptor: | 4-(2-AMINOETHYL)BENZENESULFONYL FLUORIDE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, glyceraldehyde-3-phosphate dehydrogenase | | Authors: | Robien, M.A, Bosch, J, Hol, W.G.J, Structural Genomics of Pathogenic Protozoa Consortium (SGPP) | | Deposit date: | 2005-09-26 | | Release date: | 2005-10-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum at 2.25 A resolution reveals intriguing extra electron density in the active site
Proteins, 62, 2006
|
|
2B4G
 
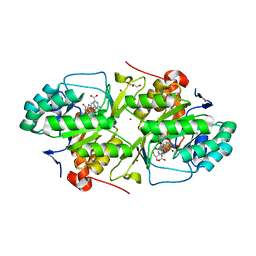 | | dihydroorotate dehydrogenase | | Descriptor: | BROMIDE ION, FLAVIN MONONUCLEOTIDE, GLYCEROL, ... | | Authors: | Arakaki, T.L, Merritt, E.A, Structural Genomics of Pathogenic Protozoa Consortium (SGPP) | | Deposit date: | 2005-09-23 | | Release date: | 2005-10-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Characterization of Trypanosoma brucei dihydroorotate dehydrogenase as a possible drug target; structural, kinetic and RNAi studies
Mol.Microbiol., 68, 2008
|
|
2A0K
 
 | | Crystal structure of Nucleoside 2-deoxyribosyltransferase from Trypanosoma brucei at 1.8 A resolution | | Descriptor: | GLYCEROL, Nucleoside 2-deoxyribosyltransferase, SULFATE ION | | Authors: | Bosch, J, Robien, M.A, Hol, W.G.J, Structural Genomics of Pathogenic Protozoa Consortium (SGPP) | | Deposit date: | 2005-06-16 | | Release date: | 2005-07-26 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Using fragment cocktail crystallography to assist inhibitor design of Trypanosoma brucei nucleoside 2-deoxyribosyltransferase.
J.Med.Chem., 49, 2006
|
|
2B4R
 
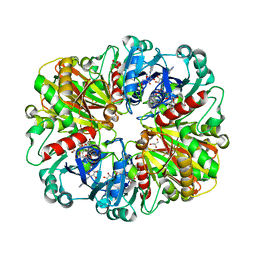 | | Crystal structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum at 2.25 Angstrom Resolution reveals intriguing extra electron density in the active site | | Descriptor: | 4-(2-AMINOETHYL)BENZENESULFONYL FLUORIDE, GLYCEROL, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Robien, M.A, Bosch, J, Hol, W.G.J, Structural Genomics of Pathogenic Protozoa Consortium (SGPP) | | Deposit date: | 2005-09-26 | | Release date: | 2005-10-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Crystal structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum at 2.25 A resolution reveals intriguing extra electron density in the active site
Proteins, 62, 2006
|
|
2AUC
 
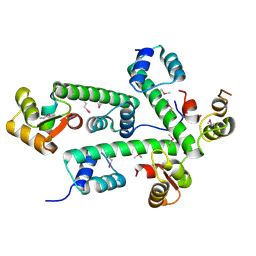 | | Structure of the Plasmodium MTIP-MyoA complex, a key component of the parasite invasion motor | | Descriptor: | Myosin A, Myosin A Tail Interacting Protein | | Authors: | Bosch, J, Turley, S, Hol, W.G.J, Structural Genomics of Pathogenic Protozoa Consortium (SGPP) | | Deposit date: | 2005-08-27 | | Release date: | 2006-01-17 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of the MTIP-MyoA complex, a key component of the malaria parasite invasion motor.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
