6GB1
 
 | | Crystal structure of the GLP1 receptor ECD with Peptide 11 | | Descriptor: | Glucagon-like peptide 1 receptor, HEXANE-1,6-DIOL, Peptide 11, ... | | Authors: | Schreuder, H.A, Liesum, A. | | Deposit date: | 2018-04-13 | | Release date: | 2018-06-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.73 Å) | | Cite: | Dual Glucagon-like Peptide 1 (GLP-1)/Glucagon Receptor Agonists Specifically Optimized for Multidose Formulations.
J. Med. Chem., 61, 2018
|
|
7Z1S
 
 | | X-ray crystal structure of SLPYL1-NIO complex | | Descriptor: | GLYCEROL, NICOTINIC ACID, SlPYL1-NIO | | Authors: | Infantes, L, Albert, A. | | Deposit date: | 2022-02-25 | | Release date: | 2022-07-13 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structure-Based Modulation of the Ligand Sensitivity of a Tomato Dimeric Abscisic Acid Receptor Through a Glu to Asp Mutation in the Latch Loop.
Front Plant Sci, 13, 2022
|
|
7Z1P
 
 | | X-ray crystal structure of SLPYL1-E151D mutant | | Descriptor: | SLPYL1-E151D | | Authors: | Infantes, L, Albert, A. | | Deposit date: | 2022-02-25 | | Release date: | 2022-07-13 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Structure-Based Modulation of the Ligand Sensitivity of a Tomato Dimeric Abscisic Acid Receptor Through a Glu to Asp Mutation in the Latch Loop.
Front Plant Sci, 13, 2022
|
|
7Z1Q
 
 | |
7Z1R
 
 | | X-ray crystal structure of SLPYL1-E151D mutant ABA complex | | Descriptor: | (2Z,4E)-5-[(1S)-1-hydroxy-2,6,6-trimethyl-4-oxocyclohex-2-en-1-yl]-3-methylpenta-2,4-dienoic acid, GLYCEROL, SlPYL1-E151D ABA | | Authors: | Infantes, L, Albert, A. | | Deposit date: | 2022-02-25 | | Release date: | 2022-07-13 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.601 Å) | | Cite: | Structure-Based Modulation of the Ligand Sensitivity of a Tomato Dimeric Abscisic Acid Receptor Through a Glu to Asp Mutation in the Latch Loop.
Front Plant Sci, 13, 2022
|
|
7OVT
 | |
7TP3
 
 | | Crystal structure of SARS-CoV-2 receptor binding domain in complex with neutralizing antibody K288.2 | | Descriptor: | CACODYLATE ION, K288.2 heavy chain, K288.2 light chain, ... | | Authors: | Yuan, M, Zhu, X, Wilson, I.A. | | Deposit date: | 2022-01-24 | | Release date: | 2022-02-23 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Broadly neutralizing antibodies to SARS-related viruses can be readily induced in rhesus macaques.
Sci Transl Med, 14, 2022
|
|
7TP4
 
 | | Crystal structure of SARS-CoV-2 receptor binding domain in complex with neutralizing antibody K398.22 | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, K398.22 heavy chain, ... | | Authors: | Yuan, M, Zhu, X, Wilson, I.A. | | Deposit date: | 2022-01-24 | | Release date: | 2022-02-23 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Broadly neutralizing antibodies to SARS-related viruses can be readily induced in rhesus macaques.
Sci Transl Med, 14, 2022
|
|
9H28
 
 | | Alternative conformation LGTV with TBEV prME | | Descriptor: | 1-PALMITOYL-2-LINOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Genome polyprotein, ... | | Authors: | Bisikalo, K, Rosendal, E. | | Deposit date: | 2024-10-11 | | Release date: | 2024-11-06 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (3.22 Å) | | Cite: | The influence of the pre-membrane and envelope proteins on structure, pathogenicity and tropism of tick-borne encephalitis virus
Biorxiv, 2024
|
|
9FK0
 
 | | LGTV with TBEV prME | | Descriptor: | 1-PALMITOYL-2-LINOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Genome polyprotein, ... | | Authors: | Bisikalo, K, Rosendal, E. | | Deposit date: | 2024-06-01 | | Release date: | 2024-11-06 | | Last modified: | 2024-11-27 | | Method: | ELECTRON MICROSCOPY (3.22 Å) | | Cite: | The influence of the pre-membrane and envelope proteins on structure, pathogenicity and tropism of tick-borne encephalitis virus
Biorxiv, 2024
|
|
9FOJ
 
 | | LGTV TP21. Langat virus, strain TP21 | | Descriptor: | 1-PALMITOYL-2-LINOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Envelope protein E, ... | | Authors: | Bisikalo, K, Rosendal, E. | | Deposit date: | 2024-06-12 | | Release date: | 2024-11-06 | | Last modified: | 2024-11-27 | | Method: | ELECTRON MICROSCOPY (3.82 Å) | | Cite: | The influence of the pre-membrane and envelope proteins on structure, pathogenicity and tropism of tick-borne encephalitis virus
Biorxiv, 2024
|
|
8E1O
 
 | | Crystal structure of hTEAD2 bound to a methoxypyridine lipid pocket binder | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 5-methoxy-N-({3-[2-(methylamino)-2-oxoethyl]phenyl}methyl)-4-{(E)-2-[trans-4-(trifluoromethyl)cyclohexyl]ethenyl}pyridine-2-carboxamide, Transcriptional enhancer factor TEF-4 | | Authors: | Noland, C.L, Dey, A, Zbieg, J, Crawford, J. | | Deposit date: | 2022-08-10 | | Release date: | 2023-08-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Targeting the Hippo pathway in cancers via ubiquitination dependent TEAD degradation
Biorxiv, 2024
|
|
6AX5
 
 | | RPT1 region of INI1/SNF5/SMARCB1_HUMAN - SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily B member 1. | | Descriptor: | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily B member 1 | | Authors: | Girvin, M.E, Cahill, S.M, Harris, R, Cowburn, D, Spira, M, Wu, X, Prakash, R, Bernowitz, M, Almo, S.C, Kalpana, G.V. | | Deposit date: | 2017-09-06 | | Release date: | 2017-10-18 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | INI1/SMARCB1 Rpt1 domain mimics TAR RNA in binding to integrase to facilitate HIV-1 replication.
Nat Commun, 12, 2021
|
|
9CDX
 
 | | Crystal structure of DLK with inhibitor bound | | Descriptor: | (5P)-5-[(4R)-6-(propan-2-yl)imidazo[1,5-a]pyridin-1-yl]-3-(trifluoromethyl)pyridin-2-amine, Mitogen-activated protein kinase kinase kinase 12 | | Authors: | Skene, R.J, Bell, J.A. | | Deposit date: | 2024-06-25 | | Release date: | 2025-02-19 | | Last modified: | 2025-02-26 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | In Silico Enabled Discovery of KAI-11101, a Preclinical DLK Inhibitor for the Treatment of Neurodegenerative Disease and Neuronal Injury.
J.Med.Chem., 68, 2025
|
|
9CDY
 
 | | Crystal structure of DLK with inhibitor bound | | Descriptor: | 1,2-ETHANEDIOL, 3-{(3M)-3-[6-amino-5-(trifluoromethoxy)pyridin-3-yl]-1-ethyl-7-oxo-1,7-dihydro-6H-pyrazolo[3,4-c]pyridin-6-yl}-1,5-anhydro-3,4-dideoxy-2-O-methyl-D-erythro-pentitol, MAGNESIUM ION, ... | | Authors: | Skene, R.J, Bell, J.A. | | Deposit date: | 2024-06-25 | | Release date: | 2025-02-19 | | Last modified: | 2025-02-26 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | In Silico Enabled Discovery of KAI-11101, a Preclinical DLK Inhibitor for the Treatment of Neurodegenerative Disease and Neuronal Injury.
J.Med.Chem., 68, 2025
|
|
8VWX
 
 | | Human Bcl-2 (G101V Mutant)/Bcl-xL Chimera Fused to Maltose-Binding Protein | | Descriptor: | Maltose/maltodextrin-binding periplasmic protein fused to apoptosis regulator Bcl-2/Bcl-xL chimera | | Authors: | Baird, J, Holliday, M. | | Deposit date: | 2024-02-02 | | Release date: | 2024-10-02 | | Last modified: | 2025-03-12 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Hydrogen/Deuterium Exchange and Protein Oxidative Footprinting with Mass Spectrometry Collectively Discriminate the Binding of Small-Molecule Therapeutics to Bcl-2.
Anal.Chem., 97, 2025
|
|
8VXN
 
 | | Human Bcl-2/Bcl-xL Chimera Fused to Maltose-Binding Protein | | Descriptor: | Maltose/maltodextrin-binding periplasmic protein fused to apoptosis regulator Bcl-2/Bcl-xL chimera | | Authors: | Baird, J, Holliday, M. | | Deposit date: | 2024-02-05 | | Release date: | 2024-10-02 | | Last modified: | 2025-03-12 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Hydrogen/Deuterium Exchange and Protein Oxidative Footprinting with Mass Spectrometry Collectively Discriminate the Binding of Small-Molecule Therapeutics to Bcl-2.
Anal.Chem., 97, 2025
|
|
8VXM
 
 | | Human Bcl-2/Bcl-xL Chimera Fused to MBP in Complex with Inhibitor S55746 | | Descriptor: | DI(HYDROXYETHYL)ETHER, Maltose/maltodextrin-binding periplasmic protein fused to apoptosis regulator Bcl-2/Bcl-xL chimera, ~{N}-(4-hydroxyphenyl)-3-[6-[[(3~{S})-3-(morpholin-4-ylmethyl)-3,4-dihydro-1~{H}-isoquinolin-2-yl]carbonyl]-1,3-benzodioxol-5-yl]-~{N}-phenyl-5,6,7,8-tetrahydroindolizine-1-carboxamide | | Authors: | Baird, J, Holliday, M. | | Deposit date: | 2024-02-05 | | Release date: | 2024-10-02 | | Last modified: | 2025-03-12 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Hydrogen/Deuterium Exchange and Protein Oxidative Footprinting with Mass Spectrometry Collectively Discriminate the Binding of Small-Molecule Therapeutics to Bcl-2.
Anal.Chem., 97, 2025
|
|
8VWZ
 
 | | Human Bcl-2 (G101V Mutant)/Bcl-xL Chimera Fused to MBP in Complex with Inhibitor S55746 | | Descriptor: | DI(HYDROXYETHYL)ETHER, Maltose/maltodextrin-binding periplasmic protein fused to apoptosis regulator Bcl-2/Bcl-xL chimera, ~{N}-(4-hydroxyphenyl)-3-[6-[[(3~{S})-3-(morpholin-4-ylmethyl)-3,4-dihydro-1~{H}-isoquinolin-2-yl]carbonyl]-1,3-benzodioxol-5-yl]-~{N}-phenyl-5,6,7,8-tetrahydroindolizine-1-carboxamide | | Authors: | Baird, J, Holliday, M. | | Deposit date: | 2024-02-02 | | Release date: | 2024-10-02 | | Last modified: | 2025-03-12 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Hydrogen/Deuterium Exchange and Protein Oxidative Footprinting with Mass Spectrometry Collectively Discriminate the Binding of Small-Molecule Therapeutics to Bcl-2.
Anal.Chem., 97, 2025
|
|
9EUU
 
 | | Structure of recombinant alpha-synuclein fibrils 1B capable of seeding GCIs in vivo | | Descriptor: | Alpha-synuclein | | Authors: | Burger, D, Kashyrina, M, Lewis, A, De Nuccio, F, Mohammed, I, de La Seigliere, H, van den Heuvel, L, Feuillie, C, Verchere, J, Berbon, M, Arotcarena, M, Retailleau, A, Bezard, E, Laferriere, F, Loquet, A, Bousset, L, Baron, T, Lofrumento, D.D, De Giorgi, F, Stahlberg, H, Ichas, F. | | Deposit date: | 2024-03-28 | | Release date: | 2024-06-05 | | Last modified: | 2025-06-18 | | Method: | ELECTRON MICROSCOPY (1.93 Å) | | Cite: | 1.94 angstrom structure of synthetic alpha-synuclein fibrils seeding MSA neuropathology
Biorxiv, 2024
|
|
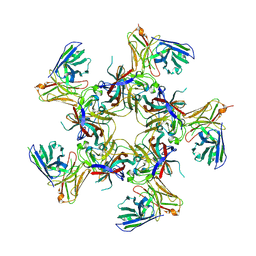
8U5L
 
 | |
4N48
 
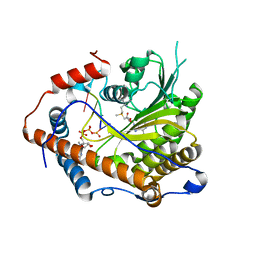 | | Cap-specific mRNA (nucleoside-2'-O-)-methyltransferase 1 Protein in complex with capped RNA fragment | | Descriptor: | 7N-METHYL-8-HYDROGUANOSINE-5'-TRIPHOSPHATE, Cap-specific mRNA (nucleoside-2'-O-)-methyltransferase 1, S-ADENOSYLMETHIONINE, ... | | Authors: | Smietanski, M, Werener, M, Purta, E, Kaminska, K.H, Stepinski, J, Darzynkiewicz, E, Nowotny, M, Bujnicki, J.M. | | Deposit date: | 2013-10-08 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.704 Å) | | Cite: | Structural analysis of human 2'-O-ribose methyltransferases involved in mRNA cap structure formation.
Nat Commun, 5, 2014
|
|
8VBL
 
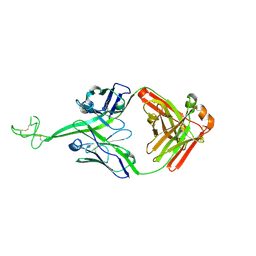 | | Structure of bovine anti-HIV Fab ElsE6 | | Descriptor: | Bovine Fab ElsE6 heavy chain, Bovine Fab ElsE6 light chain | | Authors: | Stanfield, R.L, Wilson, I.A. | | Deposit date: | 2023-12-12 | | Release date: | 2024-09-04 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Immunization of cows with HIV envelope trimers generates broadly neutralizing antibodies to the V2-apex from the ultralong CDRH3 repertoire.
Plos Pathog., 20, 2024
|
|
8VBR
 
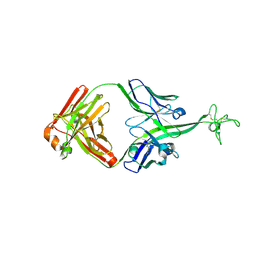 | | Structure of bovine anti-HIV Fab ElsE11 | | Descriptor: | Bovine Fab ElsE11 light chain, Bovine Fab Else11 heavy chain | | Authors: | Stanfield, R.L, Wilson, I.A. | | Deposit date: | 2023-12-12 | | Release date: | 2024-09-04 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Immunization of cows with HIV envelope trimers generates broadly neutralizing antibodies to the V2-apex from the ultralong CDRH3 repertoire.
Plos Pathog., 20, 2024
|
|
8VBM
 
 | | Structure of bovine anti-HIV Fab ElsE7 | | Descriptor: | Bovine Fab ElsE7 heavy chain, Bovine Fab ElsE7 light chain | | Authors: | Stanfield, R.L, Wilson, I.A. | | Deposit date: | 2023-12-12 | | Release date: | 2024-09-04 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Immunization of cows with HIV envelope trimers generates broadly neutralizing antibodies to the V2-apex from the ultralong CDRH3 repertoire.
Plos Pathog., 20, 2024
|
|
